Part:BBa_K1907002
Microcystinase (MlrA) for E. coli
Introduction
This part can be used to introduce microcystinase (MlrA) from Sphingomonas sp. ACM-3962 to Escherichia coli. MlrA is an enzyme degrades some cyanobacterial toxins, most notably microcystin-LR (MC-LR). To facilitate expression detection and purification of the enzyme, a 6X Histidine tag is fused on the C-terminus of MlrA, with a short linker in between. The full protein-coding sequence has been codon-optimized for expression in E. coli by GeneArt.This BioBrick is an improvement on the one created by the Peking 2014 team (BBa_K1378001). This new BioBrick has the identical MlrA amino acid sequence as the GenBank entry from Sphingomonas sp. ACM-3962 (accession number AF411068.2), which refers to the article by Bourne et al., 2001. On the other hand, the sequence of the older BioBrick differs by one amino acid, but there are no available MlrA sequences in GenBank that match with this difference. Therefore, it seems likely that the difference is due to an error in cloning, or entering the sequence.
In this new BioBrick, there is also a 6X His-tag connected to the C-terminus by a linker. The incorporation of this tag follows the example of Dziga et al. (2012), who successfully expressed the same MlrA protein with a C-terminal His-tag. This tag will allow for the enzyme to be purified with affinity chromatography and also detected with anti-His antibodies in western blotting. Without the ability to use antibodies and western blot, it can be difficult to detect or quantify the expression of the enzyme.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 105
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 244
Illegal AgeI site found at 373 - 1000COMPATIBLE WITH RFC[1000]
Background
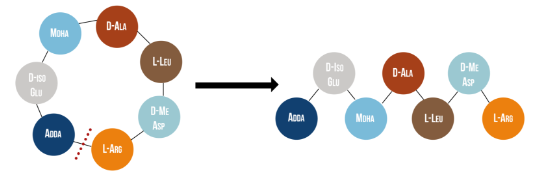
Microcystinase, or MlrA, is a member of a set of enzymes that degrade microcystins. These enzymes - MlrA, MlrB, MlrC and MlrD - are found naturally in some gram-negative bacteria, for example Sphingomonas sp. They each have a different function in the overall process of microcystin degradation. MlrA is first in the sequence to act. It hydrolyzes the peptide bond between the Adda and arginine amino acids in microcystin-LR (MC-LR). The breakage of this bond linearizes the toxin (Figure 1). Following this, MlrB (a serine protease) and MlrC (a metalloprotease) are able to further degrade the linearized product into individual amino acids, which are then used in the cell’s metabolism. Therefore, the toxin is likely used as a carbon source. MlrD’s function is not fully understood at this point, but it may have a role in importing the toxins. (Bourne et al., 2001)
Based on a homologous structure model (Figure 2) obtained from the Phyre2 tool (Kelley et al., 2015), it seems that the enzyme is a transmembrane protein consisting of eight hydrophobic α-helices that form a channel, with the presumed active site (Bourne et al., 2001) residing in the middle. It is worth pointing out, however, that the model does not cover the whole length of the MlrA sequence, as only very few homologs were found for the sequence - the homology model is mostly based on two homologs that didn’t cover the very ends of either terminus. Other than this evidence, there are previous studies pointing to the direction of MlrA being a membrane protein (Pei et al., 2011). However, when the enzyme was heterologously produced in E. coli , it was noticed that it had localised mostly in the soluble fraction of the cytosol, in an active state (Dziga et al., 2012).
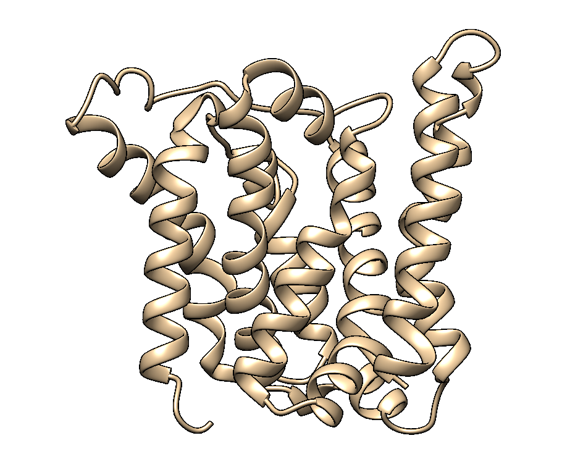
The mechanism of MlrA’s function is not yet fully understood. The homologous structure model gives some suggestions as to how the enzyme might work: it seems that one end of the channel is wider than the other, and that the active site resides in the middle of it. Consequently, if the enzyme is located on the outer membrane of the cells, it might actually be a transporter which the cells use to import the toxin and simultaneously linearize it.
Expression
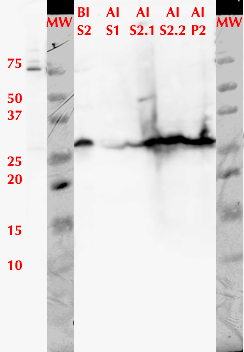
To express the enzyme, a 5 mL overnight-grown LB-kanamycin culture of E. coli BL21(DE3) was expanded to 50 mL. After the bacteria reached the mid-log phase (OD600 between 0.6-0.8), protein production was induced with 1 mM IPTG. During induction, bacteria were grown at +22 °C for 20 hours. Figure 1 presents results from analyzing different cellular fractions over the course of expression and lysis. After cell lysis, the enzyme is observed to localize in both the soluble and insoluble fractions.
Expression of the protein was also done in Magic Media auto-induction medium, in a total volume of 0.5 L in a 2 L flask covered with an air-permeable membrane. This expression was conducted in +30 °C.
FPLC purification
We used FPLC to purify the enzyme we had grown on the Magic Media auto-induction medium (0.5 L culture volume). Half of the supernatant obtained after cell lysis was used for purification. Figure 2 presents the chromatogram from the FPLC and selected fractions were analysed by SDS-PAGE and western blotting (Figure 3).
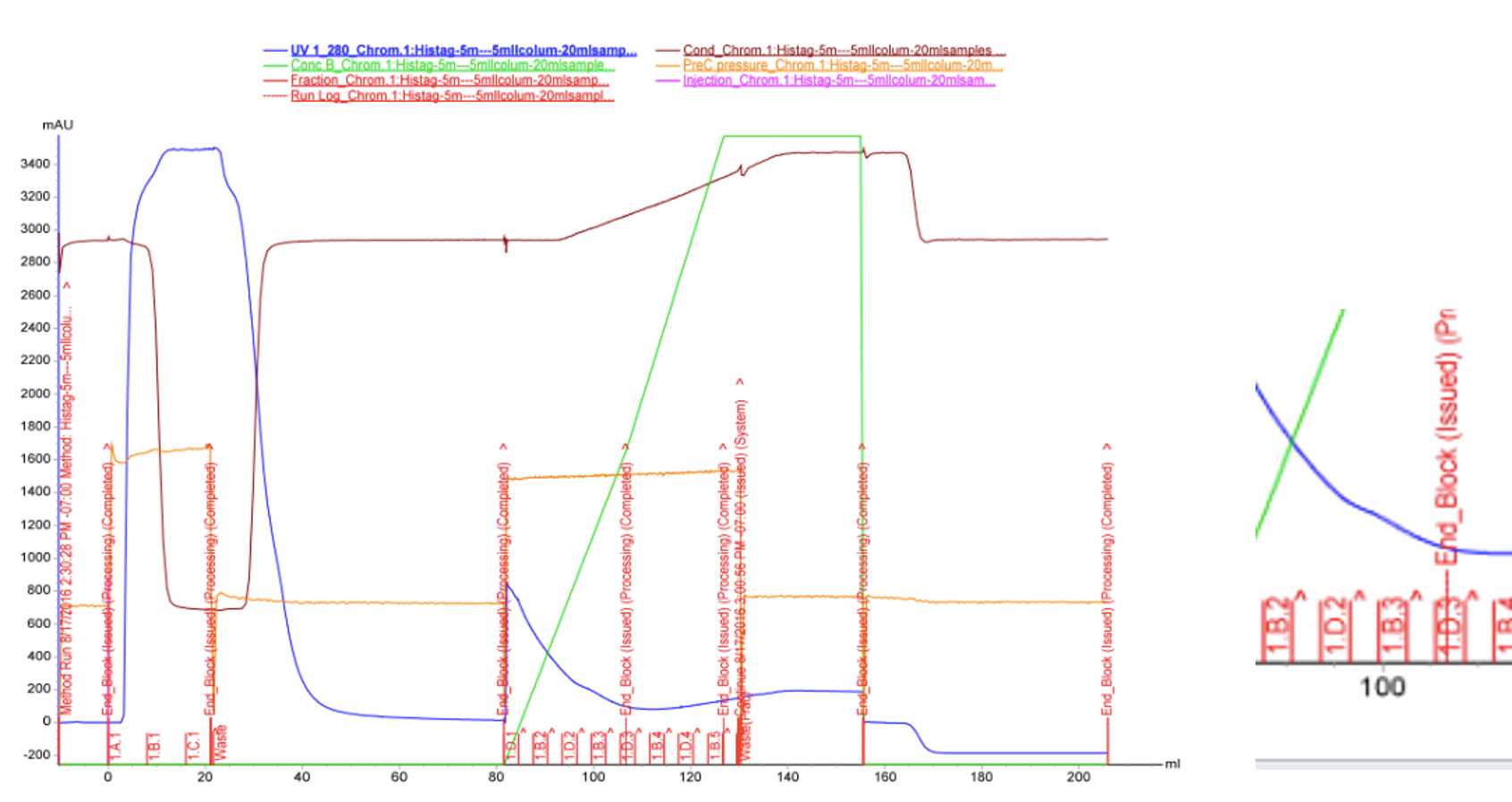
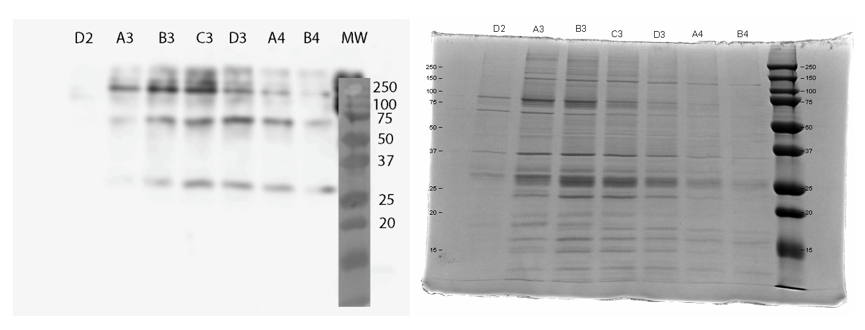
Based on these results, it seems that affinity chromatography purification of MlrA is possible, although optimization of the purification conditions would be necessary to obtain better purity and higher yields of purified protein. We didn’t have time to try these improvements ourselves, but possibilities include a very slow initial elution buffer gradient to wash out unspecific binding. In addition, optimizing the pH of the sample could improve binding; a higher pH might allow for better binding to the nickel in the column, as the number of histidines with a complementary charge would increase. Diluting the sample in phosphate buffer and desalting the sample could improve its binding to the column. Additionally, binding could be facilitated by extending the 6X His tag to a 10X His tag.
Activity measurements
The activity of MlrA was measured by incubating the enzyme with microcystin extract. The final concentration of microcystin was estimated to be 1 μg/L. Samples were taken at different time points and they were analysed with liquid chromatography/mass spectrometry (LC-MS). This gives data of the relative abundance of microcystin-LR and -RR in each sample. The results were then plotted against time, together with a negative control which consisted of microcystin extract and PBS.
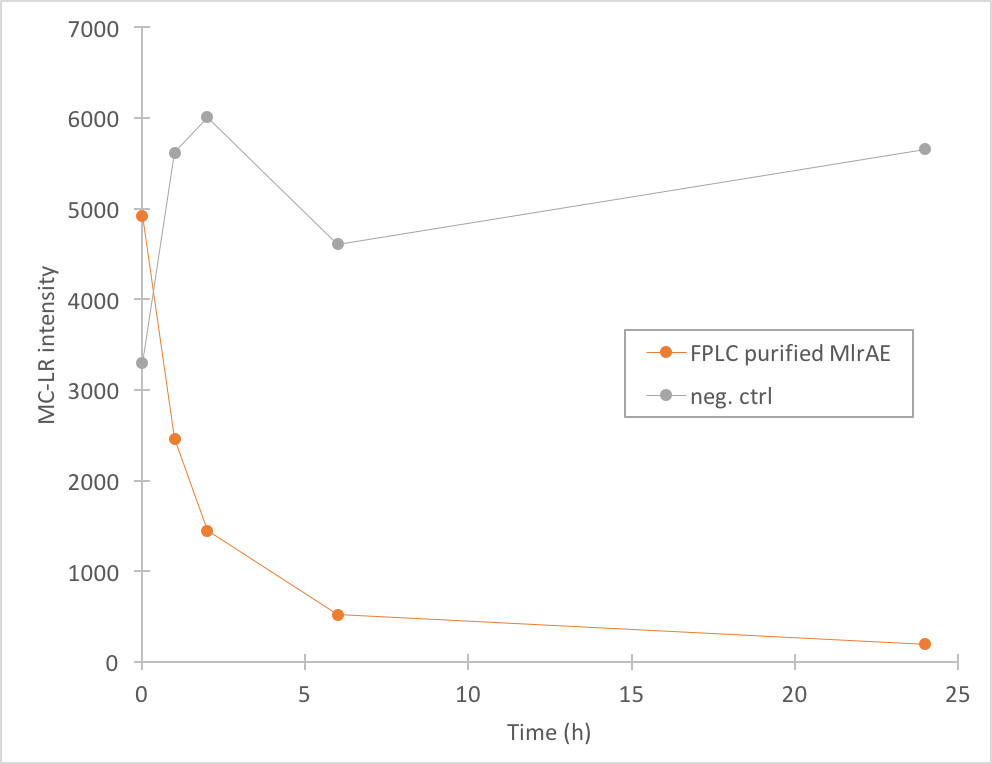
The results we acquired from this construct show that the amount of microcystin-LR decreases, which means that the enzyme is active (Figure 4). We used 970 ul of FPLC purified elution fraction in the MlrA-MC reaction, but as we don’t have exact information on the quantity of the enzyme, we were not able to quantify the activity activity any further.
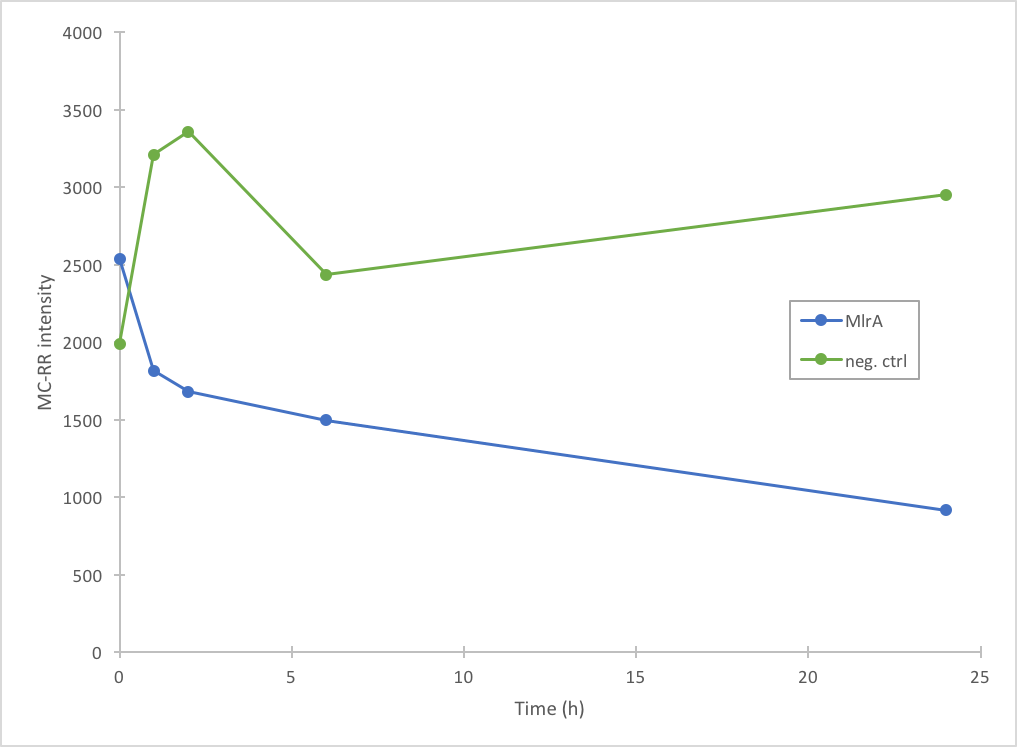
In addition to improving the previous MlrA BioBrick, we also managed to prove that our construct degrades microcystin-RR (Figure 5). Based on our results, it seems that MlrA has higher degradation activity against MC-LR than MC-RR.
For more details on the experiments done with this BioBrick, visit http://2016.igem.org/Team:Aalto-Helsinki/Laboratory
References
Bourne DG, Jones GJ, Blakeley RL, Jones A, Negri AP, Riddles P., 1996. Enzymatic Pathway for the Bacterial Degradation of the Cyanobacterial Cyclic Peptide Toxin Microcystin LR. Appl. Environ. Microbiol. 62(11), pp. 4086-4094.
Bourne DG, Riddles P, Jones GJ, Smith W, Blakely RL., 2001. Characterisation of a gene cluster involved in bacterial degradation of the cyanobacterial toxin microcystin LR. Environ Toxicol. 16(6), pp. 523-34.
Dziga D, Wladyka B, Zielińska G, Meriluoto J, Wasylewski M., 2012. Heterologous expression and characterisation of microcystinase. Toxicon, 59(5), pp. 578-586.
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJE., 2015. The Phyre2 web portal for protein modeling, prediction and analysis. Nature Protocols 10, pp. 845–858.
Pei J, Mitchell DA, Dixon JE, Grishin NV., 2011. Expansion of type II CAAX proteases reveals evolutionary origin of g-secretase subunit APH-1. J. Mol. Biol. 410:18–26.
//chassis/prokaryote/ecoli
| None |