Part:BBa_K2457001
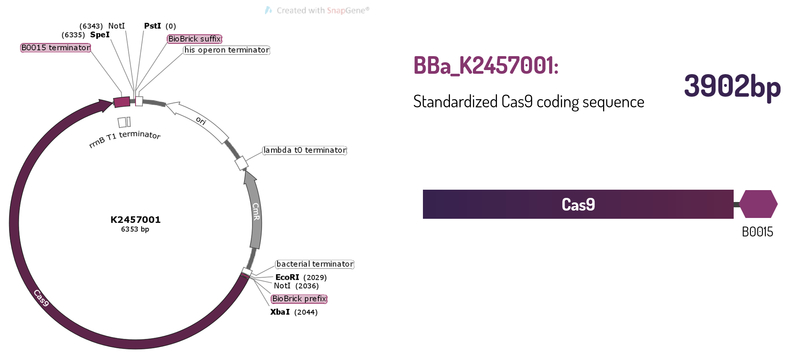
Standardized Cas9 coding sequence + B0015 terminator
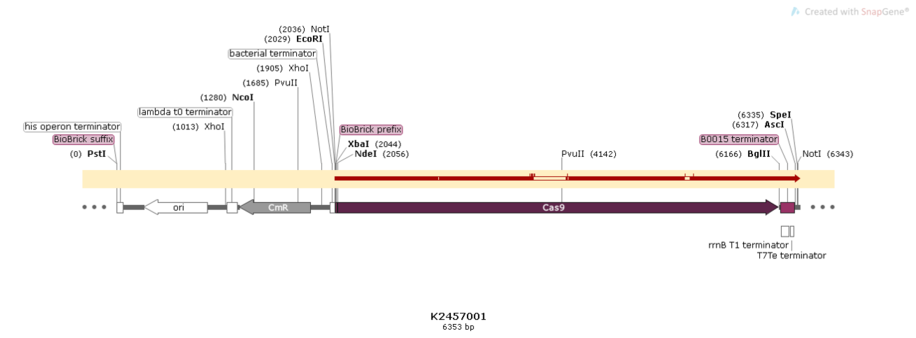
This BioBrick was engineered to be a building block of the standard BioBrick framework for bacterial genome engineering: the ([http://2017.igem.org/Team:Amazonas_Brazil pCRISPeasy]). BBa_K2457001 is composed of the Cas9 coding sequence from Streptococcus pyogenes + BBa_B0015 double terminator. It was rationally designed to be interchangeable AND standardized, providing a BioBrick to attend multiple SynBio purposes. For that matter, the RBS and promoter are easily switchable through 3A Assembly. It’s also standard restriction enzymes safe for all RFCs assemblies.
Figure 1: BBa_K2457001 circuit.
Usage and Biology
In bacteria and archaea, the CRISPR system associated to the Cas proteins is, shortly, a clustered regularly interspaced short palindromic repeats: repeated sequences interspaced by small unrepeated sequences (spacers) resulted from exogenous DNAs segments (protospacers) (Nishimasu et al., 2014). A natural mechanism that promotes immunity through invasors bacteriophage fragments transduction or plasmidial DNA incorporation in prokaryotes CRISPR locus (Church, 2013). The Cas9 (CRISPR-associated protein 9) originated from CRISPR system type II is a tool that has been very explored in genome edition methodologies, it does a sequence-specific cleavage of DNA. Compared to other genomic editing systems, this protein detaches itself for the site-specific cleavage programming possibilities through a single guide RNA (sgRNA). Due to its versatility, it is unnecessary to design a new protein to each target sequence, as in the traditional methods, like ZFNs and TALENs. Thus, the genetic edition by Cas9 has been integrating innovative applications.
The crystallography made by Jinek (2014) and Nishumasu et al. (2015) revealed the presence of two lobules in Cas9 protein: the first one is Recognition (REC) dividing itself into three parts, an extensive alpha helix, called bridge helix (60-93 residues), REC1 domain (94-179 and 308-713 residues) and REC2 domain (180-307 residues); the second lobule is from nucleases (NUC), splitting in RuvC in aminoterminal which is divided into three discontinuous segments (RuvC-I to RuvC-III), (1-59, 718-769 and 909-1098 residues), HMN in central part (775-908 residues) and the PAM-interacting domains, which is related to the carboxyterminal (1099-1368 residues).
This part codes for a standardized Cas9-version and a B0015 terminator. It can be applied to genomic editing, especially in bacteria, since Cas9 is central to this tool.
Assembly
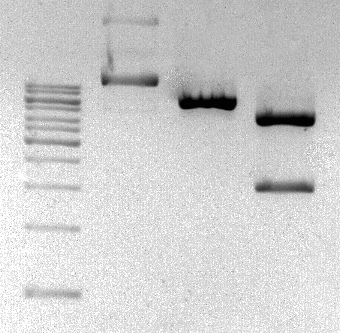
The sequence was ordered as two gBlocks from Integrated DNA Technologies and it was assembled by PvuII restriction enzyme site.
Figure 2: Eletrophoretic profile of restriction digestion of BBa_K2457001.
Characterization
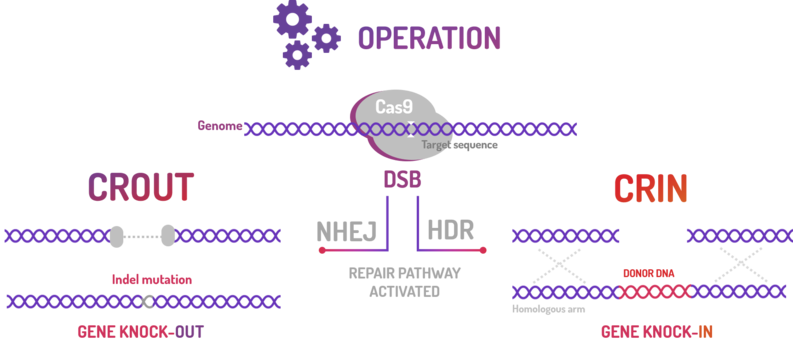
BBa_K2457001 was used by Amazonas_Brazil team in two methods based on standard BioBricks modules for bacterial genome engineering mediated by CRISPR-Cas9 machinery: the CRISPeasys. The approaches constructed were 1) for non-homologous end joining (NHEJ) repair pathway, the CRISPeasy-out method (or crOUT), referring to knock-out and the 2) for homologous directed repair (HDR), the CRISPeasy-in method (or crIN), referring to knock-in.
Figure 3: CRISPeasy subdivisions.
was used by Amazonas_Brazil team in two methods based on standard BioBricks modules for bacterial genome engineering mediated by CRISPR-Cas9 machinery: the CRISPeasys. The approaches constructed were 1) for non-homologous end joining (NHEJ) repair pathway, the CRISPeasy-out method (or crOUT), referring to knock-out and the 2) for homologous directed repair (HDR), the CRISPeasy-in method (or crIN), referring to knock-in. </p>
1. CROUT (CRISPeasy-out method): Designed for knock-out strategy for bacterial genome editing based on CRISPR/Cas9
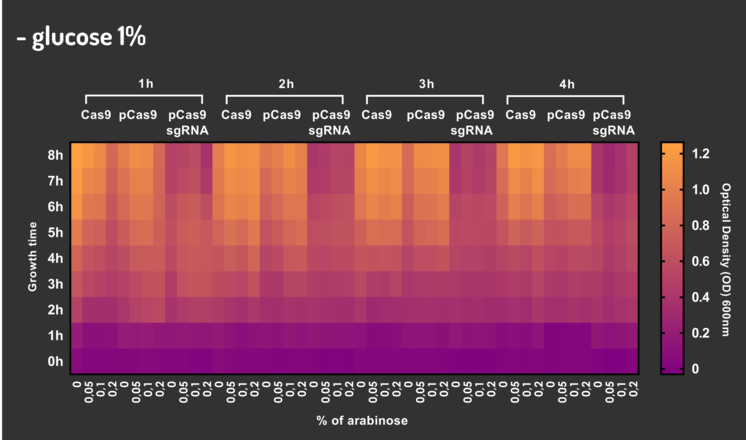
In order to characterize our genetic modules, It was transformed by electroporation E. coli K12 MG1655 with BBa_K2457006 (pBAD_Cas9_sgRNA). We took isolated colonies to pre-inoculate in 5mL of Luria-Bertani (LB), incubating overnight in a shaker at 37ºC. After that, we measured the Optical Density (OD) on spectrophotometer 600 nm aiming to standardize the cell ratio to 1 OD. All the samples were inoculated to 0.05 OD. Our experiment was performed in 96-well plates with 200µL of LB + 25µg/mL chloramphenicol, measuring the cell growth for over 8h in a range of conditions (see the layout plate below). We implemented Hidex Chameleon spectrophotometer to measure the OD and incubate the 96-well plates into shaker at 37°C. As a control, we employed MG1655 carrying two genetic circuits: BBa_K2457001 (Standardized Cas9 coding sequence without regulatory modules) AND BBa_K2457005 (Cas9 device without sgRNA).
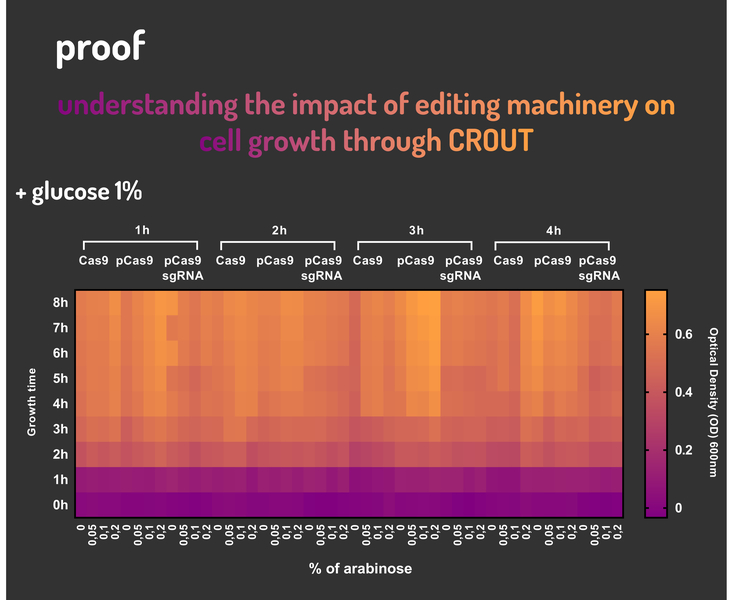
Glucose 1% was applied as a repressor to understand the activation and deactivation of the Cas9 expression and was added in 50% of the samples. The L-Arabinose solution is used as the inductor of the AraC_pBAD regulatory module controlling the Cas9 expression and was added in 75% of the samples, on respective concentrations: 0,05%, 0,1% and 0,2%. It was added at 1, 2, 3 and 4 hours of cell growth.
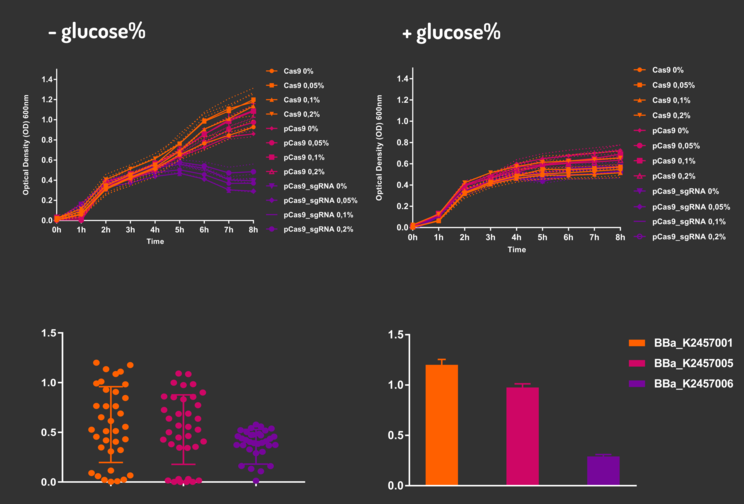
Figure 4: CRISPeasy-out results.
Based on a range of experiments involving the knocking-out process, we could propose that in the glucose (repressor) absence AND in the arabinose presence (inductor), MG1655 transformed with the BBa_K2457006 vector, carrying the Cas9 device and constitutively expressing sgRNA, reported a meaningful lower growth in contrast with the samples without the regulatory module (BBa_K2457001) OR without the sgRNA (BBa_K2457005). And, in the glucose presence, repressing the Cas9 expression, all the samples displayed a similar growth performance. The samples which carried the sgRNA (i.e., directing the Cas9 sequence to lead the DSB into a specific point of the genome) presented a low cell viability, supporting the hypothesis that even occurring the Non-Homologous End Joining (NHEJ) cell repair, not all cells could recover themselves, leading to the cellular death. Thus, we founded our proof of concept on the well-characterized impact caused by the Cas9 activity into the cellular machinery AND cell growth.
2. CRIN (CRISPeasy-in method): Designed for knock-in strategy for bacterial genome editing based on CRISPR/Cas9
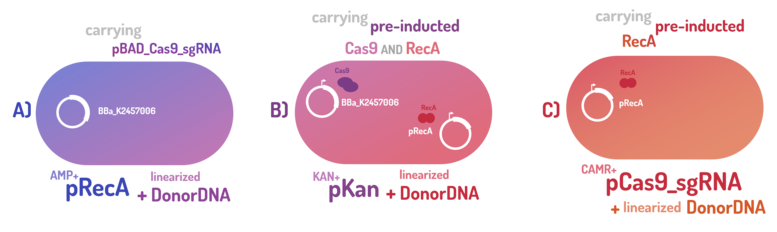
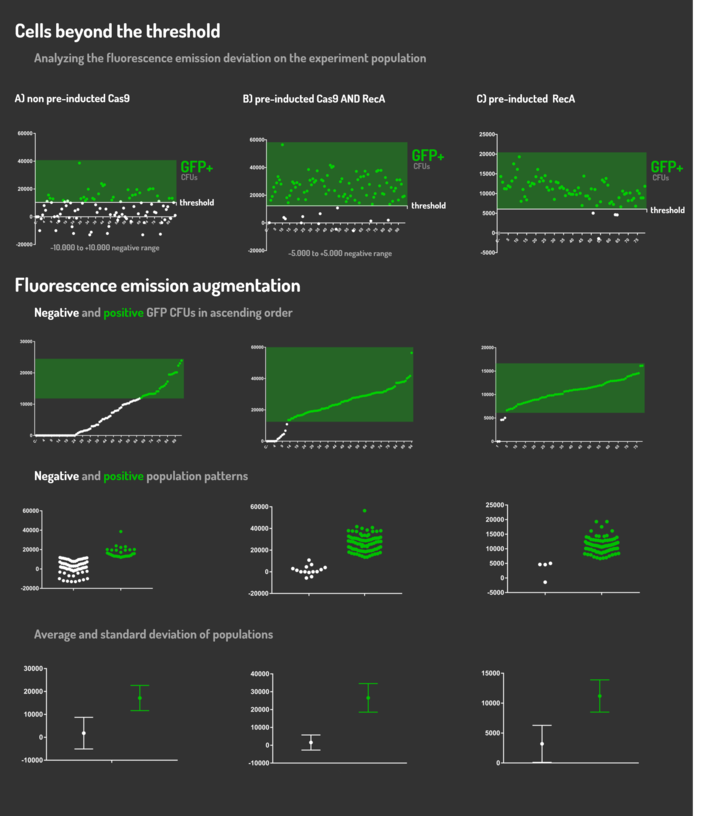
As the main key to this method, there is an AND logic gate composed of Cas9 coupled with RecA, which operates the Donor DNA insertion into E. coli genome through Direct Homology Repair (HDR). In this way, the arabinose AND IPTG (the inputs) activate the gene expression program encoding Cas9 and RecA. Then, there is the execution of their hardware's (physical parts, e.g. proteins) operation. As an output, Cas9 AND RecA activate the DNA insertion into the genome. In order to standardize a reliable datasheet of genome engineering efficiency, we tested a range of electrocompetent E. coli K12 MG1655 cells with different circuits: test A) carrying a non-pre-inducted Cas9 device + sgRNA (BBa_K2457006) plasmid; B) carrying a pre-inducted Cas9 AND RecA; C) carrying a pre-inducted RecA. The donor DNA was electroporated in all the tests.
Figure 5: CRISPeasy-in methodology.
A) carrying a non-pre-inducted Cas9 device + sgRNA;
We transformed through electroporation E. coli K12 MG1655 carrying a non-pre-inducted pBAD_Cas9_sgRNA plasmid CamR+ (BBa_K2457006) with pRecA (AmpR+) and a linearized Donor DNA (BBa_K2457004). We screened over 94 transformants colonies, inoculating in 200µL of LB + 25µg/mL chloramphenicol + 100µg/mL ampicillin + 0.2% L-Arabinose + 1mM IPTG to activate the expression of RecA AND Cas9. We measured the initial OD (600nm) and initial fluorescence (exc. 340 nm em. 530 nm) on 96-wells plates applying the Hidex Chameleon V spectrofluorimeter. The process was repeated for each hour over 10 hours while the 96-wells plate kept growing on a shaker at 37ºC.
B) carrying a pre-inducted Cas9 + sgRNA AND RecA;
We transformed through electroporation E. coli K12 MG1655 carrying a pre-inducted pBAD_Cas9_sgRNA plasmid CamR+ (BBa_K2457006) AND pRecA plasmid AmpR+ with a linearized Donor DNA (BBa_K2457004) + pKana. We screened over 94 transformants colonies, inoculating in 200µL of LB + 50µg/mL kanamycin + 25µg/mL chloramphenicol + 100µg/mL ampicillin + 0.2% L-Arabinose + 1mM IPTG to activate the expression of RecA AND Cas9. We measured the initial OD (600nm) and initial fluorescence (exc. 340 nm em. 530 nm) on 96-wells plates applying the Hidex Chameleon V spectrofluorimeter. The process was repeated for each hour over 10 hours while the 96-wells plate kept growing on a shaker at 37ºC.
C) carrying a pre-inducted RecA.
We transformed through electroporation E. coli K12 MG1655 carrying a pRecA (AmpR+) with pBAD_Cas9_sgRNA plasmid CamR+ (BBa_K2457006) and a linearized Donor DNA (BBa_K2457004). We screened over 80 transformants colonies, inoculating in 200µL of LB + 25µg/mL chloramphenicol + 100µg/mL ampicillin + 0.2% L-Arabinose + 1mM IPTG to activate the expression of RecA AND Cas9. We measured the initial OD (600nm) and initial fluorescence (exc. 340 nm em. 530 nm) on 96-wells plates applying the Hidex Chameleon V spectrofluorimeter. The process was repeated for each hour over 10 hours while the 96-wells plate kept growing on a shaker at 37ºC.
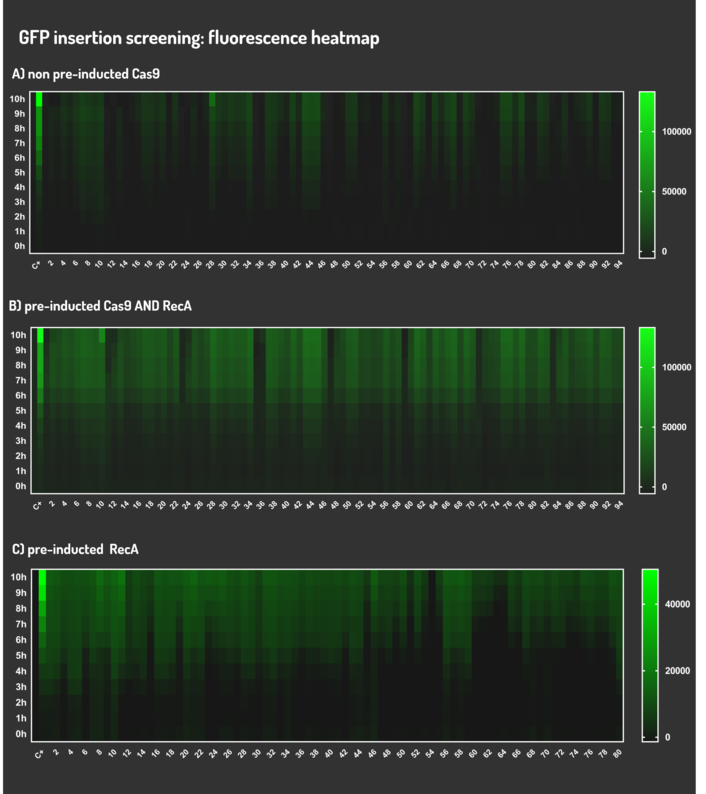
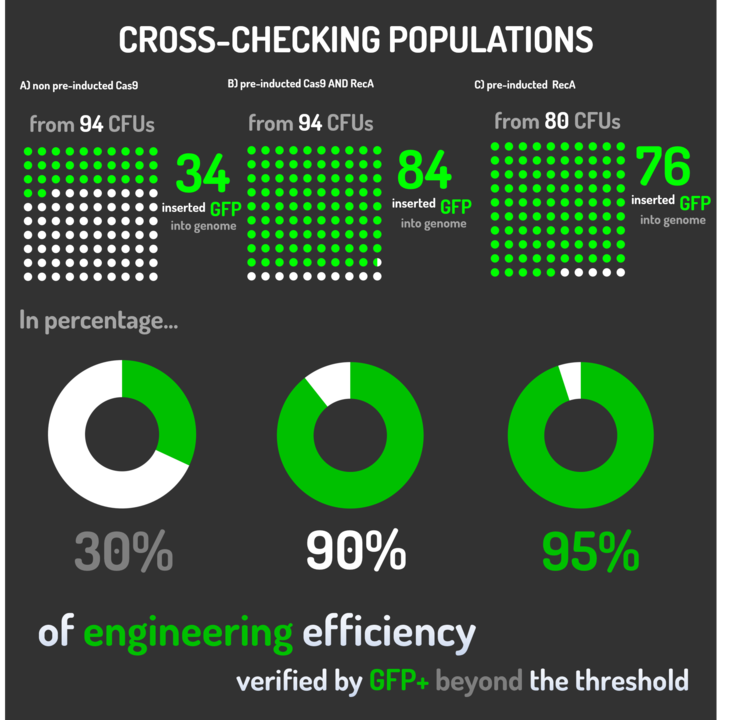
Figure 6: CRISPeasy-in results.
Leading from the achieved results, it was noted that has certain variations between the methodologies performances: with non-induced Cas9 (30% recombinants), with pre-induced Cas9 and RecA (90% recombinants) and with exclusively pre-induced RecA (95% recombinants).
The non-induced Cas9 method possibly do not have the best performance (30%) due to the absence of Cas9 protein when the Donor DNA is inserted into genome, causing a delay in the target sequence cleavage by Cas9, and, therefore, also the delay in homologous recombination, then the major linear Donor DNA is degraded in the bacterial cytosol.
The pre-induced Cas9 and RecA presented a very good result and similar to the most effective approach. When Donor DNA is inserted into the genome, is already happening the cleavage by Cas9 and RecA is investigating a homologous sequence to the broken strand, then Donor DNA is quickly incorporated, having less DNA degradation.
We found out that the most efficient method was the one which had only RecA pre-induced, since Cas9 presence affects the cells growth, therefore, when Cas9 and Donor DNA are added simultaneously, it will have the normal cell growth, beyond the further homologous recombination.
The CRISPeasy framework development can be a boost for an easily AND reliable way to use bacterial genome editing based on CRISPR/Cas9. We encourage teams to dive into CRISPR's standardization with us and expect to be able to see in the next years more and more projects with technique's improvement!
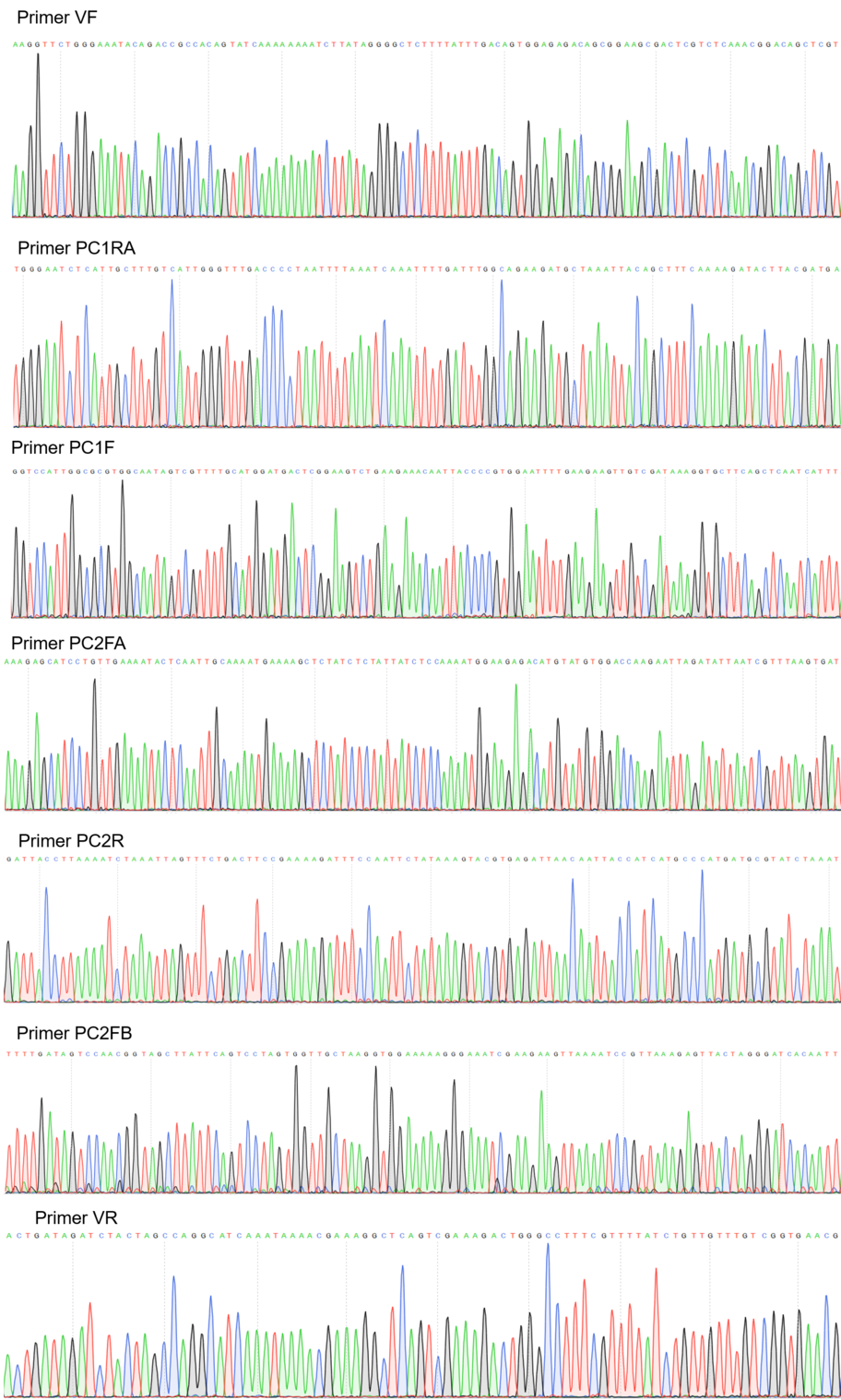
Sequencing
Figure 2:Sequencing electropherogram from BBa_K2457001.
Figure 3:Alignment of the designed sequence and our final construction from BBa_K2457001.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
| None |