Part:BBa_K2621000
CAT-Seq Esterase
CAT-Seq Esterase is a hydrolase that was used in Catalytic Activity Sequencing system for its promiscuous capability to hydrolyse N4-acyl-2'-deoxycytidine triphosphates (Substrate Nucleotides). It has been found that the enzyme accepts various N4-acyl-2'-deoxycytidine triphosphates as substrates for hydrolysis, leaving a 2'-deoxycytidine triphosphate (Product Nucleotide).
Sequence-wise it is the most similar to tannases (EC 3.1.1.20) and feruloyl esterases (EC 3.1.1.73), however to date there has been no enzyme characterized hydrolysing N4-amidic bond in modified deoxycytidine triphosphates.
It is the main component of Catalytic Activity Sequencing (CAT-Seq) method. CAT-Seq is a method for high-throughput catalytic biomolecule and genetic regulatory part activity-sequence relationship assessment toolkit.
See how this part is used in the CAT-Seq by pressing here!
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 671
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 487
- 1000COMPATIBLE WITH RFC[1000]
Contents
Introduction
Biology
Description of the CAT-Seq esterase
CAT-Seq Esterase is a hydrolase that has a promiscuous capability to hydrolyse N4-acyl-2'-deoxycytidine triphosphates (Substrate Nucleotide). Sequence-wise it is the most similar to tannases (EC 3.1.1.20) and feruloyl esterases (EC 3.1.1.73), however to date there has been no enzyme characterized hydrolysing N4-amidic bond in modified deoxycytidine triphosphates.
To further elaborate on the fact, it has been found that the enzyme accepts various N4-acyl-2'-deoxycytidine triphosphates as substrates for hydrolysis, leaving a 2'-deoxycytidine triphosphate (Product Nucleotide).
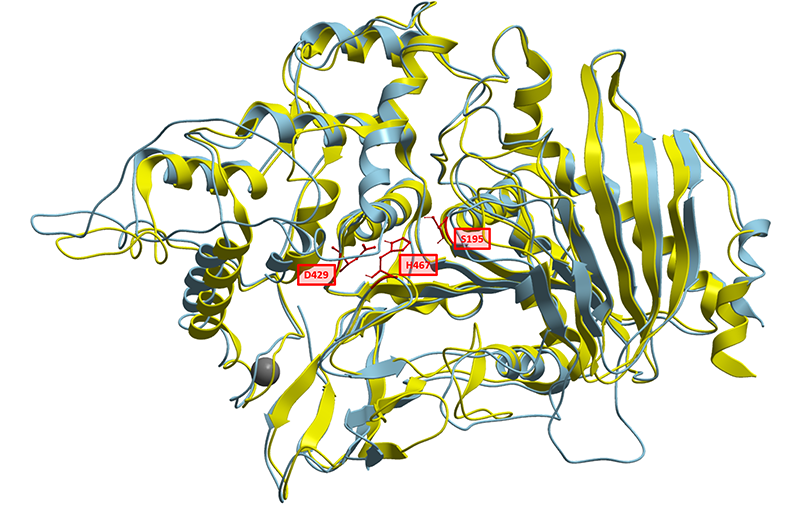
To better understand the nature of this biological part, we have generated a structural homology model based on solved structures of homologous feruloyl esterases. Currently known feruloyl esterases contain a catalytic triad in their active sites consisting of Serine, Aspartate and Histidine. Aligning CAT-Seq Esterase to the structural 3WMT template, we could also identify the catalytic triad consisting of Ser195, Asp429, His467. The sequence similarities and the positions Catalytic Triad amino acids are highlighted in the illustration below (Fig. 1).
In-Silico design of the CAT-Seq esterase mutants
The reason of making esterase mutants was to benchmark the accuracy of CAT-Seq system. We have aimed to make Esterase mutants that would have slightly different activities. After deriving the mutations, we have measured their activities using standard methods, and then compared them with CAT-Seq measurements. Therefore, the esterase and its mutants can now be used for accuracy calibration when setting up the CAT-Seq for the first time.
The residues chosen for mutation were scattered around the catalytic triad of the esterase and involved mainly polar and aromatic side chains. In literature, some of these side chains are assumed to form bonds stabilizing the binding of the substrate (Fig. 2).

After carefully selecting the mutations in-silico, we have synthesized and measured the Activity of those esterase mutants in the laboratory. The mutations have successfully altered the activity of the original Esterase. Also, there were no cases of complete inactivation of enzyme catalytic activity from the mutants we managed test. Please head to the results section for further information.
Usage with CAT-Seq (Catalytic Activity Sequencing)
About CAT-Seq
CAT-Seq stands for Catalytic Activity Sequencing - a system designed and built for high-speed, simultaneous characterization of Catalytic and Regulatory biological parts. You can learn more about CAT-Seq [http://2018.igem.org/Team:Vilnius-Lithuania-OG by clicking this link]
Determining the accuracy of CAT-Seq
Trying to build a CAT-Seq pipeline in your own laboratory will require the CAT-Seq esterase in order to troubleshoot the system and assess the measurement accuracy and precision. In other words, the esterase can be used to calibrate the CAT-Seq.
Together with the Esterase, its substrate attached to a nucleotide is required (substrate nucleotide). In the standard case, the Substrate Nucleotide is N4-benzoyl-2'-deoxycytidine triphosphate. If the Esterase catalyzes the removal of the substrate from the nucleotide, it becomes the Product Nucleotide - 2'-deoxycytidine triphosphate.
Genetic Regulatory Part activity and cross-interaction assessment
While Catalytic Activity Sequencing began as a method for catalytic biomolecule activity recording, we have also create a way to adjust CAT-Seq to record activities of regulatory part. In addition to the activities, cross-interactions of different regulatory parts can also be measured.
When assessing the activities and sequences of libraries of catalytic biomolecules in CAT-Seq , the activity is measured and recorded as a function of Product Nucleotide that was produced in each droplet.
Yet, the activity of the catalytic biomolecule is not the only aspect that can influence the amount of Product Nucleotides that are produced. If all of the droplets would contain the same catalytic biomolecule , but each droplet would have a different concentration of that biomolecule, we would in result get different amounts of Product Nucleotides. For example, droplets with large amount of biomolecules may produce a large number of Product Nucleotides and vice-versa. The default and well-characterized Catalytic Biomolecule in CAT-Seq for regulatory part charectation would be the CAT-Seq Esterase.
Yet, the activity of the catalytic biomolecule is not the only aspect that can influence the amount of Product Nucleotides that are produced. If all of the droplets would contain the same catalytic biomolecule , but each droplet would have a different concentration of that biomolecule, we would in result get different amounts of Product Nucleotides. For example, droplets with large amount of biomolecules may produce a large number of Product Nucleotides and vice-versa. The default and well-characterized Catalytic Biomolecule in CAT-Seq for regulatory part charectation would be the CAT-Seq Esterase.
Example for RBS activity strength determination
We can now explore an example in which the CAT-Seq esterase can be used to assess the strengths of different ribosome binding site (RBS) sequences.
While keeping all the workflow from the main Catalytic Activity Sequencing design, all we need to do is to change the way we prepare the library. Firstly, instead of using a library of catalytic biomolecules, we can use a single CAT-Seq Esterase enzyme. Then, specific or random RBS sequence library can be prepared and attached to the esterase enzyme sequence. After that, every step of CAT-Seq method is the same.
After the library is encapsulated into droplets, depending on the RBS sequence strength some droplets may have different amounts of the esterase. For example, the droplets with more esterase will produce more Product Nucleotides and vice-versa. After the amplification and sequencing steps we can learn two main things about the measured library - the RBS sequence and its relative RBS strength in the form of Product Nucleotide concentration for every library variant screened.
Example for Toehold cross-interaction determination
Next, we want to give an example on how to record regulatory part interactions using CAT-Seq. In this case, we will be using Toehold Switches.
The Toehold Switch systems are composed of two RNA strands referred to as the Switch and Trigger. The Switch RNA contains the coding sequence of the gene being regulated. The Switch RNA forms a hairpin structure that includes the RBS site. While the hairpin structure is formed, the translation is inhibited. The Trigger RNA is a molecule that can selectively bind to the Switch RNA region and expose the gene RBS site for ribosomes. Once that happens, the protein translation can be initiated.
Once again, only the first part of general CAT-Seq design needs to be changed - the library preparation. Instead of using a catalytic biomolecule library, a single enzyme is used. Then, libraries need to be prepared - one for Toehold Switches and another for Trigger RNA. The Trigger RNA libraries also require a separate T7 promoter for RNA expression. Then, both of those libraries must be ligated to the enzyme DNA fragment. Resulting library contains fragments which have the same catalytic biomolecule (In the general case - the CAT-Seq Esterase), yet each of them have a random combination of a specific Toehold Switch and RNA Trigger.
After the encapsulation, the droplets which have the correct Trigger RNA and Toehold Switch combination (in which Trigger RNA binds to that specific Toehold Switch), can produce catalytic biomolecules . In turn, those catalytic biomolecules can produce the Product Nucleotides.
In other droplets, where the Trigger RNA binds to the Toehold Switch with weaker affinity, there are also less catalytic biomolecules and Product Nucleotides produced. Finally, the droplets in which the Trigger RNA does not bind to the Toehold Switch, there are no biomolecules or product nucleotides produced.
Characterization of the CAT-Seq Esterase (Vilnius-Lithuania Overgraduate 2018)
Kinetic characterization of the CAT-Seq Esterase
Spectrofotometic kinetic data based on decay of absorbance at 310 nm due to substrate nucleotide catalytic conversion was gathered using a range of starting substrate nucleotide concentrations. A Michelis Menten curve for the CAT-Seq esterase was plotted using the aquired data. Data shown in 1 Fig., show perfect fit (R2 = 0.9449) to a standard Michaelis Menten curve.
Michaelis Menten plot transformations were generated for CAT-Seq esterase enzyme (BBa_K2621000). 2 Fig., display Lineweaver-Burk and Hanes-Woolf transformation plot. Based on the equation koeficients in the Hanes-Woolf transformation, the expermentally determined Vmax value is 17.2 µM/min, Km value is 86 µM.
Activity measurements of CAT-Seq Esterase and Esterase mutants
The 10 esterase mutants housing mutations at bioinformatically predicted sites were created. To see why and how mutations were designed in-silico, [http://2018.igem.org/Team:Vilnius-Lithuania-OG/Model please click this link]. Each of the mutant was constructed utilizing PCR and synthesized using In vitro transcription and translation kit and their catalytic activity towards N4-benzoyl-2'-deoxycytidine triphosphate were tested.
The reaction kinetics were measured using the spectrophotometer as a decrease of absorbance. 3 Fig. displays the relative hydrolysis speed of each mutant generated. As seen from these results, a variety of mutants, showing different catalysis speeds were produced. Some of the amino acids changes affected the activity drastically, for example Trp224 to Tyr, Lys227 to Arg or Glu509 to Lys. Other mutations only modulated the activity Asn107 to Asp or Glu194 to Ala. Additionally, large 8 amino acid deletion at position Pro348-Hy356 caused only a moderate decrease in enzymes activity.
Ribosome binding site activity measurements
The three ribose binding sites (BBa_B0030, BBa_B0032, BBa_B0034) already found in biobrick registry and a new CAT-Seq RBS BBa_K2621038 were place upstream the CAT-Seq esterase gene. The strength of each ribosome binding site was investigated by expressing the esterase enzyme using the In vitro transcription and translation kit and spectrophotometrically measuring the as a decrease of absorbance due to the hydrolysis of substrate nucleotides.
<grafikas>
X Fig. displays the relative hydrolysis speed (normalized to BBa_B0034) of CAT-Seq esterase expressed using 4 different ribosome binding sites. Based on the fact that the same enzyme and substrate nucleotide concentration are used, the difference in activity corresponds to difference in synthesised biomolecule concentrations. The expression of an enzyme depends on the particular strength of used ribosome binding sites. Taking this into consideration, the measured relative activity corresponds to strength of ribosome binding sites. The results fit well with already determined relative ribosome binding strength given in biobrick registry - Bba_B0034 - 1; BBa_B0032 - 0.48; Bba_B0032 - 0.045.
Cross-interaction measurements of Toehold Switches
The 9 library members, composed of 3 unique Toehold Switch sequences and 3 unique activating RNA sequences, termed Trigger RNA, were designed to test the orthogonality and regulatory characteristics of each part.
Each of the regulatory part, consisting of one toehold sequence upstream of the CAT-Seq esterase gene (BBa_K2621000) and one trigger sequence were constructed. First of all, the orthogonality of each toehold:activating RNA pair and they regulatory characteristic have been tested in bulk.
The constructed library members were synthesized using In vitro transcription and translation kit and their catalytic activity towards N4-benzoyl-2'-deoxycytidine triphosphate were tested. The reaction kinetics were measured using the spectrophotometer as a decrease of absorbance due tue hydrolyzed substrate nucleotide.
<grafikas>
4 Fig. displays the relative hydrolysis speed of each regulatory part variant in a form of matrix. The decrease of absorbance shown in Y graph corresponds to the catalytic conversion of substrate nucleotides. As seen from control experiment, in which standard esterase was expressed, the decline of absorbance over time is seen.
Taking these results into consideration, the same decrease of absorbance is only seen in the diagonal of the matrix. This means, that active catalytic molecules is expressed only when both regulatory molecules of the same group are present - Toehold1 with Trigger1, Toehold2 with Trigger2 and Toehold3 with Trigger3. None of the regulatory sequences show any cross talk with the other group. These results conclude the generation of working toehold riboswitches control library for regulatory sequences parameter and orthogonality screening.
References
| None |