Part:BBa_K117002
LsrA promoter (indirectly activated by AI-2)
AI-2 quorum sensing system
The integral part of the AI-2 quorum-sensing (QS) system consists of a transporter complex, LsrABCD; its repressor, LsrR; and a cognate signal kinase, LsrK. Once uptaken into the cell, AI-2 will be phosphorylated under the effect of LsrK. Without the presence of AI-2, LsrA promoter is inhibited by the repressor, LsrR. Phosphorylated AI-2 will prevent LsrR from inhibiting this promoter, hence activating it so that the downstream sequence can be expressed. To prevent the cell from self-inducing this promoter, we must suppress its AI-2 producing capability. It can be done by knocking out LuxS, one of the vital gene required for producing AI-2. By doing that, LsrA promoter can only be activated under the presence of AI-2 from other sources than the host cell which is LuxS mutant (say, by contact with normal cells).
More information on AI-2 quorum-sensing (QS) system please refer to: http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1952038
Modelling The Lsr Promoter
Below are differential equations which can be used to model the production rate of the Lsr promomter, source here: https://doi.org/10.1371/journal.pcbi.1002172. To use these, teams must estimate a value of extracellular AI-2 concentration. For our own project we assumed that the Lsr-operon concentration produced would be the same as our protein being expressed by the Lsr promoter. We also created equations to model bacterial population and AI-2 production which can be found here: https://2021.igem.org/Team:Manchester/Model. It should be mentioned that the concentration given is an intracellular concentration so any secreted proteins must undergo further modelling or assumptions.
Variables
| Symbol | Description | Initial Value |
|---|---|---|
| t | Time | 0 minutes |
| [OP] | Lsr-operon concentration | 0 M |
| [REG] | Lsr Regulator Cocnentration | 0 M |
| [R] | LsrR Concentration | 0 M |
| [Ap] | Intracellular Phosphorylated AI-2 concentration | 0 M |
| [Aout] | Extracellular AI-2 Concentration | Estimate/model this uM |
Parameters
| Parameter | Description | Value |
|---|---|---|
| kop | Lsr-synthesis rate | 7 uM-1 min-1 [2] |
| kr | LsrR synthesis rate | 2 min-1 [2] |
| k1 | Repression Coefficient (Lsr-operon) | 0.2 uM [2] |
| k2 | Repression coefficient (LsrR) | 0.1 uM [2] |
| k3 | AI-2/LsrR binding rate | 0.5 uM-1 min-1 [2] |
| k4 | Repression coefficient (Lsr-regulator) | 65 uM [2] |
| k5 | REG/AI-2 interaction | 0.0001 uM-1 min-1 [2] |
| kf | AI-2 fluc through alternative pahways | 0.01 uM-1 min-1 [2] |
| kimp | AI-2 import via LasrABCD | 0.01 uM-1 min-1 [2] |
| nOP | Cooperativity coefficient (Lsr-operon) | 4 [2] |
| knR | class="model-th"Cooperativity coefficient (LsrR) | 4 [2] |
| kdeg | Protein decay rate | 0.02 min-1 [2] |
Equations
Equation 1
<img src="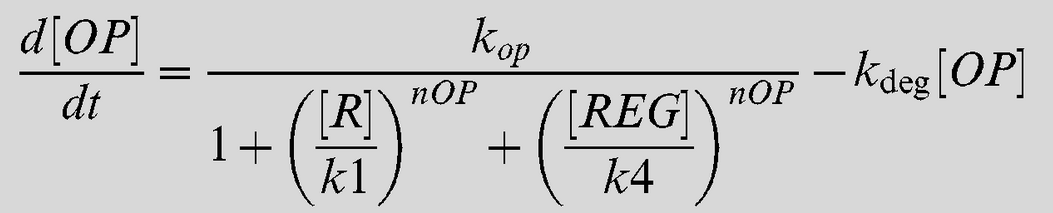
Equation 2
<img src='https://2021.igem.org/wiki/images/5/5d/T--Manchester--Equation_2.PNG'>
Equation 2 is describing the rate of change of a hypothetical Lsr-regulator molecule suggested to exist in the source literature. [2]
Equation 3
<img src='https://2021.igem.org/wiki/images/7/7e/T--Manchester--Equation_3.PNG'>
Equation 3 describes the rate of synthesis of LsrR
Equation 4
<img src='https://2021.igem.org/wiki/images/1/1c/T--Manchester--Equation_4.PNG'>
This equation describes the concentration of phosphorylated AI-2 inside the cell. This is contributed to by flux of AI-2 through the LsrABCD channel complex and alternative pathways. Ap is depleted by AI-2 binding to the LsrR protein and the theoretical Lsr-regulator molecule.
Equation 5
<img src='https://2021.igem.org/wiki/images/2/21/T--Manchester--Equation_5.PNG'>
Equation 5 gives us the rate of decrease of the exrracellular AI-2 concentration, this assumes that we have estimated an initial concentration of AI-2 which is not very realistic. In our model we added equations to decribe bacterial concentration and AI-2 production, further details can be found here:https://2021.igem.org/Team:Manchester/Model
Equation 6
<img src='https://2021.igem.org/wiki/images/6/6d/T--Manchester--Equation_6.PNG'>
This equation gives us the rate of change of the Lsr-regulator/AI-2 complex.
Equation 7
<img src='https://2021.igem.org/wiki/images/9/9e/T--Manchester--Equation_7.PNG'>
This equation gives us the rate of change of the LsrR/AI-2 complex.
</div>
</div>
Important notice:
- This part is only the promoter region of LsrA gene which is normally inhibited by LsrR. It is NOT the entire LsrA gene.
- It is INDIRECTLY induced by AI-2, through a complex network involving LsrK,LsrR.
How this promoter works:
By ligating LsrA promoter with a protein coding sequence downstream, we can regulate the expression of that protein by AI-2 as the inducer. To prevent self-inducing by the host cell, the plasmid must be transformed into cell with LuxS gene knocked out.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
//direction/forward
//chassis/prokaryote/ecoli
//promoter
//regulation/positive
| negative_regulators | |
| positive_regulators | 1 |

 1 Registry Star
1 Registry Star
