Part:BBa_K4140008
lacZ alpha
Part Description
The LacZ-alpha fragment is encoded by this region and is derived from the pUC19 cloning vector. Because it is smaller than the LacZ-omega region, the LacZ-alpha fragment can be easily incorporated into a plasmid when the two non-functional LacZ gene fragments (alpha and omega) are co-expressed. X-gal (5-bromo-4-chloro-3-indoyl-d-galactopyranoside), a soluble colourless molecule that is a substrate of ß-galactosidase and generates a blue product when cleaved, is used to detect the alpha-complementation
Usage
alpha domain of ß-galactosidase (our reporter protein) which is produced in response to high levels of phenylalanine and generates a blue product when X-gal is cleaved which is a soluble colorless molecule that is a substrate of ß-galactosidase and generates a blue product when cleaved, is used to detect the alpha-complementation
Improvements
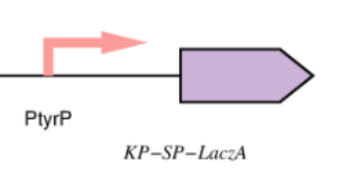
Improvement by KP-SP tagging
This year, we enhanced the LacZ alpha gene by incorporating a peptide signal that controls the secretion of B galactosidase extracellularly in an effort to improve the cleavage capacity of the X gal, heighten the intensity of the dark blue color emitted by the X gal product, and reduce the amount of time required to receive the results after performing the test on the chip
Improvement of LacZ alpha gene which encodes for the alpha domain of B-galactosidase that acts as a reporter protein in our therapeutic circuit :
Lactose converting enzyme (B-gal) beta-galactosidase that is originated from bacteria And have been studied as a reporter protein for many years ,It's a polypeptide of 1029 amino acid, and its tetramerization carries out the functional form of the enzyme.
The deletion of the tetramerization domain which is the N-terminal region spanning the first 50 residues will abolish its enzymatic activity, which could be fully restored by the alpha complementation of the intact N- terminal portion of the (Beta-gal) this alpha complementation is detected by using a beta-galactosidase colorless substrate as X gal to be cleaved into a dark blue product.
This year we employed this approach to indicate the presence of phenylalanine within our whole cell biosensor on the test line with media containing X gal substrate of our platform to diagnose phenylketonuria
Improvement by Directed Evolution
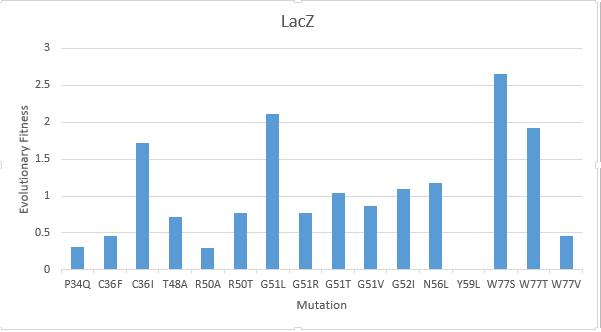
After creating a multiple sequence alignment of the protein sequence and predicting mutational landscapes, the effect of these mutations on the evolutionary fitness of the protein is tested. The prediction of the mutational landscape by saturation mutagenesis of the LacZ protein. The (W77S ) mutation, as depicted in the chart, had the greatest score when compared to other mutations. On the other hand, it's clear that the (Y59L) had the least evolutionary fitness for LacZ protein. As displayed in Figure(2)
References
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 93
Illegal XhoI site found at 144 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
| None |