Part:BBa_K3629016
Modified T. reesei EGI expression construct with gibson homology sequences and 6X His tag
Usage and Biology
Fully functional cellulase is composed of:
- Endoglucanases (EG) which randomly cleave internal beta-bonds of cellulose polymers to make them shorter
- Cellobiohydrolases (CBH or exoglucanases) which cleave the shorter polymers to make cellobiose
- CBHI= Acts on reducing end of sugar molecule
- CBHII= Acts on non-reducing end of sugar molecule
- Beta-glucosidases (BGS) which cleave the cellobiose disaccharide to free glucose units
These proteins must be in the correct proportions to each other to efficiently degrade cellulose.
This expression construct can be used in the assembly of a EG gene cassette containing Modified TrEGI (BBa_K3629016) and TrEGII (BBa_K3629017). This gene cassette is then intended to be transformed into Y. lipolytica to create a EG- producing strain. This strain should then be co-cultured with two other strains with either a CBH or BGS gene cassette. The three strains together will be able to survive on cellulose media.
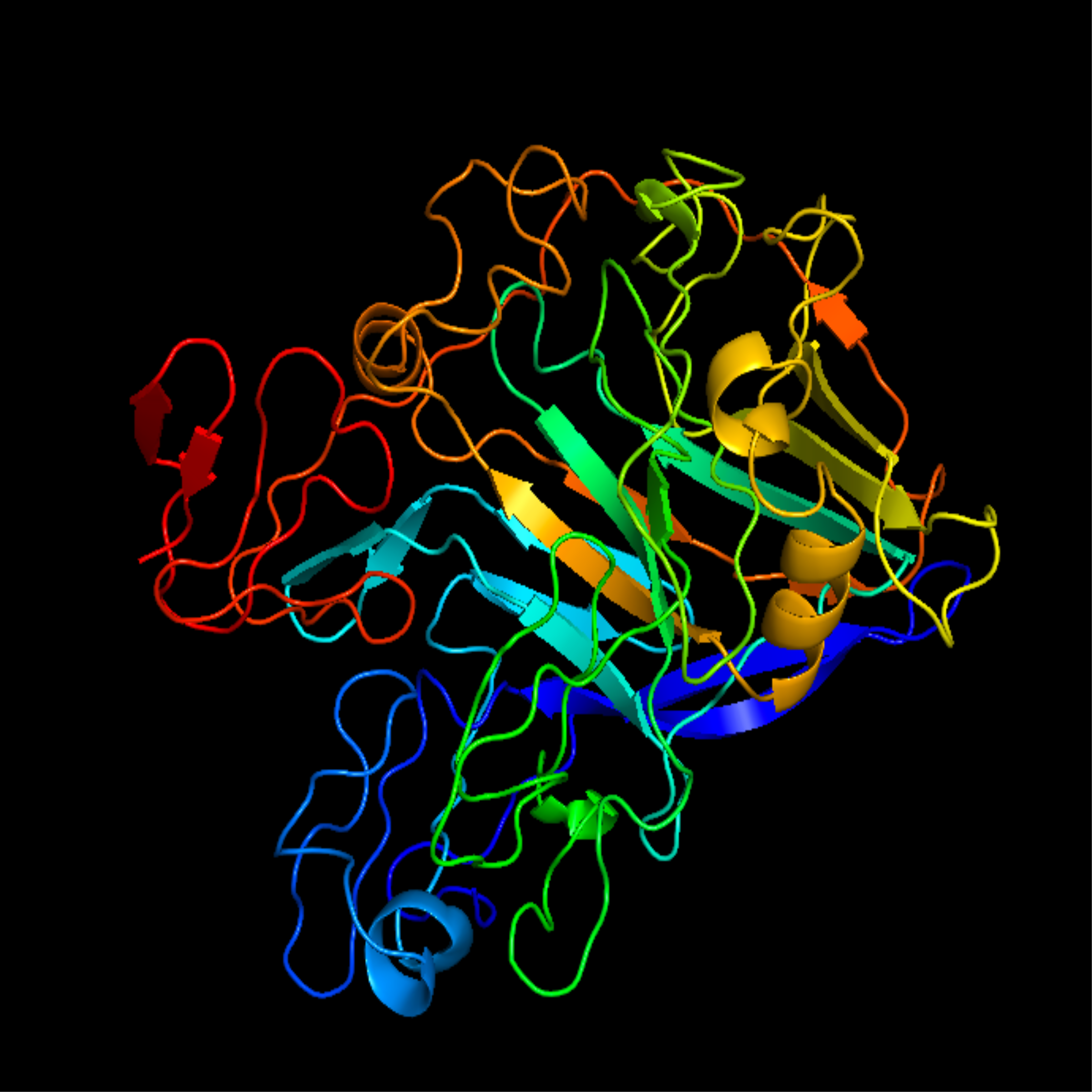
CHIMERIC PROTEIN CREATION AND MODELLING
When developing the sequence for the modified TrEGII (called SEGI8), we first focused on the catalytic domain. We utilized the SEGI-8 modified domain as it has been proven to work with higher efficiency in a broader range of conditions (3). For the linker, we opted to move forward with the ApCel5A linker. The ApCel5A linker was selected due to its flexibility and length, which have been shown to enhance protein efficiency with a feedstock that contains lignin. Finally, the wild type Endoglucanase I cellulose-binding module was selected to ensure the synergy between the catalytic domain and binding module was maintained.
For the modified and wild type enzyme, we predicted the three-dimensional structure using homology modelling. From these structures, we completed molecular dynamic simulations. These simulations were then characterized by GausHaus to determine if the changes increased variance in the enzyme dynamics. The modifications actually had a net lowering of the variance in the enzyme allowing the team to move forward confident in the modifications.
Design
The native signal peptide from T. reesei was removed so it would not interfere with the fused Lip2 secretion tag native to Y. lipolytica.
Sequence and Features
- 10INCOMPATIBLE WITH RFC[10]Illegal prefix found in sequence at 1
Illegal suffix found in sequence at 2385
Illegal EcoRI site found at 2204 - 12INCOMPATIBLE WITH RFC[12]Illegal EcoRI site found at 1
Illegal EcoRI site found at 2204
Illegal NheI site found at 81
Illegal SpeI site found at 2386
Illegal PstI site found at 2400
Illegal NotI site found at 7
Illegal NotI site found at 2393 - 21INCOMPATIBLE WITH RFC[21]Illegal EcoRI site found at 1
Illegal EcoRI site found at 2204
Illegal BamHI site found at 2334
Illegal XhoI site found at 132 - 23INCOMPATIBLE WITH RFC[23]Illegal prefix found in sequence at 1
Illegal suffix found in sequence at 2386
Illegal EcoRI site found at 2204 - 25INCOMPATIBLE WITH RFC[25]Illegal prefix found in sequence at 1
Illegal EcoRI site found at 2204
Illegal XbaI site found at 16
Illegal SpeI site found at 2386
Illegal PstI site found at 2400
Illegal NgoMIV site found at 1326 - 1000COMPATIBLE WITH RFC[1000]
| None |