Part:BBa_K2728001
pfrmR - An Engineered Formaldehyde-Inducible Promoter
Basic Description
This promoter is an engineered formaldehyde-inducible promoter. Escherichia coli has a native formaldehyde-inducible promoter, pfrm, which is found upstream of the frmRAB formaldehyde detoxification operon. FrmR, the first product of the operon, is a member of the DUF156 family of DNA-binding transcriptional regulators. It binds the frmRAB promoter region and is negatively allosterically modulated by formaldehyde. FrmR is specific to formaldehyde, responding to acetaldehyde, methylglyoxal, and glyoxal to far lesser degrees and not at all to a range of other aldehydes and alcohols tested. The genes frmA and frmB encode a formaldehyde dehydrogenase and S-formylglutathione hydrolase, respectively, and are responsible for detoxifying formaldehyde to formic acid in a glutathione-dependent pathway. The negative-feedback regulation of the frmRAB operon is similar to that of many other prokaryotic operons, whereby the transcription factor represses its own transcription.
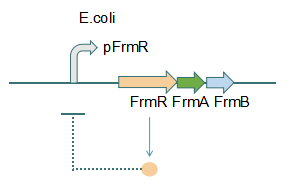
Fig 1: Without Formaldehyde
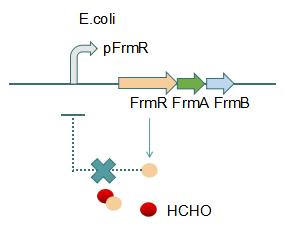
Fig 2: With Formaldehyde
Features
- It’s a formaldehyde-inducible promoter from E.coli.
- It’s an engineered promoter. It retains formaldehyde responsiveness, with 2-fold higher GFP expression in response to 100 μM formaldehyde than the native pfrm. Application of this promoter with higher basal and induced expression levels before methanol assimilation genes achieves higher biomass titers than the native E. coli pfrm.
Origins
Escherichia coli
Improvements and Experimental Characterization
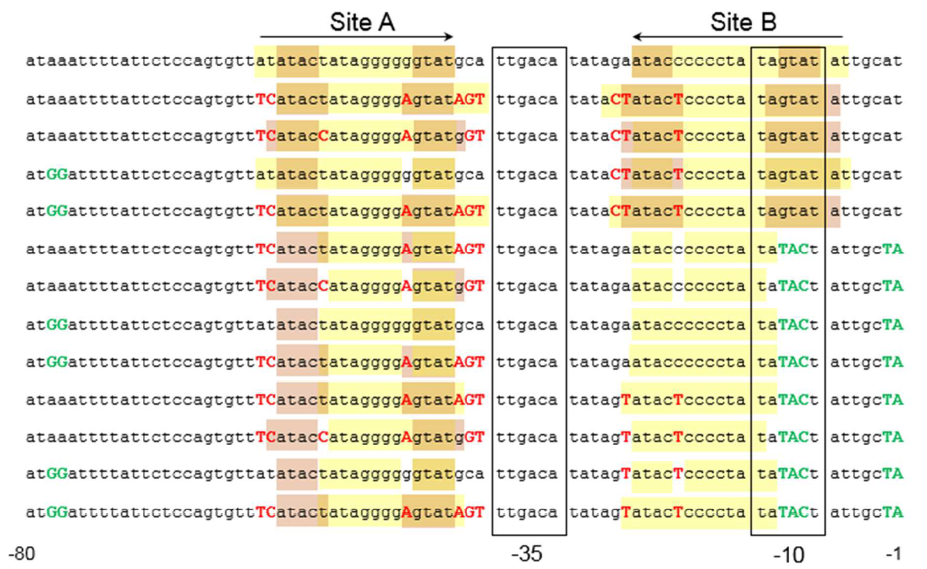
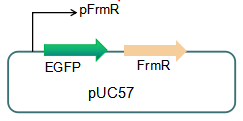
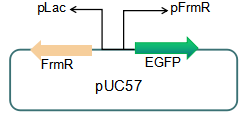
The sequence of this part was taken from the research of Rohlhill J. etc. The 2 binding sites of variations are -35 and -10 (Fig 1). We ordered synthesized plasmid with pFrmR and EGFP from Gensceipt and constructed pFrmR-EGFP-FrmR reporter system on plasmid pUC57. After dosing formaldehyde of 100 to 400uM, we tested the EGFP expression to identify the activity of the formaldehyde induced response of this prompter.
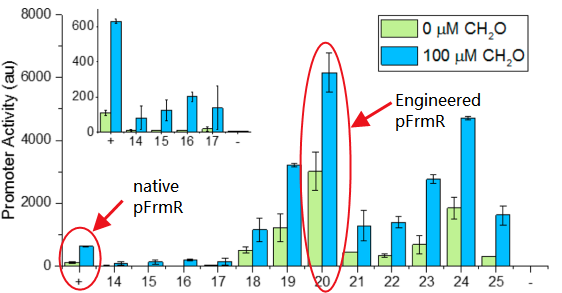
The research conducted by Rohlhill J. etc. has proved that, pFrmR retains formaldehyde responsiveness, with 2-fold higher GFP expression in response to 100 μM formaldehyde than the native pfrm (BBa_K749008 )[4].
Compared with the research of Rohlhill J. etc. (Fig 2), unfortunately we were not able to detect any fluorescence after several rounds of repeating experiments (Fig 4). This result did not necessarily indicate that there was no activity of this promoter. One of the probable reason to cause this, might be that the concentration of formaldehyde that entered the cells was not strong enough, thus the expression level of FrmR and the level of formaldehyde was unbalanced. Most of the FrmR binded with pFrmR, therefore repressed the activity of pFrmR, which means, pFrmR was at a inhibited status.
Future Improvements:
We plan to optimize our reporter vector by introducing an independent promoter pLac to regulate the expression of FrmR.
Fig 1: Sites of mutants
Fig 2: Comparison of activity
Fig 3: Current reporter system
Fig 4: Fluorescent strength vs Formaldehyde concentration
Fig 5: Future reporter system
Potential Application
- To construct a formaldehyde sensor with this promoter.
- To enable higher growth under formaldehyde pressure with the application of the engineered formaldehyde responsive promoter.
Parts Verification Before Submission
We verified our parts in the lab before submission. They are reliable! Please feel free to apply them onto your project.=)
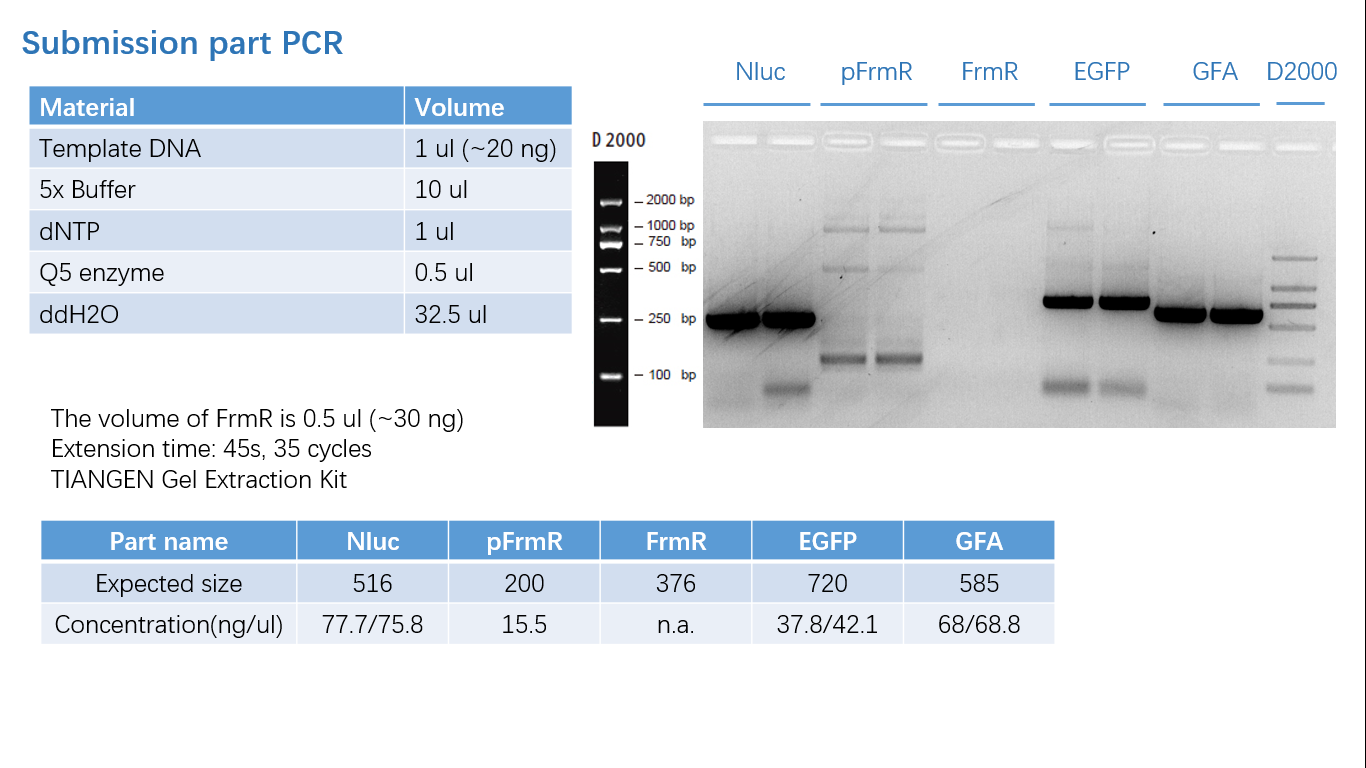
Fig 1: PCR (to get targeted genes)
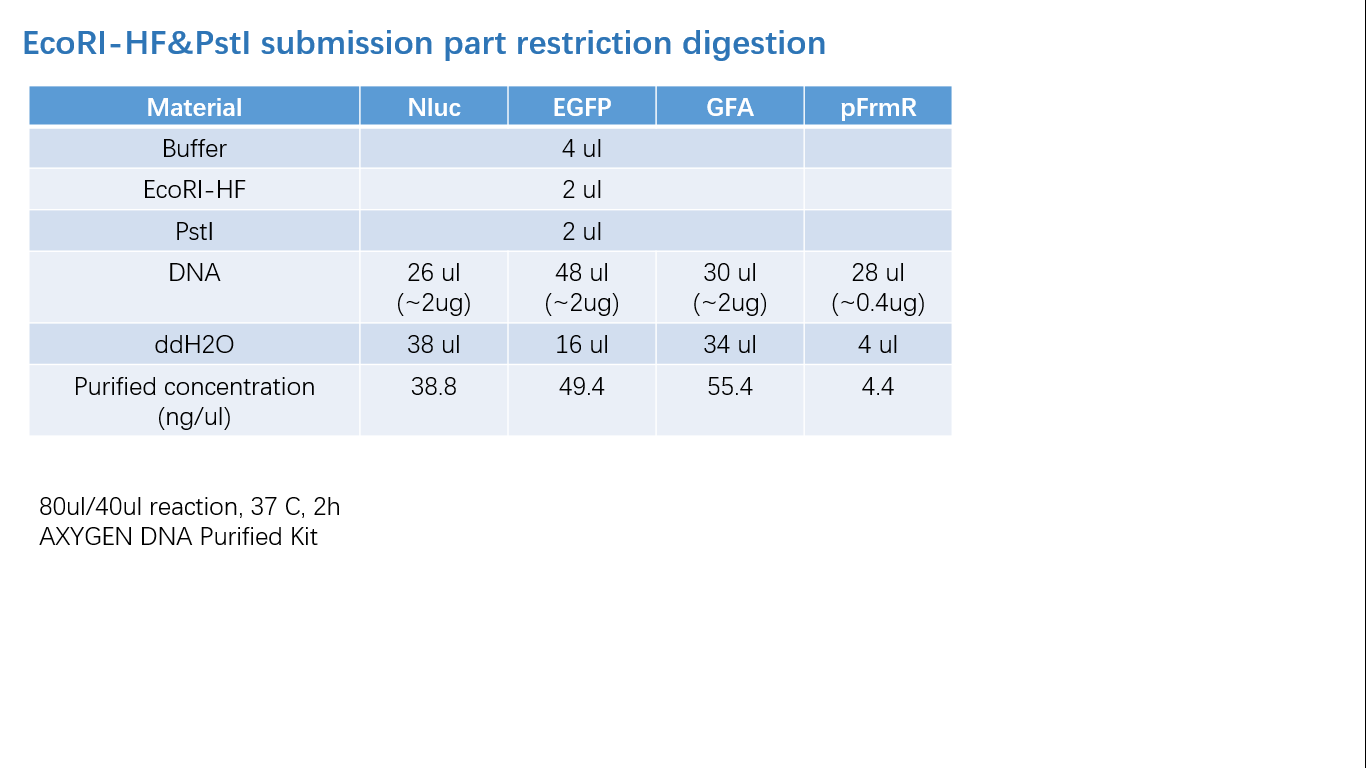
Fig 2: Restriction Digestion
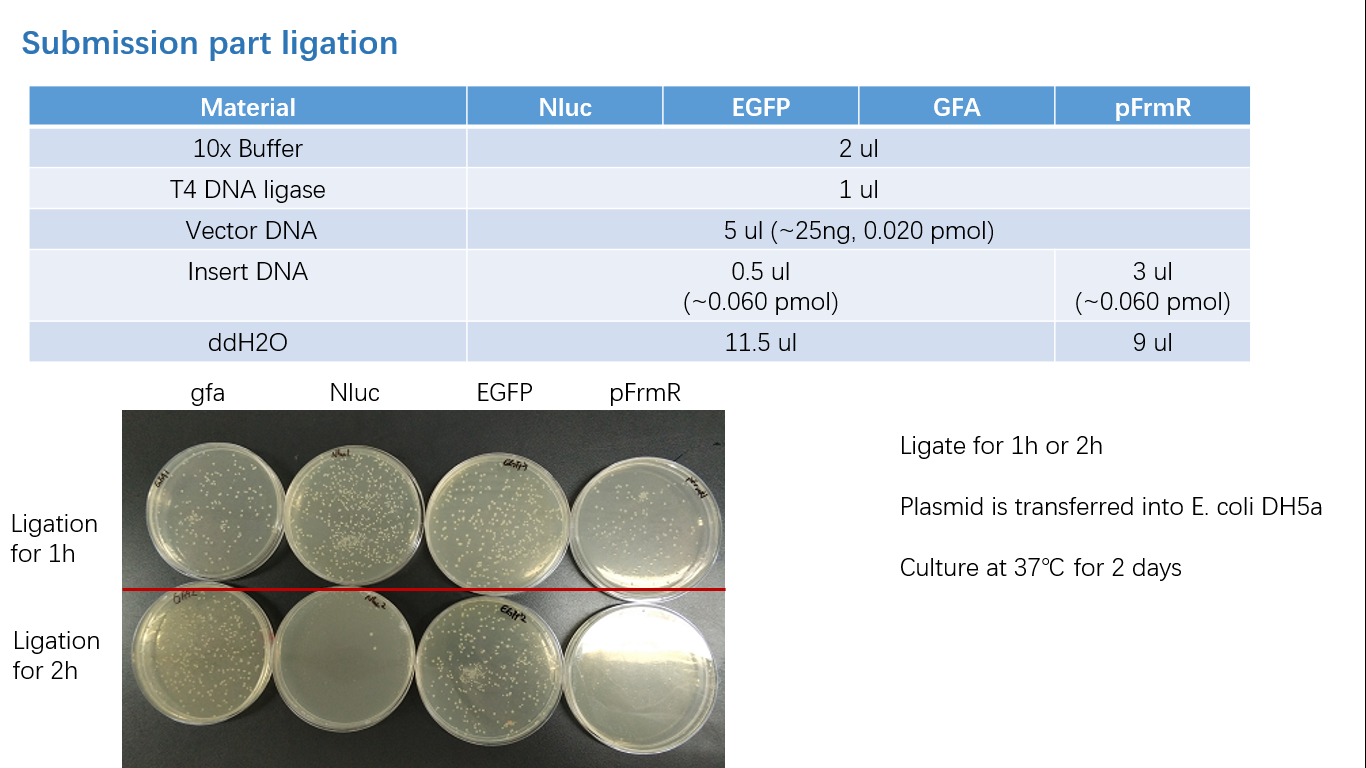
Fig 3: Ligation
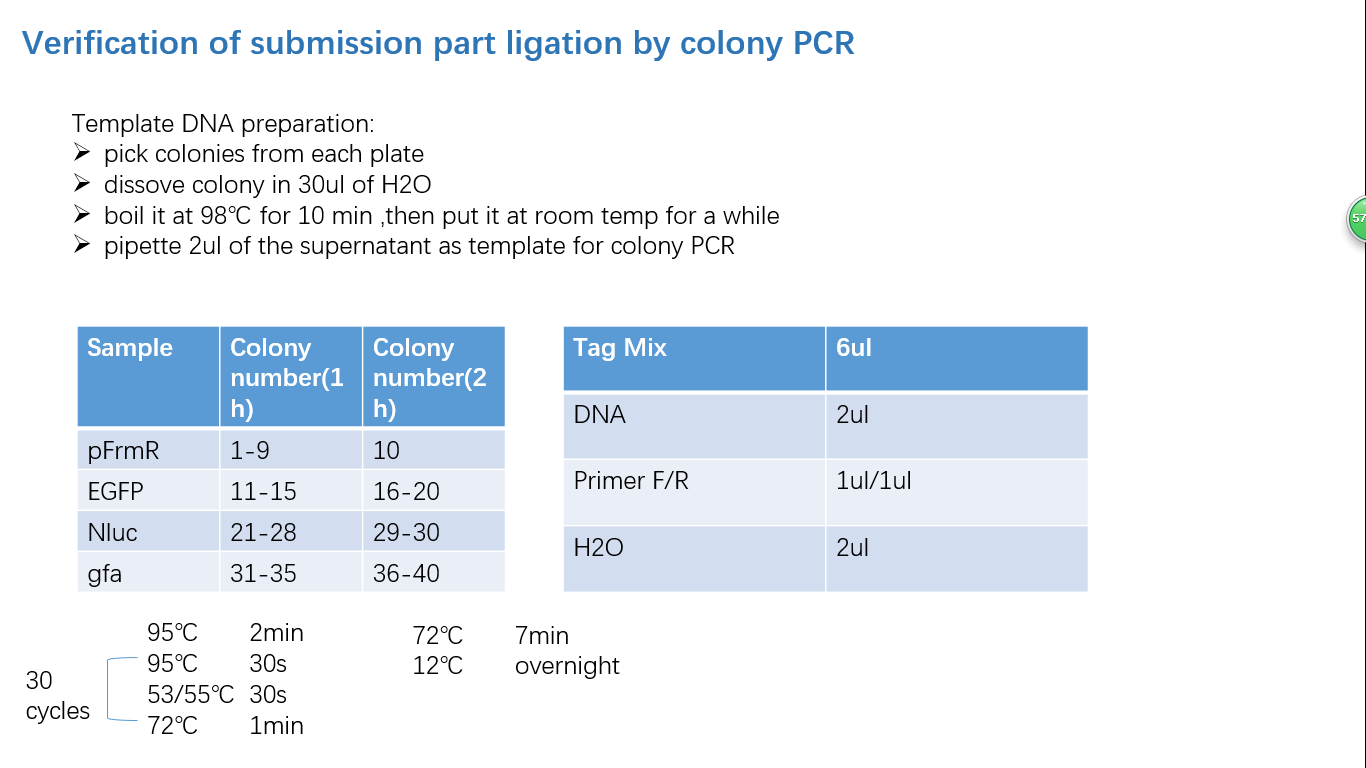
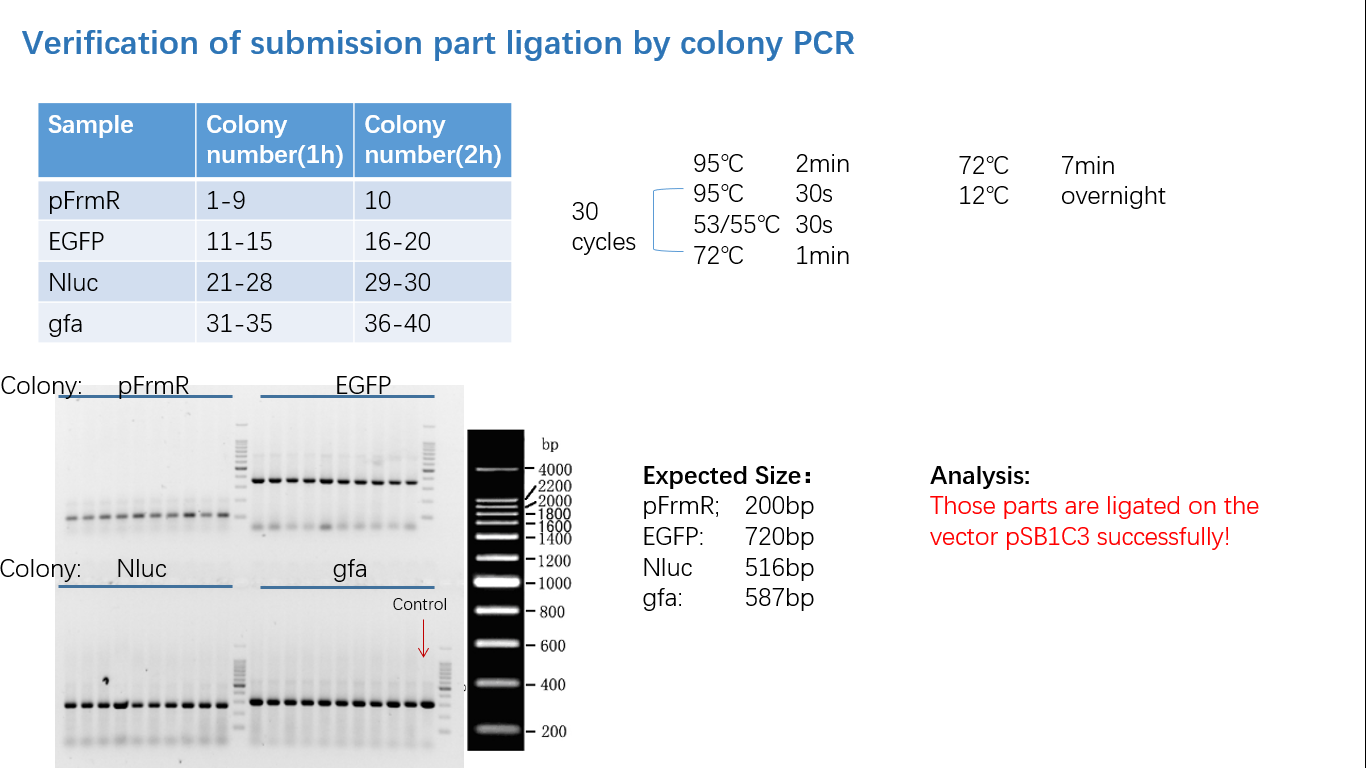
Fig 4: Colony PCR
Fig 5: Gel Verification
References
- Osman, D., Piergentili, C., Chen, J., Sayer, L. N., Uson, I., Huggins, T. G., Robinson, N. J., and Pohl, E. (2016) The Effectors and Sensory Sites of Formaldehyde-Responsive Regulator FrmR and Metal-Sensing Variant. J. Biol. Chem. 291, 19502-19516
- Denby, K. J., Iwig, J., Bisson, C., Westwood, J., Rolfe, M. D., Sedelnikova, S. E., Higgins, K., Maroney, M. J., Baker, P. J., Chivers, P. T., and Green, J. (2016) The mechanism of a formaldehyde-sensing transcriptional regulator. Sci. Rep. 6, 38879
- Gonzalez, C. F., Proudfoot, M., Brown, G., Korniyenko, Y., Mori, H., Savchenko, A. V., and Yakunin, A. F. (2006) Molecular basis of formaldehyde detoxification: Characterization of two S-formylglutathione hydrolases from Escherichia coli, FrmB and YeiG. J. Biol. Chem. 281, 14514-14522
- Rohlhill J, Sandoval N R, Papoutsakis E T. Sort-seq approach to engineering a formaldehyde-inducible promoter for dynamically regulated Escherichia coli growth on methanol.[J]. Acs Synthetic Biology, 2017, 6(8)
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
| None |