Part:BBa_R0061
Promoter (HSL-mediated luxR repressor)
This part involves the -10 binding site, the -35 binding site, and the twenty nucleotides between that constitute the lux box. With this part, LuxR functions as a acyl-homoserine lactone-dependent repressor. LuxR resonds to the HSL produced by LuxI, N-(3-oxohexanoyl)-HSL. The Lux box is positioned such that it partially overlaps the consensus -35 and -10 hexamers of an RNA polymerase binding site.
Usage and Biology
A quorum-sensing system involving LuxR, the transcriptional activator, and an acyl-homserine lactone signal regulate the lux operon in vibrio fischeri. In [http://en.wikipedia.org/wiki/Vibrio_fischeri vibrio fischeri], the lux box, which is a 20-base inverted repeat unit, is positioned 42.5 bases upstream of the transcriptional start of the lux operon and is required for transcriptional activation. Egland and Greenberg constructed an artificial lacZ promoter with the lux box positioned between and overlapping the -35 and -10 hexamers of the RNA polymerase binding site was constructed, and LuxR functioned as an acyl-homoserine lactone-dependent repressor at this promoter. It was found that full-length LuxR by itself can bind to lux box-containing DNA. (see reference below)
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Contribution
Group: Valencia_UPV iGEM 2018
Author: Adrián Requena Gutiérrez, Carolina Ropero, Carlos Andreu Vilarroig
Summary: We adapted the part to be able to assemble transcriptional units with the Golden Gate assembly method
Documentation:
In order to create our complete [http://2018.igem.org/Team:Valencia_UPV/Part_Collection part collection] of parts compatible with the Golden Gate assembly method, we made the part BBa_K2656002 which is this part adapted to the Golden Gate technology.
In order characterize this promoter we constructed the composite part BBa_K2656116 using the following RBS, CDS and terminator:
- RBS BBa_K2656009: the B0030 RBS in its Golden Braid compatible version from our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection Part Collection]
- CDS BBa_K2656022: the E0040 GFPmut3b sequence in its Golden Braid standardized version from our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection Part Collection]
- Terminator BBa_K2656026: the B0015 transcriptional terminator in its Golden Braid compatible version from our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection Part Collection]
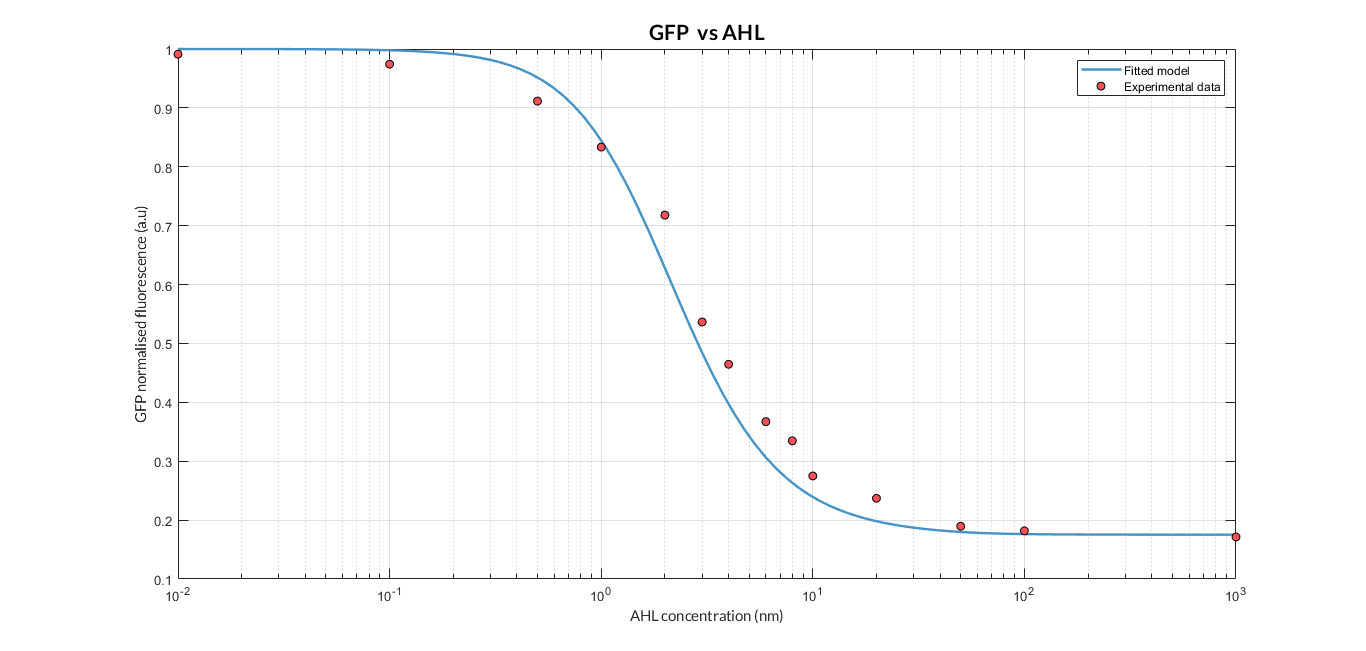
By using this [http://2018.igem.org/Team:Valencia_UPV/Experiments#exp_protocol experimental protocol] and inducing at different AHL concentrations, we have obtained the following experimental measures (in blue dots). These measurements were compared with our [http://2018.igem.org/Team:Valencia_UPV/Modeling#models constitutive AHL-LuxR model] to test it.
Characterization by BS_United_China 2021
See detail in part BBa_K3882002 In our design, we also used LuxR regulatory protein to identify AHL in the environment. Through a series of experiments, we proved that LuxR protein and LuxR operon can correctly activate downstream genes.The downstream PVDQ-eGFP protein activated by the LuxR operon smoothly initiates transcription
//chassis/prokaryote/ecoli
//classic/regulatory/uncategorized
//direction/forward
//promoter
//regulation/negative
//rnap/prokaryote/ecoli/sigma70
| biology | |
| direction | Forward |
| negative_regulators | 1 |
| positive_regulators |

 1 Registry Star
1 Registry Star