Part:BBa_K525221
S-layer cspB from Corynebacterium halotolerans with TAT-Sequence and lipid anchor, PT7 and RBS
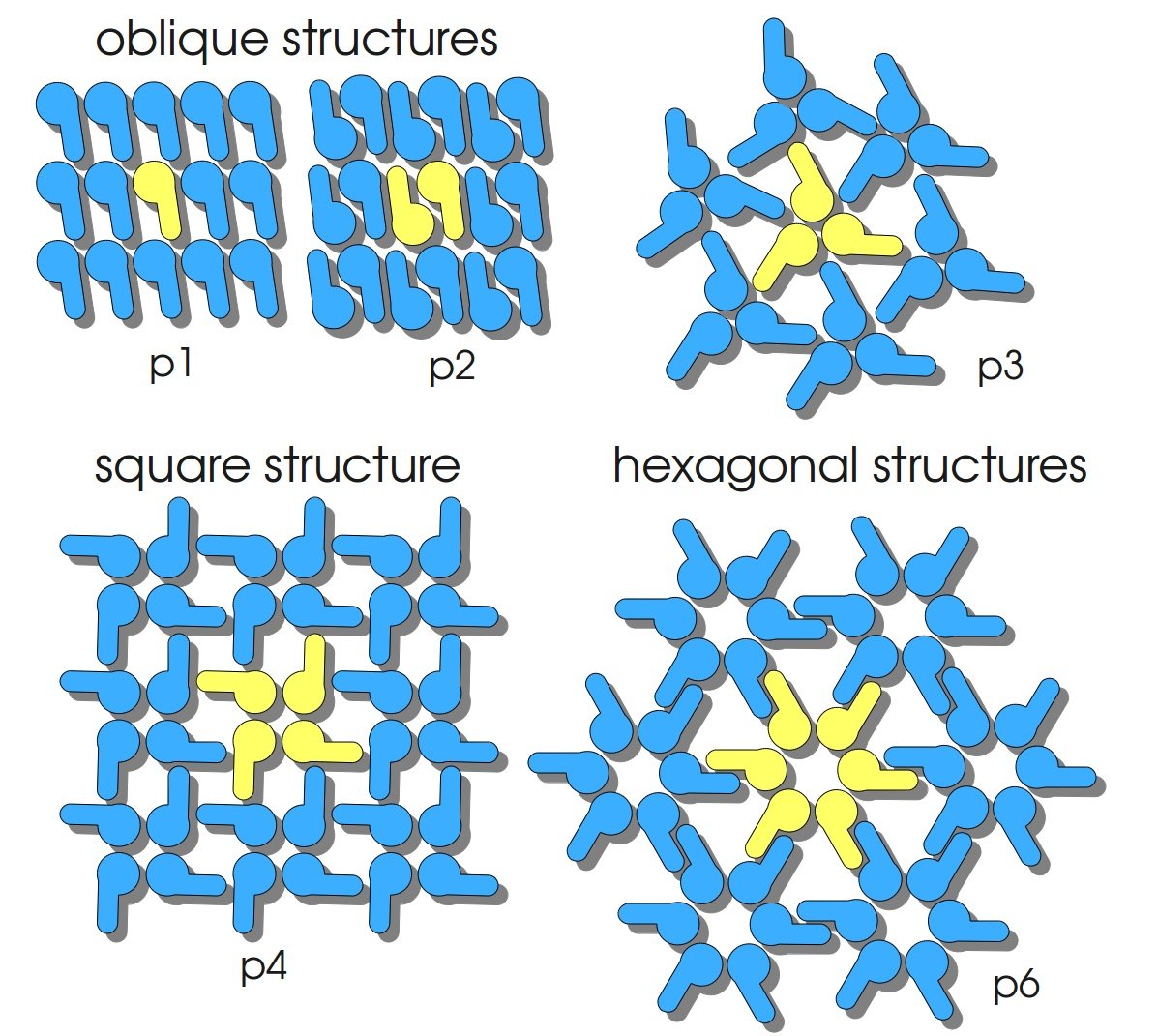
S-layers (crystalline bacterial surface layer) are crystal-like layers consisting of multiple protein monomers and can be found in various (archae-)bacteria. They constitute the outermost part of the cell wall. Especially their ability for self-assembly into distinct geometries is of scientific interest. At phase boundaries, in solutions and on a variety of surfaces they form different lattice structures. The geometry and arrangement is determined by the C-terminal self assembly-domain, which is specific for each S-layer protein. The most common lattice geometries are oblique, square and hexagonal. By modifying the characteristics of the S-layer through combination with functional groups and protein domains as well as their defined position and orientation to eachother (determined by the S-layer geometry) it is possible to realize various practical applications ([http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.2006.00573.x/full Sleytr et al., 2007]).
Usage and Biology
S-layer proteins can be used as scaffold for nanobiotechnological applications and devices by e.g. fusing the S-layer's self-assembly domain to other functional protein domains. It is possible to coat surfaces and liposomes with S-layers. A big advantage of S-layers: after expressing in E. coli and purification, the nanobiotechnological system is cell-free. This enhances the biological security of a device.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 1191
Illegal XhoI site found at 647 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 314
Illegal NgoMIV site found at 1403
Illegal NgoMIV site found at 1514
Illegal AgeI site found at 89
Illegal AgeI site found at 305
Illegal AgeI site found at 546
Illegal AgeI site found at 593 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 995
Illegal BsaI.rc site found at 302
Illegal BsaI.rc site found at 680
Illegal BsaI.rc site found at 1082
//proteindomain/internal
//rnap/bacteriophage/t7
| n/a | S-layer cspB from Corynebacterium halotolerans with TAT-Sequence and lipid anchor, PT7 and RBS |