Part:BBa_K2559003
Ecoli promoter of sgRNA(PrplJ)
The BBa_K2559003 , PrplJ promoter is a strong endogenous promoter in Escherichia coli K 12. We used it to express single guide RNA in our CRISPR/cas9 system.
Usage and Biology
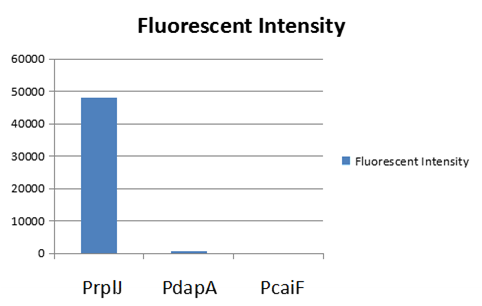
We obtained this promoter from a database named PromEC ( http://margalit.huji.ac.il/promec/index.html) .PromEC is an updated compilation of E. coli mRNA promoter sequences. It includes detailed information of genesites which have been experimentally identified mRNA transcriptional starting position in the E. coli chromosome, as well as the actual sequences of the promoters. We have built up a model to measure the expression intensity of these promoters. Eventually, we selected three promoters including: PrplJ, Pdapa and PcaiF based on the results of modeling PrplJ, Pdapa and PcaiF.We further made chimeric fluorescent fusion proteins between the promoter and GFP respectively and tested the fluorescent intensity driven by different promoters. The combination of PromEC and our modeling can be used as a efficient tool to figure out the information about interested promoters. The GFP expression in E.coli (DH5 α) and was driven by PrplJ, Pdapa and PcaiF promoter.
Basing on the model predition and experiment testing results,we used PrplJ as the promoter of sgRNA
The part is facilitate the in-depth research for other teams!
GZHS-United 2019
At NRP-UEA, we aimed to produce a prebiotic to prevent colon cancer. To do this, we attempted to acylate/butrylate starch in plants. Methods to chemically acylate starch purified from plants already exist, but as they use strong chemicals and require heating they are not environmentally friendly. Using various acyltransferases, we hope to acylate starch in plants. We’ll be using a model plant, Nicotinia benthamiana, for initial tests because we can get results within a few days. We used Golden Gate Cloning and the Plant Standard Syntax described in RFC106 and Patron et al (2015) to make our constructs.
All of our constructs contained the 35s Constitutive Promoter. This was done to drive the constitutive expression of our parts in the plants. We also added a chloroplast transit peptide, acyl transferase, and a 35s terminator to our constructs (BBa_K1618033‐036). In a second set of parts, we added a fluorescent tag, to confirm the function of our chloroplast transit peptide (BBa_K1618029‐032).
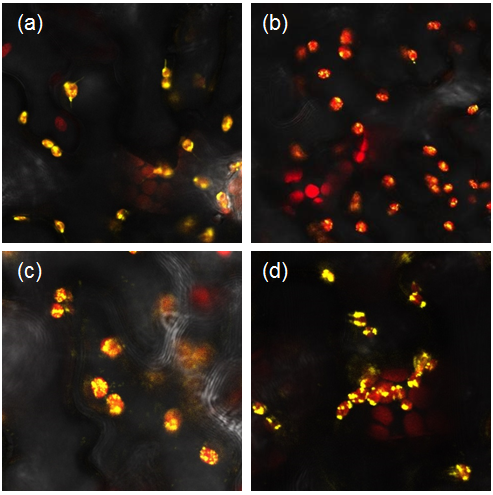
We transformed our constructs into Agrobacterium tumefaciens and then infiltrated it into our plants. We used the YFP tagged constructs to confirm the localisation of parts to the chloroplast (Figure 1) and the untagged constructs to analyse the starch content of our leaves.
Figure 1: Constructs BBa_K1618029‐032 contain a yellow fluorescent protein, as well as a chloroplast transit peptide. These are confocal microscopy images of the constructs infiltrated into Nicotiana benthamiana, in which the red structures are the chlorophyll within the chloroplast, and the yellow is the fluorescent fusion protein expressed from constructs (a) BBa_K1618029, (b) BBa_K1618031, (c) BBa_K1618032, and (d) BBa_K1618030.
The above results not only indicate that our chloroplast transit peptide works, but also suggest that the 35s promoter that we used is GoldenGate compatible and functioning.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
| None |