Part:BBa_K510022
attTn7
TnsABC+D transposition machinery causes the site-specific insertion of the Tn7 transposon in the attTn7 (from attachment site) sequence, which is found in many bacteria. Our [http://2011.igem.org/Team:UPO-Sevilla/Foundational_Advances/MiniTn7/Bioinformatics/attTn7_Insertion_Site Bioinformatic analysis] reveal that the insertion site is well conserved in Gram-negative bacteria, an even in higher organisms until humans. However, here we add a portable attTn7 to allow using the [http://2011.igem.org/Team:UPO-Sevilla/Foundational_Advances/MiniTn7/Overview miniTn7 BioBrick toolkit] in any organism after portable attTn7 integration, as well as in plasmids.
All the derivatives of the miniTn7 BioBrick toolkit could be inserted in the attTn7 site by transposition.
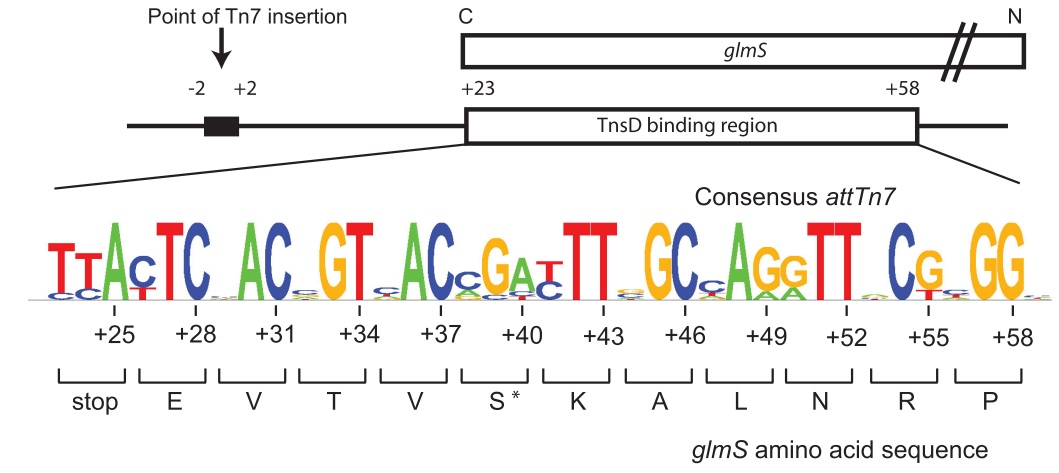
The organization of attTn7 (from [http://www.mobilednajournal.com/content/1/1/18 Rupak et al, 2010]). A schematic of the attTn7 at the C-terminus of the glmS gene with the TnsD binding site is shown. The sequence of the E. coli GlmS protein, and the consensus attTn7 sequence were derived as described in the text (note that the least conserved nucleotides correspond to the third position of each codon).
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
| None |