Part:BBa_K4273020
pTDH3-Np5598-tTDH1-pPGK1-NlmysD-tPGK1
Np5598 is a gene found in cyanobacteria Nostoc punctiforme that encodes for AGL whereas NlmysD encodes for AlaL in Nostoc linckia. AGL-AlaL allows 4-DG to be converted into the MAAs shinorine or porphyra-334. In our experiment, we found that AlaL encoded by NlmysD has a strong preference toward the amino acid threonine. Therefore, this part could efficiently produce the MAA porphyra-334 primarily, making it the first successful case of producing pure samples of porphyra-334. This part is within our part collection that allows efficiently production of MAAs in S. cerevisiae.
Our part collection contains necessary genes to produce gadusol and the MAAs shinorine, porphyra-334, and palythine at a high rate. Xyl1, Xyl2, and Xyl3 are genes that allow S. cerevisiae to utilize xylose to produce S7P. DDGS and OMT converts S7P to the precursor of MAA, 4-deoxygadusol (4-DG). AGL converts 4-DG into M-glycine (MG). AlaL, by adding either serine or threonine, produces shinorine and porphyra-334, respectively. MysH could be added to the circuit of shinorine to produce palythine. S7P could also be converted into gadusol under the catalyzation of EEVS and M-Tox. In this part collection, we included multiple pathways and methods to increase the production of the upstream S7P and downstream MAAs.
The part collection includes BBa_K4273017, BBa_K4273018, BBa_K4273019, BBa_K4273020,
BBa_K4273021 , BBa_K4273016 and BBa_K4273000. This collection can provide inspiration and efficient methods to utilize the penta phosphate pathway or to produce other types of MAAs in S. cerevisiae for other teams.
Usage and Biology
We selected promoters pTDH3 and pPGK1 due to their stability expression in S. cerevisiae (Apel et. al., 2016). These promoters are shown to have stable and strong expression in YPD culture mediums. Among the three, pTDH3 has highest stability and strength, followed by pPGK1. For expression of AGL and AlaL enzymes, we used pTDH3 for AGL and pPGK1 for AlaL. In order to optimize our production, we inserted this part into the yeast’s genome at position 106, chromosome I (Apel et. al., 2016).
Experiment
Design and Absorbance of Nine AGL-AlaL Combinations
We expressed AGL-AlaL genes first using plasmid vector and then through genome insertion. Due to the existence of multiple different types of AGL and AlaL with different efficiency and amino acid preference in nature, we selected ligases from three different marine organisms: Nostoc punctiform (Np5598 and Np5597), Nostoc linckia (NlmysC and NImysD) and Actinosynnema mirum (Am4257 and Am4256), and expected to create nine combinations of AGL-AlaL. We used Lee's yeast toolkit to produce Level1 plasmids containing only AGL or AlaL, and used Golden Gate assembly on the basis of Level1 to construct Level2 plasmid containing the 9 combinations of AGL and AlaL. Finally, we transformed these Level2 plasmids into SC.L3 strains, which are S. cerevisiae which could produce 4-DG efficiently from the addition of DDGS and OMT, to form strain SC.L5.

Shinorine and porphyra-334 has similar absorption curves with an absorption peak at 334 nm. We cultivated the SC.L5 yeast using a SC-Ura culture medium with 1% glucose and 1% xylose and tested the absorption spectrum of the supernatant of the fermentation broth. Other than the Am4257-Am4256 combination yeast which failed to grow, there is an obvious absorption peak of 334 nm for the other 8 combinations. We found that the absorption peak of AGL Np5598 series was significantly higher than that of NlmysC and Am4257. In ALAL series, NlmysD has the highest absorption peak, followed by Am4256 and Np5597. We selected the 6 samples with higher peak and lysed them to preform OD scanning on the lysate. The scanning results after lysis further performed that Np5598 is the most efficient AGL and NlmysD is the most efficient AlaL. We think we have found an optimal combination of these two enzymes and have successfully produced MAAs.
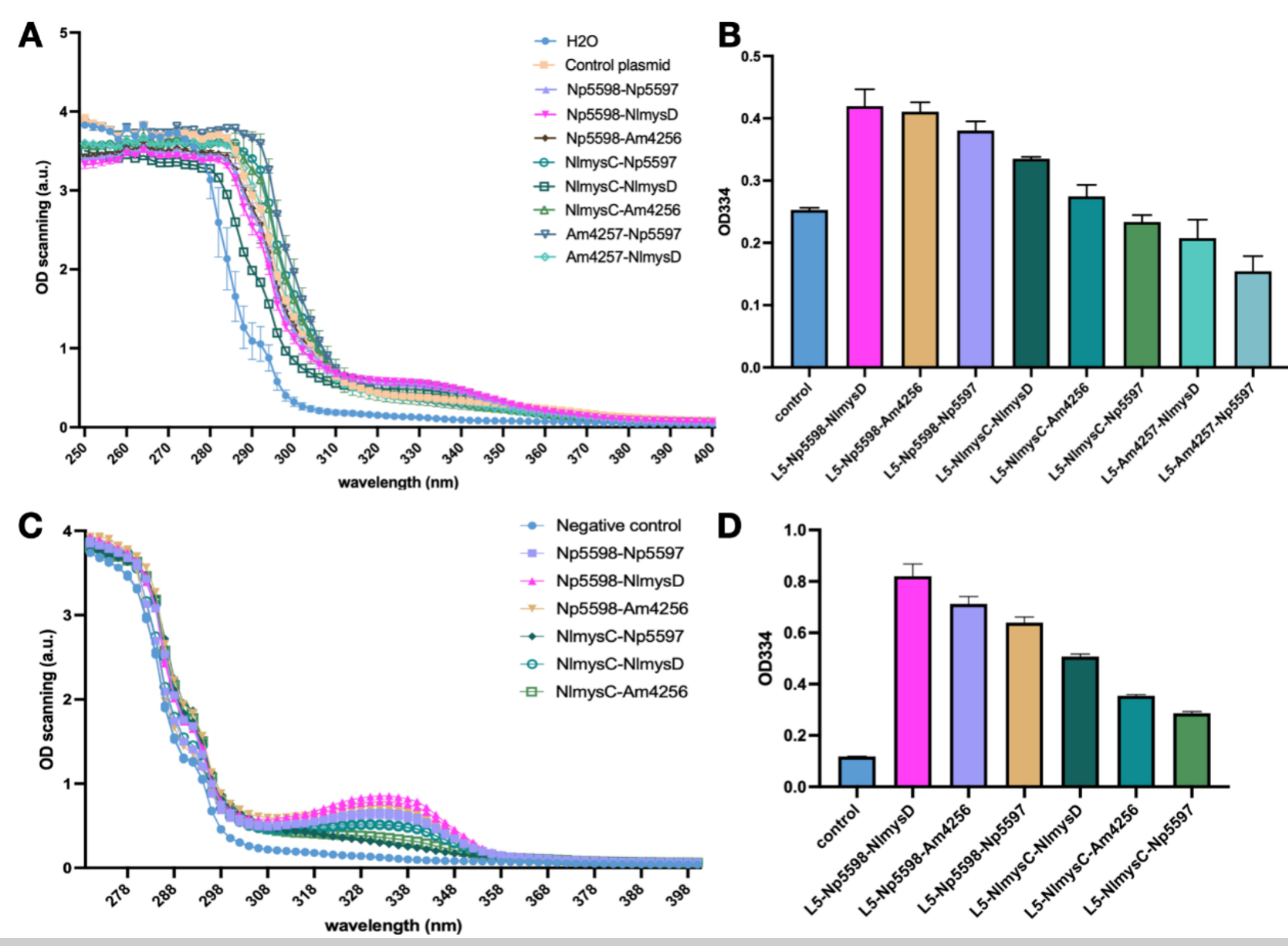
Determining the Amino Acid Preference of Np5597 and NlmysD
Moreover, because AlaL has a preference for different amino acids to produce shinorine and porphyra-334, we need to further confirm the type of MAA produced by different strains. To do so, we used High-performing Liquid Chromatology (HPLC) and Mass Spectrometer (MS) technology. Due to the lack of standard samples of these MAAs in the market, we extracted MAAs from nori samples using methods we learnt through our human practice activities and used the extractives as standard. We found that the metabolite of Np5598-NlmysD mainly has the absorption spectrum of porphyra-334 according to HPLC, and the result of MS also proved that mainly porphyra-334 exist in the metabolite (m/q = 347). This shows that NlmysD has a strong selective preference toward the amino acid Threonine, and thus will mainly produce porphyra-334 if both Threonine and Serine are present in the environment. This is the first successful case of producing mainly 334 only. Np5597, on the other hand, shows shinorine's absorption peak, meaning it has preference toward serine. Therefore, we decided to use Np5598-NlmysD for porphyra-334 production and Np5598-NlmysD for shinorine production.

Optimization Through Genome Insertion
Considering the instability and lower efficiency of using plasmid expresssion, we decided to insert Np5598-NlmysD and Np5598-Np5597, the most efficient combinations to produce porphyra-334 and shinorine respectively (Figure 9 & 10) into SC.L6's genome at chromosome I, position 106 (SC.L6 is SC.L5 with OMT-DDGS inserted at Nqm1 in order to increase MAAs production). We named the new strains SC.L8 series. After fermentation for 72 hours in YPD medium, we compared the OD 334 absorption of the broth's supernatant between SC.L7 series, SC.L6 with the plasmids pYT-pTDH3-Np5598-tTDH1-pPGK1-AlaL-tPGK1 (AlaL includes Np5597 and NlmysD) and SC.L8 series. We found that the OD value of L8 series (for both Np5598-NlmysD and Np5598-Np5597) with genome recombinant, showed a steady increase of approximately 20% compared with that of L7 series with plasmid expression. Finally, the nori extract with a higher purity of shinorine and porphyra-334 is used as the standard sample to obtain rough yield data of SC.L8 series. Porphyra-334’s yield reached 270.7mg/L in L8-Np5598-NlmysD.
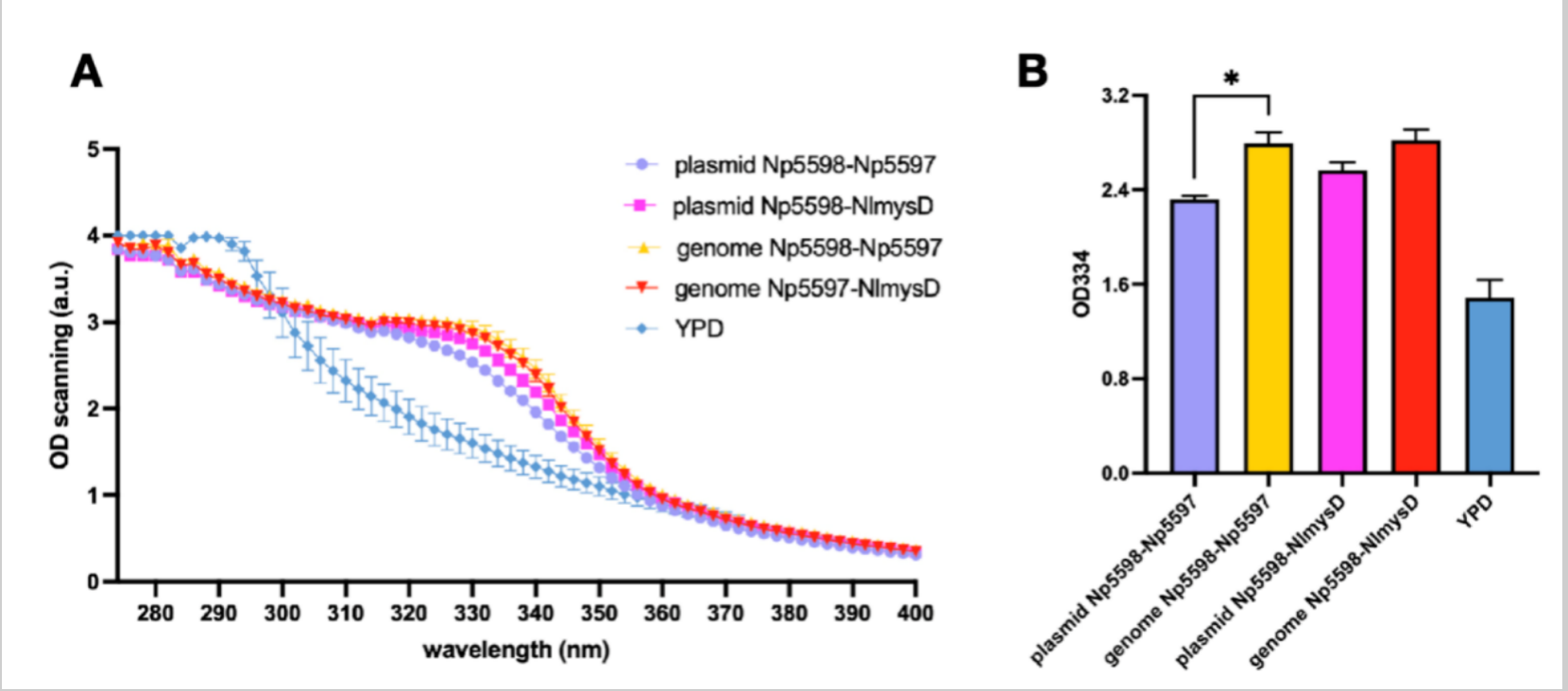
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 3423
Illegal XhoI site found at 2075 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 2321
Illegal BsaI.rc site found at 3433
Reference
Park SH, Lee K, Jang JW, Hahn JS. Metabolic Engineering of Saccharomyces cerevisiae for Production of Shinorine, a Sunscreen Material, from Xylose. ACS Synth Biol. 2019;8(2):346-357.
Jin C, Kim S, Moon S, Jin H, Hahn JS. Efficient production of shinorine, a natural sunscreen material, from glucose and xylose by deleting HXK2 encoding hexokinase in Saccharomyces cerevisiae. FEMS Yeast Res. 2021;21(7):foab053.
Chen M, Rubin GM, Jiang G, Raad Z, Ding Y. Biosynthesis and Heterologous Production of Mycosporine-Like Amino Acid Palythines. J Org Chem. 2021 Aug 20;86(16):11160-11168.
Osborn AR, Almabruk KH, Holzwarth G, Asamizu S, LaDu J, Kean KM, Karplus PA, Tanguay RL, Bakalinsky AT, Mahmud T. De novo synthesis of a sunscreen compound in vertebrates. Elife. 2015 May 12;4:e05919.
Reider Apel A, d'Espaux L, Wehrs M, Sachs D, Li RA, Tong GJ, Garber M, Nnadi O, Zhuang W, Hillson NJ, Keasling JD, Mukhopadhyay A. A Cas9-based toolkit to program gene expression in Saccharomyces cerevisiae. Nucleic Acids Res. 2017 Jan 9;45(1):496-508.
Zhang H, Jiang Y, Zhou C, Chen Y, Yu G, Zheng L, Guan H, Li R. Occurrence of Mycosporine-like Amino Acids (MAAs) from the Bloom-Forming Cyanobacteria Aphanizomenon Strains. Molecules. 2022 Mar 7;27(5):1734.
Cress BF, Toparlak ÖD, Guleria S, et al. CRISPathBrick: Modular Combinatorial Assembly of Type II-A CRISPR Arrays for dCas9-Mediated Multiplex Transcriptional Repression in E. coli. ACS Synth Biol. 2015;4(9):987-1000.
| None |
