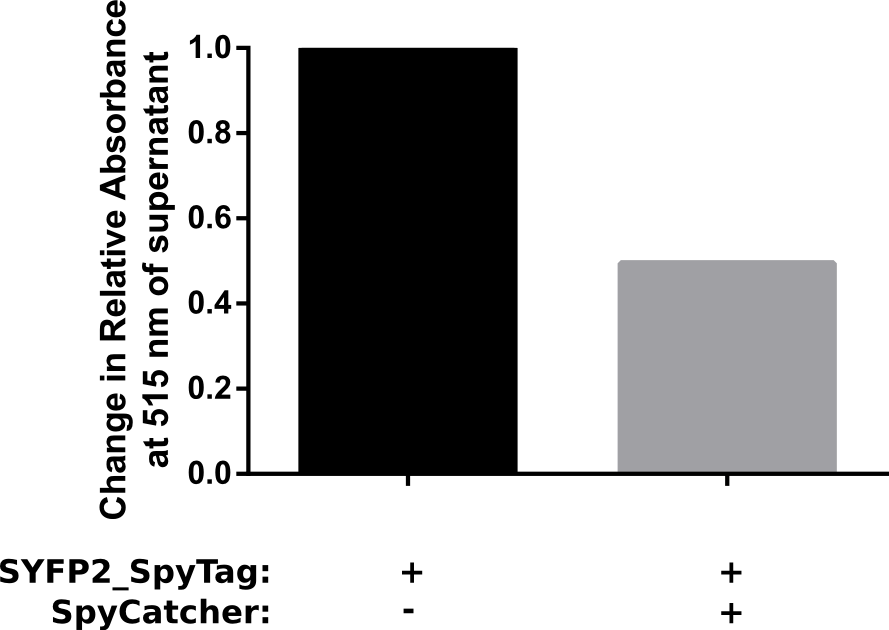Part:BBa_K1159201
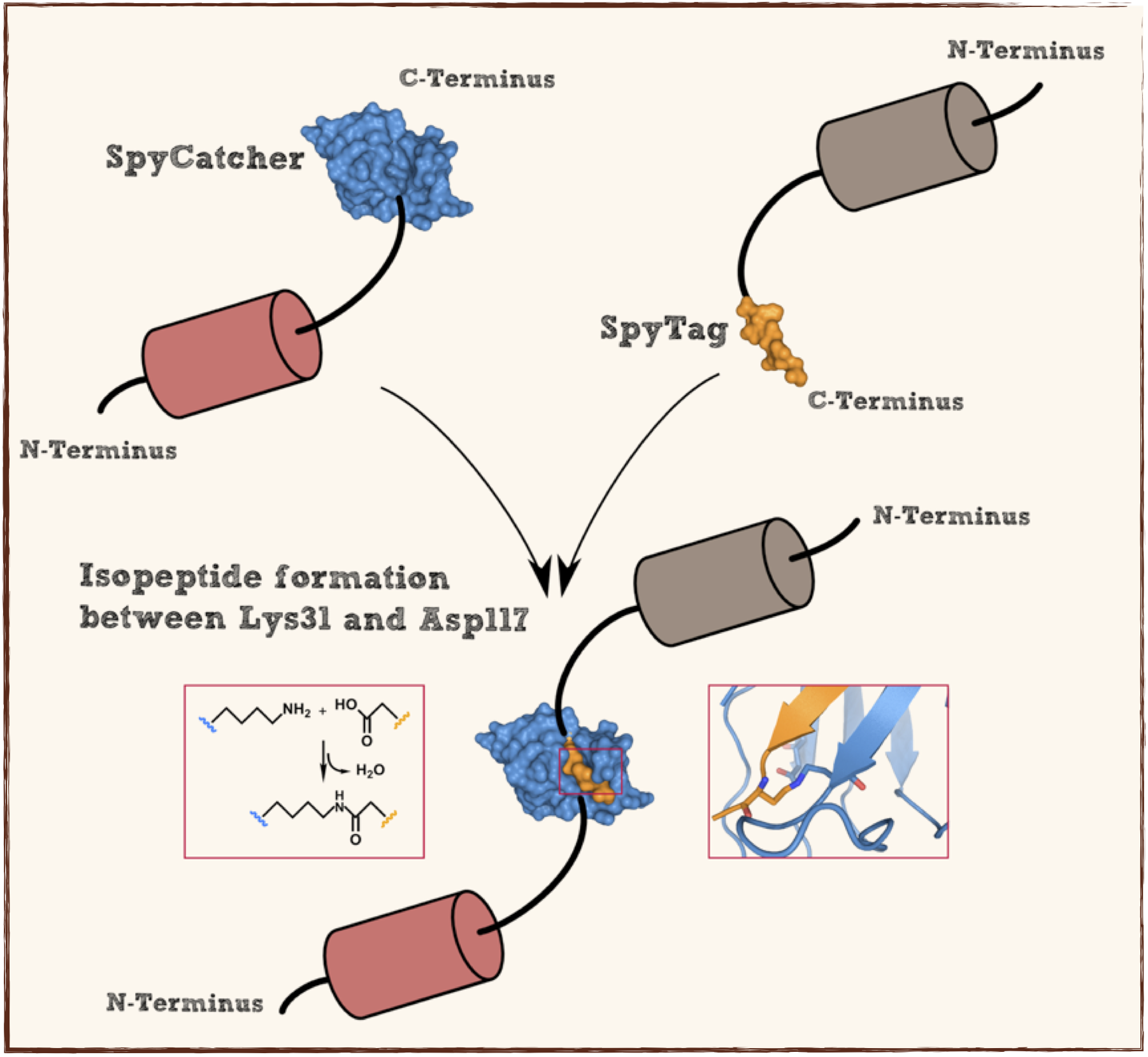
Splitted and engineered C-terminal FbaB for isopeptide bound formation (SpyTag) in RFC[25]
This part codes for a oligopeptide that is recognized and bound by a covalent isopeptide bound to a protein, called SpyCatcher. That catcher protein for this oligopeptdie can be found under BBa_K1159200. This part is flanked by RFC[25] pre- and suffix for further protein fusions.
This part codes for a protein that recognizes and forms a covalent isopeptide bound to a oligopeptide. That oligopeptide can be found under BBa_K1159200. In contrast to the original SpyCatcher this part only contains the essential part for forming isopeptide bounds and does not contain a N-terminal His-tag and TEV cleavage site. Furthermore two amino acids were exchanged to improve the function, Ile34 was replaced by Glu34 which enhances the reaction rate, also the exposed hydrophobic redidue Met69 was replaced by Tyr69. This part is flanked by RFC[25] pre- and suffix for further protein fusions.
Experimental Evidence
Proper funtion of the construct was verified in a pulldown assay using 10 μM of SpyCatcher with N-terminal HisTag which was incubated with 5 μM of C-terminal spytagged SYFP2 for 1 h in PBS. Afterwards HisTag beads were added to bind the fusion product, resulting in a decrease of spytagged SYFP2 in the supernatant. The results of this assay are shown in Figure 2. The results show that after one hour at room temeperature, a two fold change in relative absorbance was observed. This verifies that the construct is working as intended.
Furthermore the two interaction partners were mixed, incubated and analyzed via SDS Page (reducing conditions) and coomassie staining.
Reference
http://www.ncbi.nlm.nih.gov/pubmed/22366317 Zakeri et al., 2012 Zakeri B, Fierer JO, Celik E, Chittock EC, Schwarz-Linek U, Moy VT, Howarth M. (2012). Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. Proc Natl Acad Sci U S A. 20;109(12)
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
| Protein data table for BioBrick BBa_ automatically created by the BioBrick-AutoAnnotator version 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide sequence in RFC 25, so ATGGCCGGC and ACCGGT were added (in italics) to the 5' and 3' ends: (underlined part encodes the protein) ATGGCCGGCGCTCATATT ... CCAACTAAGACCGGT ORF from nucleotide position -8 to 45 (excluding stop-codon) | ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid sequence: (RFC 25 scars in shown in bold, other sequence features underlined; both given below)
| ||||||||||||||||||||||||||||||||||||||||||||||
Sequence features: (with their position in the amino acid sequence, see the list of supported features)
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid composition:
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid counting
| Biochemical parameters
| |||||||||||||||||||||||||||||||||||||||||||||
| Plot for hydrophobicity, charge, predicted secondary structure, solvent accessability, transmembrane helices and disulfid bridges | ||||||||||||||||||||||||||||||||||||||||||||||
Codon usage
| ||||||||||||||||||||||||||||||||||||||||||||||
Alignments (obtained from PredictProtein.org)
| ||||||||||||||||||||||||||||||||||||||||||||||
| Predictions (obtained from PredictProtein.org) | ||||||||||||||||||||||||||||||||||||||||||||||
Subcellular Localization (reliability in brackets)
| Gene Ontology (reliability in brackets)
| |||||||||||||||||||||||||||||||||||||||||||||
Predicted features:
| ||||||||||||||||||||||||||||||||||||||||||||||
| The BioBrick-AutoAnnotator was created by TU-Munich 2013 iGEM team. For more information please see the documentation. If you have any questions, comments or suggestions, please leave us a comment. | ||||||||||||||||||||||||||||||||||||||||||||||
| None |