Coding
cspB
Part:BBa_K525222
Designed by: Anna Drong Group: iGEM11_Bielefeld-Germany (2011-09-11)
Revision as of 16:27, 21 September 2011 by JSchwarzhans (Talk | contribs)
S-layer cspB from Corynebacterium halotolerans
S-layer Corynebacterium halotolerans
Sequence and Features
Assembly Compatibility:
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 1103
Illegal XhoI site found at 559 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 226
Illegal NgoMIV site found at 1315
Illegal AgeI site found at 217
Illegal AgeI site found at 458
Illegal AgeI site found at 505 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 907
Illegal BsaI.rc site found at 214
Illegal BsaI.rc site found at 592
Illegal BsaI.rc site found at 994
Expression in E. coli
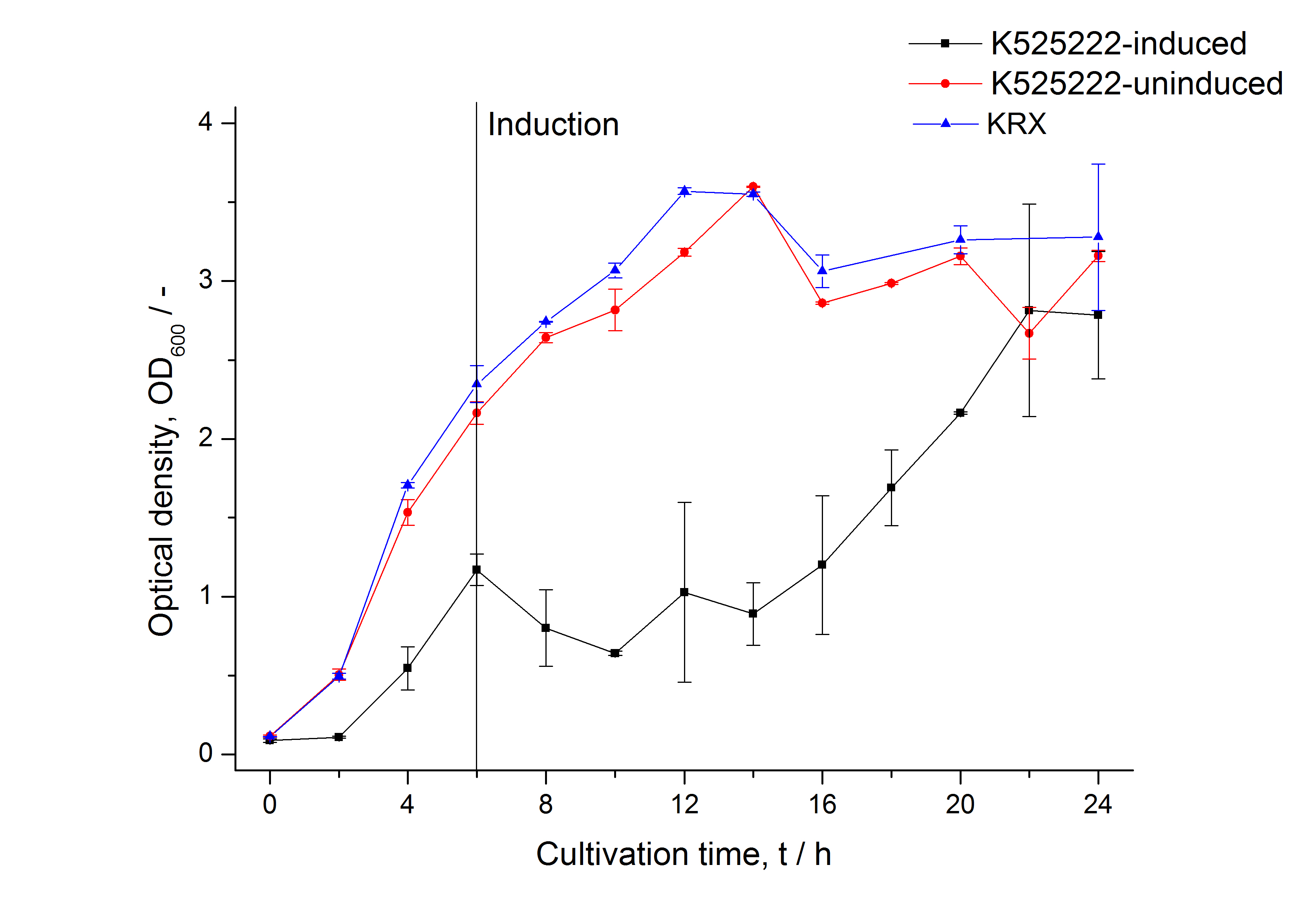
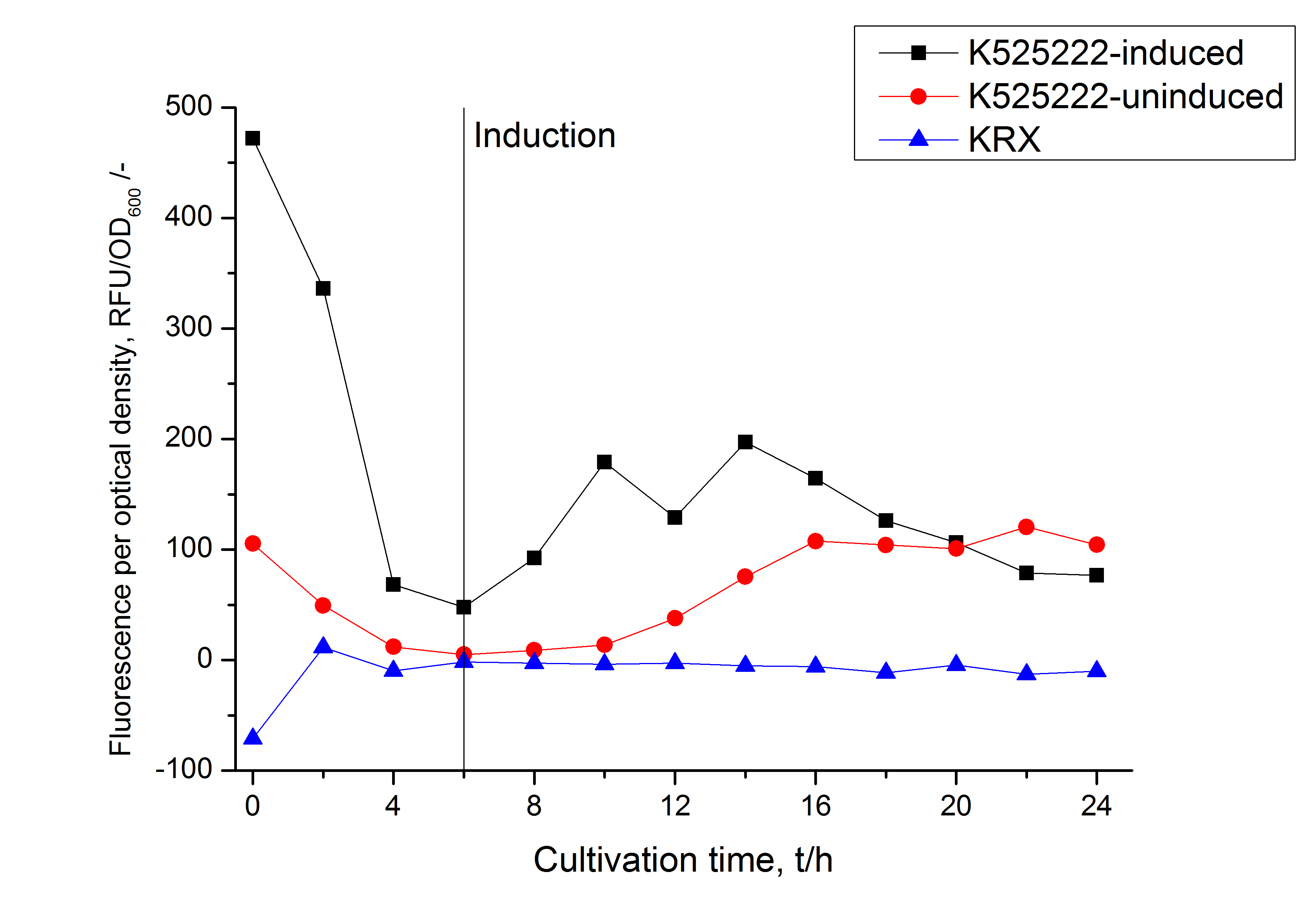
The CspB gen was fused with a monomeric RFP (BBa_E1010) using [http://2011.igem.org/Team:Bielefeld-Germany/Protocols#Gibson_assembly Gibson assembly] for characterization.
The CspB|mRFP fusion protein was overexpressed in E. coli KRX after induction of T7 polymerase by supplementation of 0,1 % L-rhamnose using the [http://2011.igem.org/Team:Bielefeld-Germany/Protocols/Downstream-processing#Expression_of_S-layer_genes_in_E._coli autinduction protocol] from promega.
[edit]
Categories
Parameters
//chassis/prokaryote/ecoli
//proteindomain/internal
//proteindomain/internal
| n/a | S-layer cspB from Corynebacterium halotolerans |