Part:BBa_K628006
Protegrin-1 Kill Switch
This part functions as a protegrin-1 (antimicrobial peptide), pBad strong (arabinose-sensitive inducible promoter) controlled E. coli kill switch.
Usage and Biology
Protegrin-1 coding region under the influence of a pBad-strong promoter. Protegrin-1 is an '18-residue beta-sheet peptide isolated from porcine leukocytes with antimicrobial activity against a broad range of microorganisms.' (Steinberg et al 1997) It has its effect by pore membrane disruption (Lam et al 2006) and possibly also by effects such as activation of membrane-damaging proteases (Bierbaum 1985) and has anti-microbial activity against E. coli (Aumelas 2004).
The presence of the pBad strong promoter allows the expression of protegrin-1 to be induced quickly, but at various rates by differing concentrations of arabinose.
The kill switch functions by producing protegrin-1 intracellularly, and concerns were raised about whether antimicrobial peptides would function integrating into the inner phospholipid bilayer leaf, rather than the outer leaf like they do in nature. Protegrin-1 integrates into membranes containing zwitterionic phospholipids, which promote electrostatic interactions between protegrin-1 and the bilayer. As these are present in both the outer and inner leaflets of prokaryotic bacterial membranes, we believe this antimicrobial effect, and therefore, the functionality of the kill switch, should still be present despite the kill switch activating intracellularly. More information on this issue can be found on our wiki: http://2011.igem.org/Team:St_Andrews/switch
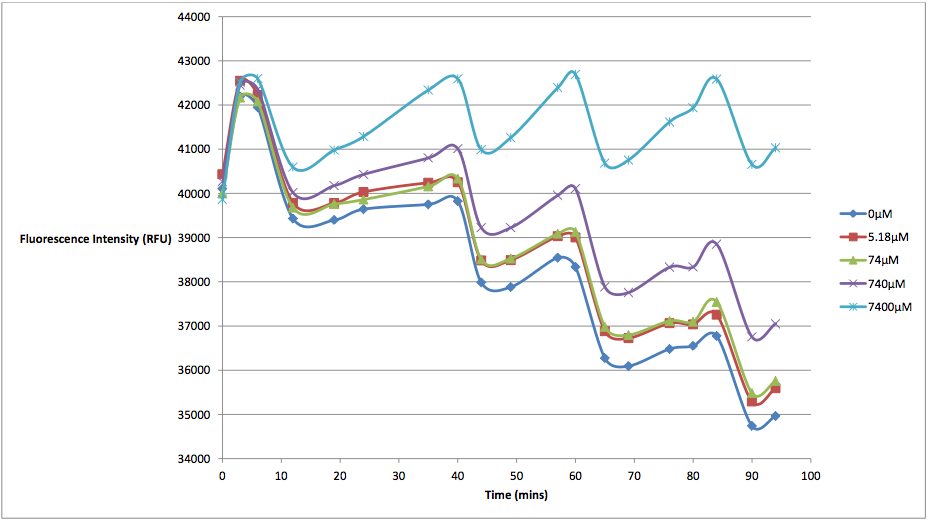
Figure 1 - This graph shows fluorescence readings at 600 nm (485 nm exciting) of K628006-transformed E. coli in solutions of various arabinose concentrations.
The above graph measures the fluorescence readings of E. coli transformed with K628006 across multiple arabinose concentrations. The y-axis represents the scale of the fluorescent output of each E. coli population, while the x-axis denotes the time of each reading. The graph doesn't look very aesthetically pleasing, and in fact, may lead some to believe that our readings of fluorescence decreased over time, and subsequently, so did cell death. However, this is due to the fact that the dye used to stain the cells reduced in fluorescing power over time, resulting in what looked like decreasing cell death. In order to show the relationship between the effects of various arabinose concentrations on fluorescence, we decided to subtract the fluorescent value of our control (E. coli transformed with K628006 in a 0 uM solution) from the rest of our values at every reading. This would remove the focus from the decreasing values, and instead place emphasis on the difference between individual arabinose concentrations. What we find is a much more interesting graph:
Figure 2 - This graph shows the fluorescence readings at 600 nm (485 exciting) of K628006-transformed E. coli in solutions of various arabinose concentrations. The value of the 0 uM control has been subtracted from the fluorescence values of each other arabinose concentration at every 5 minute reading, in order to show the scale of difference between each reading.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 125
Illegal NotI site found at 164 - 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 65
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
References
Steinberg et al (1997) 'Protegrin-1: a broad-spectrum, rapidly microbicidal peptide with in vivo activity Antimicrobial Agents and Chemotherapy', Aug 1997, 1738-1742, Vol 41, No. 8
Lam et al (2006) 'Mechanism of Supported Membrane Disruption by Antimicrobial Peptide Protegrin-1' J. Phys. Chem. B, 2006, 110 (42), pp 21282–21286
Bierbaum, G. & Sahl, H.-G (1985) 'Induction of autolysis of Staphylocci by the basic peptide antibiotics pep5 and nisin and their influence on the activity of autolytic enzymes.' Arch. Microbiol. 141, 249 ± 254
Aumelas et al. (2004) 'Synthesis and Solution Structure of the Antimicrobial Peptide Protegrin-1' European Journal of Biochemistry Volume 237, Issue 3, pages 575–583, May 1996
//collections/antimicrobial
| None |