Part:BBa_K2997008
Native promoter-pchR-sfGFP
BBa_K2997008 hello
pchR Achievements:
Construction of pchR-GFP BBa_K2997008 and confirm by double digestion.
The pchR-GFP (BBa_K2997008) was constructed from IDT DNA synthesis in the form of 3 gBlock gene fragments via overhang PCR. Then, we performed digestion on the finished part and ligated it into the pSB1C3 vector. Subsequently, we transformed the pchR-GFP construct (BBa_K2997008) into E. coli Top 10 using electroporation. After allowing the transformed E. coli Top 10 to grow on LB with Cm (Chloramphenicol) plates, colony PCR and sequencing was carried out to determine successful clones. The colony PCR results indicated that the E. coli has pSB1C3 with pchR-GFP. However, after several attempts to construct pchR-GFP, all the sequencing resulted in mutations.
<<Insert pchR sequencing result2.png below the above paragraphs>>
Fig. 15. Sequencing results of the pSB1C3 w/ pchR-GFP construct.
These results led us to hypothesize that the pchR protein is toxic to our E. coli. Moreover, according to a research paper [9] we have found, it is suggested in the study that p-Cresol is toxic to Gram-negative bacteria, which includes E. coli. Hence, this plasmid construct has been dropped.
Therefore, the construction of pchR-GFP was carried out again by re-doing overlap PCR, where we combine all the three IDT gene fragments into the full length pchR-GFP construct. For this time, the finished full length pchR-GFP part was then inserted into the vector pBBR1 MCS-4 (carries ampicillin resistance gene).
<<Insert 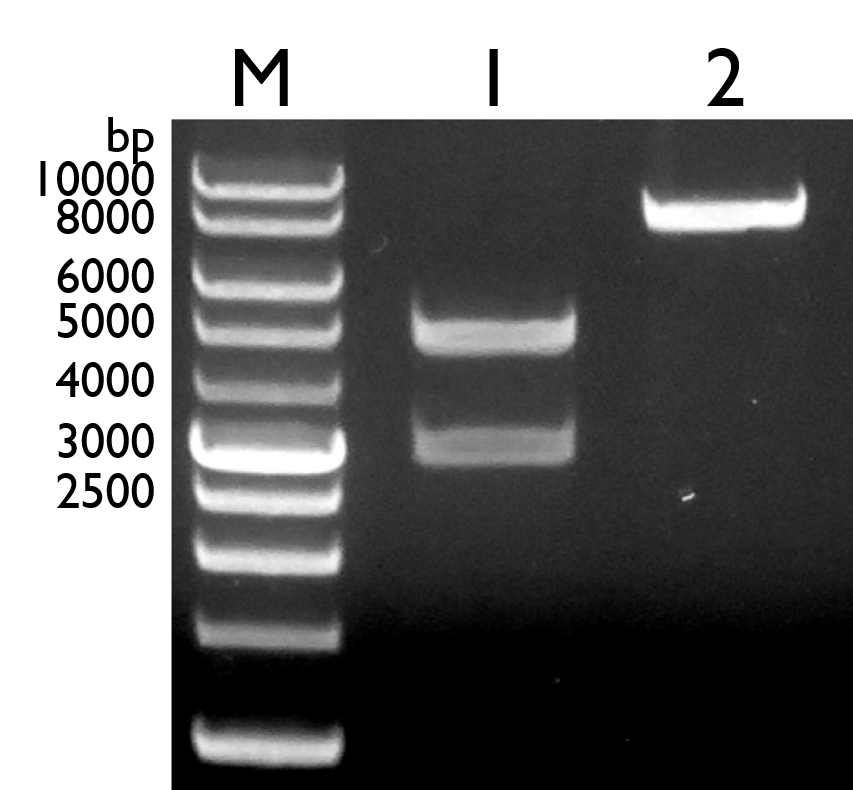
below the above paragraphs>>
Fig. 16. Gel electrophoresis result of pBBR1 MCS-4 with full length pchR-GFP. Lane 1: Marker; Lane 2: Pre-ligation of vector pBBR1 MCS-4 (4950bp) and insert pchR-GFP (3138bp); Lane 3: Post-ligation of pBBR1 MCS-4 with full pchR-GFP (8098 bp)
After that, we proceeded into transforming the construct into Pseudomonas fluorescens 55 and Pseudomonas aeruginosa PAD1 via electroporation.
However, due to the complications surrounding the selection marker being used (Ampicillin), we were not able to get a definitive result that indicates that we have positive colonies. This is due to the fact that P.fluorescens 55 and P.aeruginosa PAD1 are naturally resistant to ampicillin and the vector we used for our construct carries ampicillin resistance gene. Hence, positive colonies cannot be differentiated with negative colonies. Besides that, our colony PCR results on the single colonies were also proven to be negative.
Therefore, with time being one of the factors, we decided to drop any further construction and functional experiments surrounding the p-Cresol sensing part (pchR-GFP).
Sequence and Features
- 10INCOMPATIBLE WITH RFC[10]Illegal prefix found in sequence at 1
Illegal suffix found in sequence at 3173 - 12INCOMPATIBLE WITH RFC[12]Illegal EcoRI site found at 1
Illegal NheI site found at 1769
Illegal SpeI site found at 3174
Illegal PstI site found at 3188
Illegal NotI site found at 7
Illegal NotI site found at 3181 - 21INCOMPATIBLE WITH RFC[21]Illegal EcoRI site found at 1
Illegal BamHI site found at 1182 - 23INCOMPATIBLE WITH RFC[23]Illegal prefix found in sequence at 1
Illegal suffix found in sequence at 3174 - 25INCOMPATIBLE WITH RFC[25]Illegal prefix found in sequence at 1
Illegal XbaI site found at 16
Illegal SpeI site found at 3174
Illegal PstI site found at 3188
Illegal NgoMIV site found at 1227
Illegal AgeI site found at 210
Illegal AgeI site found at 1110 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 1136
Illegal BsaI.rc site found at 3096
| None |
