Part:BBa_K2601004
HOTag3 (Homo-Oligomeric Tag3)
Introduction
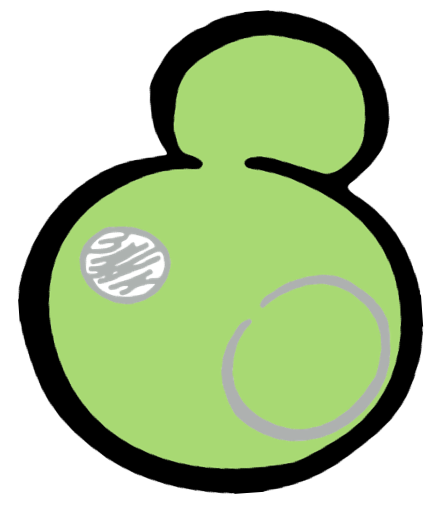
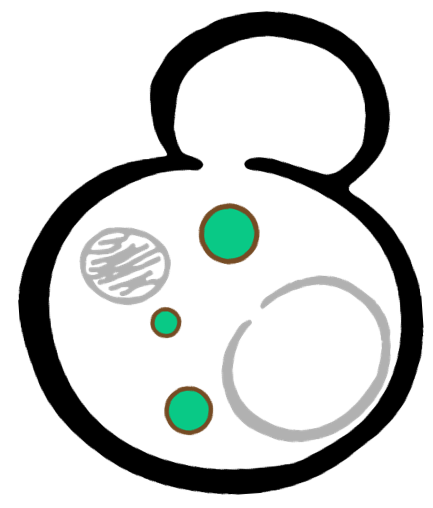
Some membrane-less organelles, such as stress granules and P bodies, have been discovered in recent years. Proteins condense into droplets and assemble these organelles through a process called phase separation. Physically, phase separation is the transformation of a one-phase thermodynamic system to a multi phase system, much like how oil and water demix from each other. According to thermodynamics, molecules will diffuse down the gradient of chemical potential instead of concentration. This is exactly why proteins will self organize into granules, diffusing from regions of low concentration to regions of high concentration. Here is an illustration of phase separation in cells.
 |  |  |
Design
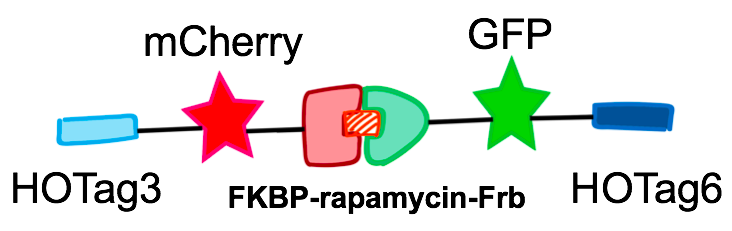
When we wanted to rationally design a synthetic organelle based on phase separation and used it as a platform to achieve multi-functions, some design principles had to be followed. Interaction can bind the parts together while multivalence can make larger assemblies. In order to drive protein phase separation, we needed a multivalent module and a protein-protein interaction module. HOTag is the biobrick that we used to introduce multivalence. In natural process, such as phase separation occurred during T cell signal transduction, multivalency depends on multiple repeats protein domains. But it was not ideal to use multiple repeat domains in our design, because it would not only make the scaffold extremely large but also be problematic for molecular cloning and making transgenic yeasts. Thus, instead of using multiple repeats, we turned to de novo-designed homo-oligomeric short peptides. These short peptides are called HO-Tag (homo-oligomeric tag). HOTags contain approximately 30 amino acids. HOTag3 has high stoichiometry, forming hexamer spontaneously.
As for the protein-protein interaction part, we chose two sets of dimerization modules. The first pair was SUMO and SIM, which can dimerize spontaneously. The second one was chemically inducible FKBP and Frb. Rapamycin was the inducer of dimerization. We chose different modules according to the different functions we wanted to achieve. The hexameric HOTag3, together with another tetrameric HOTag (HOTag6), could robustly drive protein phase separation upon protein interaction (achieved by the protein-protein interaction module). Thus, HOTag3/6 pair is a useful tool to investigate protein phase separation and design a synthetic organelle. To verify the feasibility of the system, we fused two fluorescence proteins with the two components of synthetic organelles. We could observe the self-organization of components and the formation of organelles under fluorescence microscope. We named our system SPOT (Synthetic Phase separation-based Organelle Platform) because it could form granules (fluorescent spots) in yeast. Here is a demonstration of our overall design.
Properties
SUMO-HOTag3 / SIM-HOTag6
Our results confirmed that SUMO/SIM module fused with HOTags can drive phase separation in yeast.
Furthermore, we used Tet07, an inducible promoter, to control the expression of the SIM component. Doxycycline is the inducer of the promoter. SIM fused with HOTag6 couldn’t express in the absence of doxycycline. Before we added doxycycline, SUMO-HOtag3 was evenly distributed in the cells and can’t phase separate. But after we added it, phase separation gradually appears. Therefore, we confirmed that the dimerization modules was essential for phase separation and HOTags alone couldn’t induce the process.
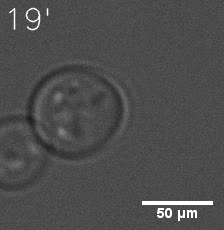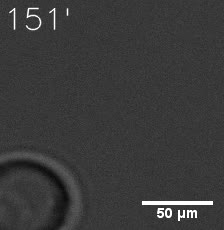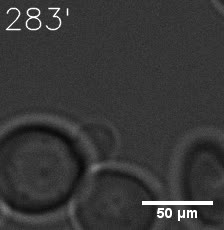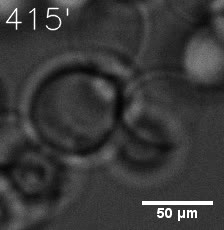 | 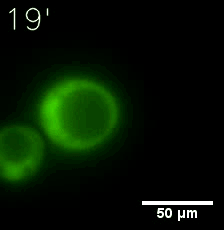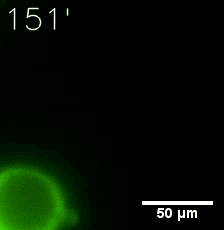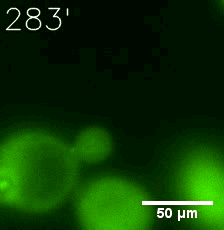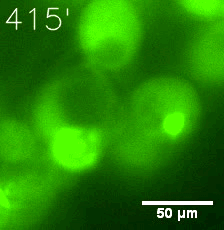 |
 | 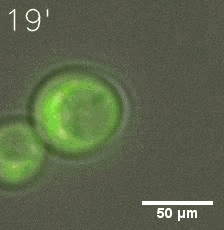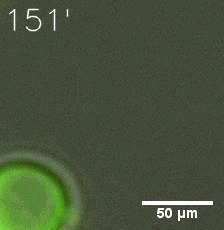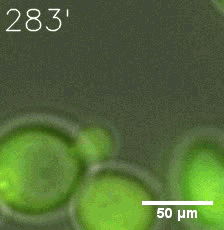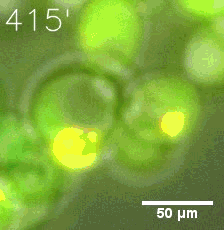 |
| Figure 6. GIF images of different channels. | |
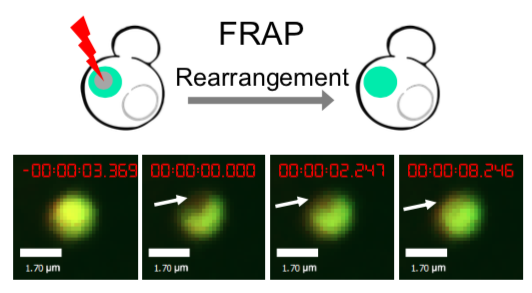
As for the biophysical properties of the system, we assumed the granules were liquid-like. A liquid-like granule is a dynamic system, exchanging mass with cytoplasm rapidly. There are three characteristics which define a liquid-like compartment. First, the compartments should be roughly spherical due to surface tension. It was proved by the 3D-rendered shape of the granules, taken by confocal microscope. Second, two droplets should fuse and coalesce into one droplet spontaneously. Third, the components should undergo rapid internal rearrangement. We used FRAP to verify this criteria. After photobleaching, the fluorescence of granules quickly recovered, which indicated that the granules had rapid mass exchange with cytoplasm.
 | 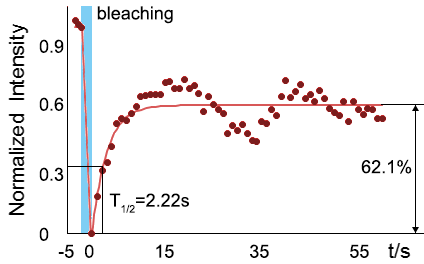 |
| Figure 10. The quantitation of FRAP was determined. The intensity of the fluorescence recovered to approximately 60 percent of the original state. | |
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
//chassis/eukaryote/yeast
//proteindomain/binding
| biology | Saccharomyces cerevisiae |