Part:BBa_J04450
RFP Coding Device
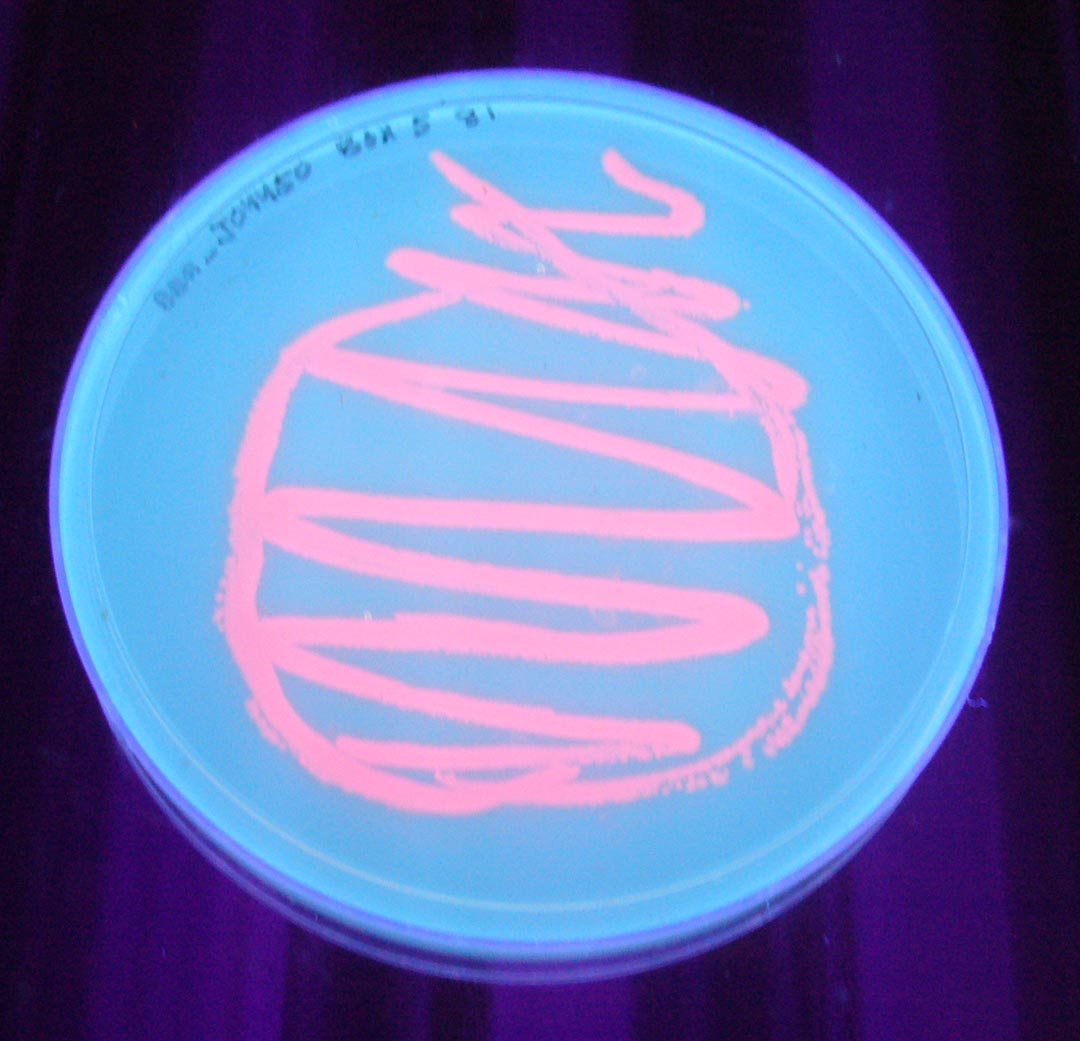
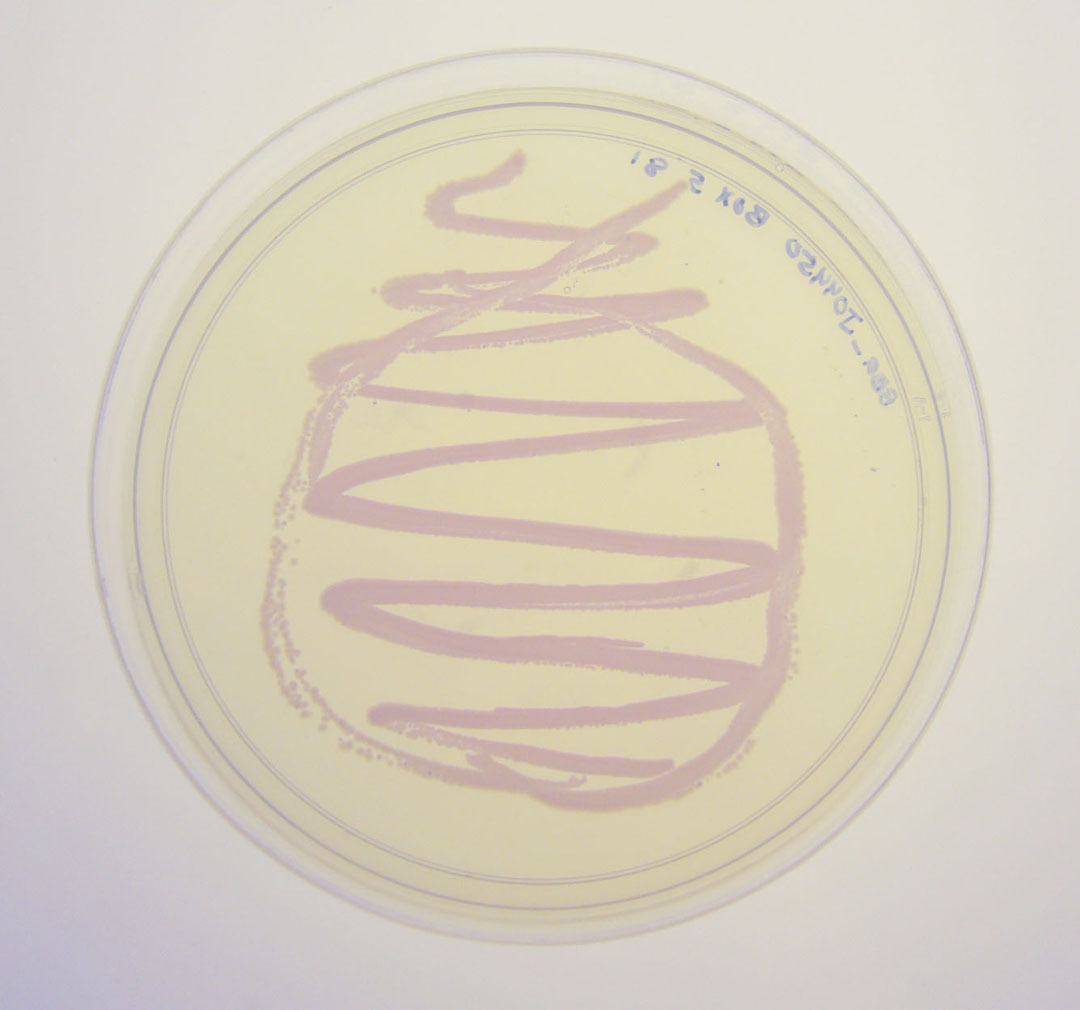
The colonies are clearly red in color under natural light after about 18 hours. Smaller colonies are visibly red under UV. The RFP part does not contain a degradation tag and the RBS is strong.
- LacI sensitive
- CAP sensitive
This part is commonly used, but can fail if the system contains LacI or CAP protein.
(--Meagan 15:39, 23 July 2009 (UTC))
[http://2012.igem.org/Team:TU_Munich Team TU_Munich 2012] improved this part by making it compatible to RFC10 and RFC25 (see: BBa_K801100)
(--VolkerMorath 15:02, 21 October 2012 (UTC))
[http://2013.igem.org/Team:NRP-UEA-Norwich Team NRP-UEA 2013] improved this part by adding a NdeI restriction site before the RFP gene. (see: BBa_K1041000)
(--holusac 20:46, 14 August 2013 (UTC))
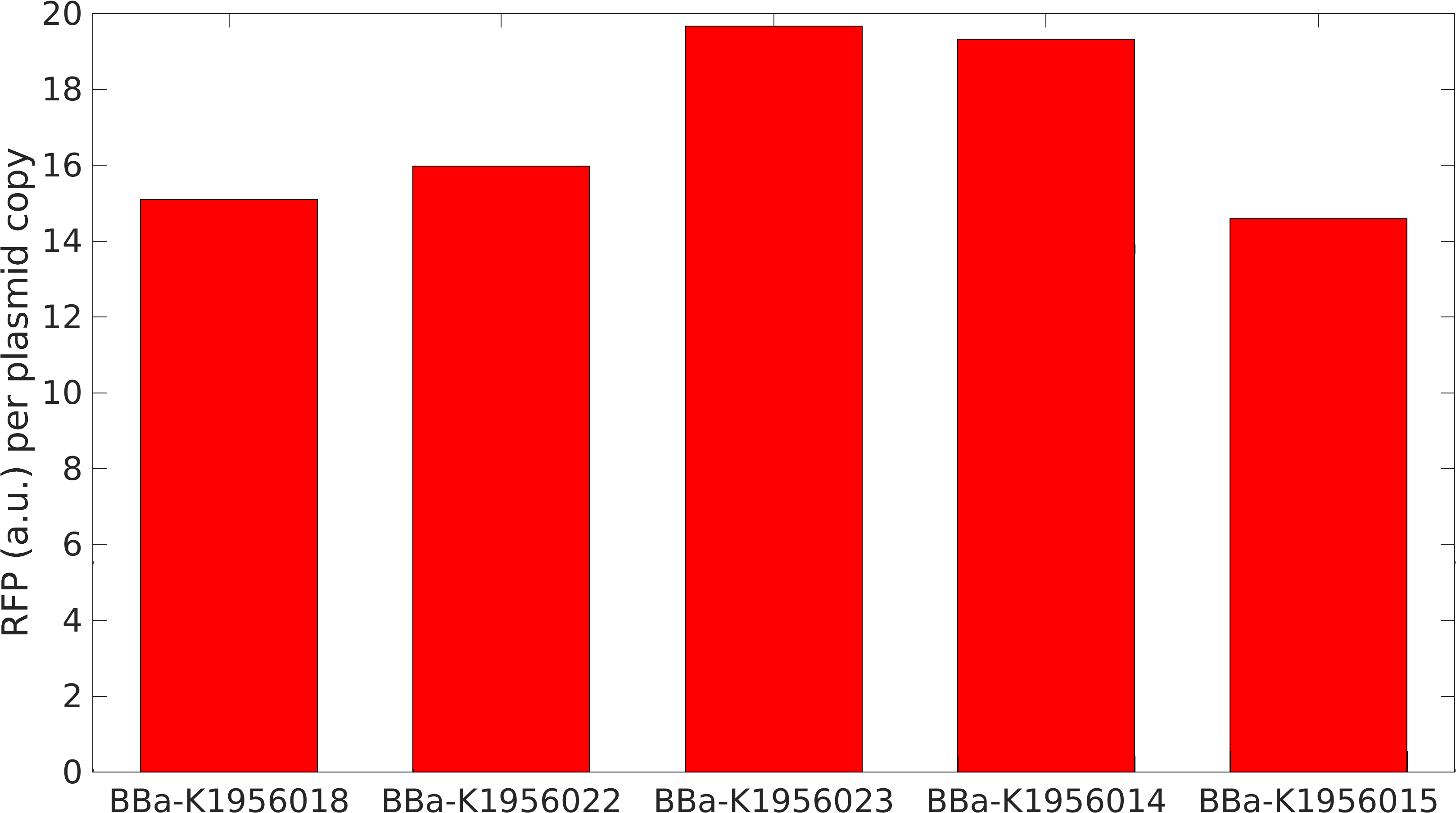
[http://2015.igem.org/Team:Warwick Team Warwick 2015] improved this part by analysing the effect of copy number on gene expression.
(--Lcarroll 20:48, 25 September 2015 (UTC))
[http://2016.igem.org/Team:Leiden Team Leiden 2016] contributed to the characterisation of this part by showing equal functionality in simulated microgravity (0g) as in the normal gravity of the Earth.
(--Valentijn 19:38, 19 October 2016 (UTC))
[http://2017.igem.org/Team:UChicago Team UChicago 2017] contributed to this part by improving/changing the documented sequence through mutagenesis to create blunt-end restriction sites for cloning not within the prefix/suffix region (created BBa_K2428000).
(--pzulueta97 21:14, 25 October 2017 (UTC) )
[http://2017.igem.org/Team:Grenoble-Alpes Team Grenoble-Alpes 2017] contributed to the characterisation of this part by testing the time of apparition of fluorescence, in presence of IPTG or not (because the promoter leaks), as well as they contributed to the improvement of this part by using its fluorescence as a detection signal to be able to detect Vibrio Cholerae.
(--NoreenLouis 20:47, 26 October 2017 (UTC) )
[http://2017.igem.org/Team:Kingsborough_NY Team Kingsborough NY 2017] contributed to the characterization of this part by showing decreased fluorescence when expressed either in a higher salt media - such as LB with 3% sodium chloride - or E. coli that lacks tmRNA, the principal component of the cell's ribosome rescue system. View the data on the experience page or [http://2017.igem.org/Team:Kingsborough_NY/RFP visit our Wiki]
(--djcamenares 17:56, 27 October 2017 (UTC) )
[http://2017.igem.org/Team:iTesla-SoundBio Team iTesla SoundBio 2017] contributed to the characterization of this part by analyzing the rate of false positives when using the coloring of transformed colonies as a red/white screen for determining experimental success.
(--gladish 01:26, 28 October 2017 (UTC) )
[http://2018.igem.org/Team:Grenoble-Alpes Team Grenoble-Alpes 2018] contributed to the characterisation of this part by testing the delay before apparition of fluorescence directly after transformation and the intensity of the leak, in three different E. Coli strains.
(--perrine 15:06, 9 October 2018 (UTC) )
Pictures
Team ITB_Indonesia 2017: Red color dynamics of cloned Escherichia coli strains in LB broth

In Team ITB_Indonesia 2017
characterization, we found in normal growth/incubation condition (37 oC, LB agar) that
BBa_J04450-transformed Escherichia coli BL21 colony appear to need longer incubation time (>18
hours) until it clearly shows red color under natural light.
We then investigate whether this phenomenon is influenced by the strain, and we try if there are
lac repressor in the system that can be released by inducing the culture with IPTG, hence
increasing the expression of mRFP.
Experimental Design
We used three different strains of transformed E. coli (BL21, DH5alpha, and Top10) for this
study. They were incubated in LB broth, 37 oC, and sampled every 4 hours for 2 days to
determine the red color absorbance at 588 nm. The amount of IPTG added for respective treatment is 500
µM.
Result and Findings
- There are no significant differences of mRFP expression in different strains of E. coli (BL21, DH5alpha, Top10)
- There are no significant effects of mRFP increased expression after IPTG induction.
- The red color absorbance under 588 nm wavelength is recorded around 2.5-3 OD units.
- The broth become red in color under natural light around 16-20 hours of incubation time.


 ==Pictures==
==Pictures==


Team INSA-UPS France 2017 : usage in Vibrio harveyi strain engineered by conjugation
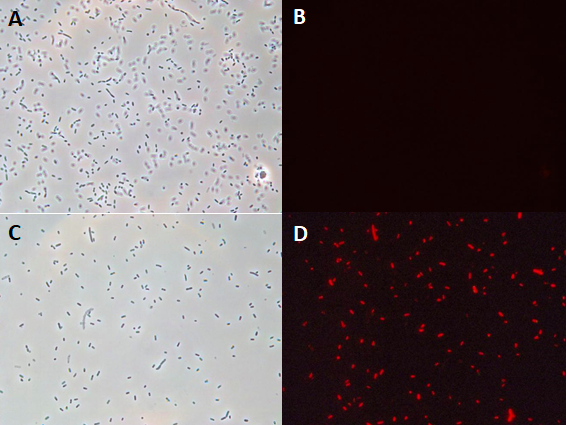
In the context of the iGEM INSA-UPS France project 2017, BBa_J04450 was tested in the Vibrio harveyi background. To the best of our knowledge, RFP has never been used in this strain. BBa_J04450 biobrick was cloned in a broad host range plasmid (pBBR1MCS-4) and conjugated into V. harveyi. The protocol of triparental mating can be found here. Its expression has been studied by fluoresence microscopy in Vibrio harveyi .
The microscopy results demonstrated the fonctional production of RFP in Vibrio harveyi, and hence, the functionality of part BBa_J04450 in this background.
IIT Madras 2016's Characterization
Experimentation
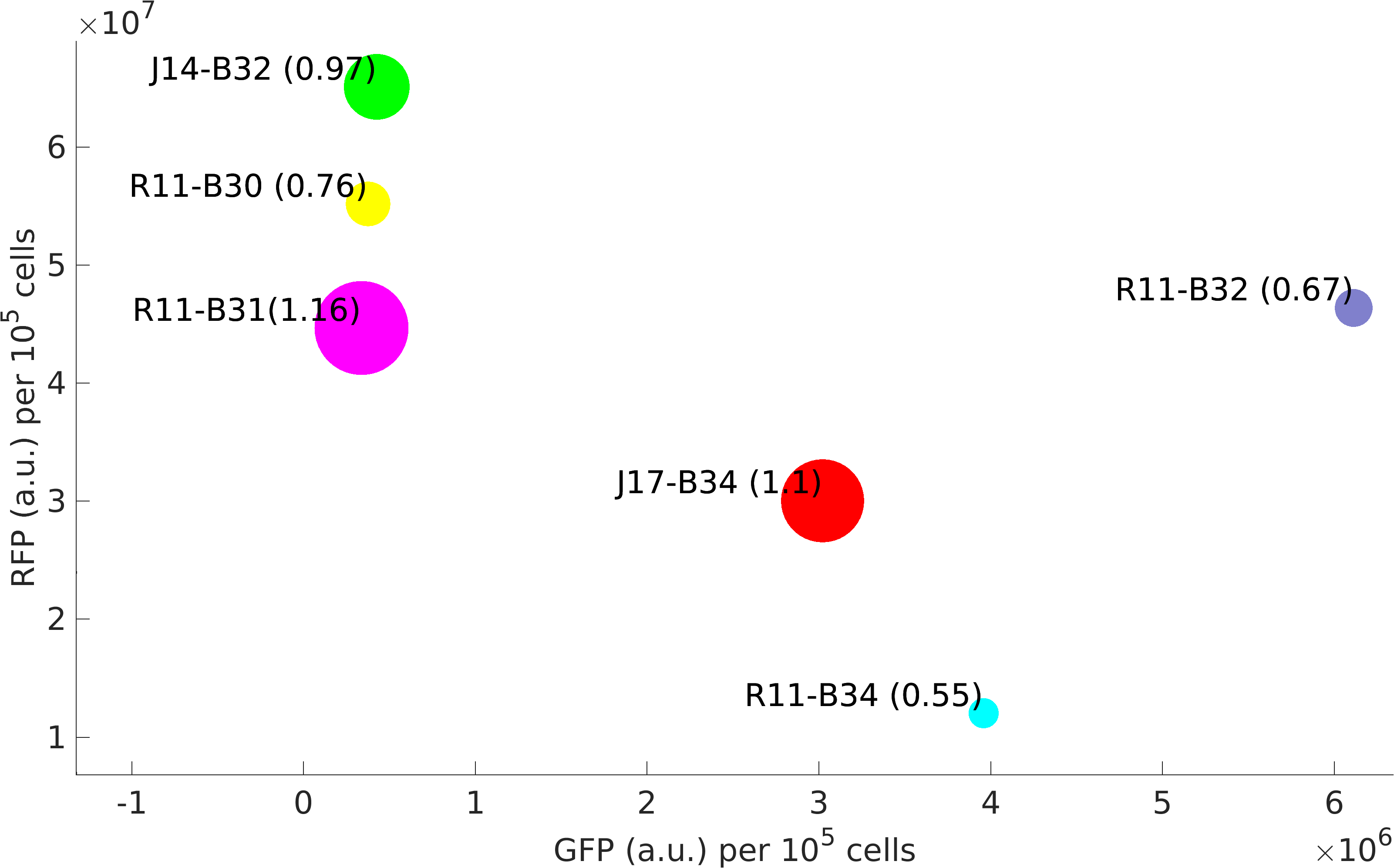
This BioBrick was used along with various GFP producing devices to understand the role of RBS and Promoter parts in giving rise to intrinsic noise in E. coli DH5alpha. Expression data for GFP and RFP proteins were obtained using flow cytometry (BD FACS Aria III) at 3hr, 6hr, 9hr and 12hr stage of growth along with cells expressing only GFP, only RFP and none. Cumulative intrinsic and extrinsic noise were measured using modified [http://2016.igem.org/Team:IIT-Madras/Model#Noise_in_Devices| Elowitz formula]. OD600 values for specific growth rate estimation were obtained using Spectrophotometer over an interval of an hour for 12 hours. Given specific growth rates are in it's logarithmic values. This BioBrick can be used to characterize noise and strength of complex devices by cloning this device with given device, which produces a different reporter protein. In graphs, we have R11-B32, R11-B34, J14-B3, J17-B34, R11-B30 and R11-B31 in pSB1A2 plasmid backbone.
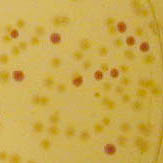
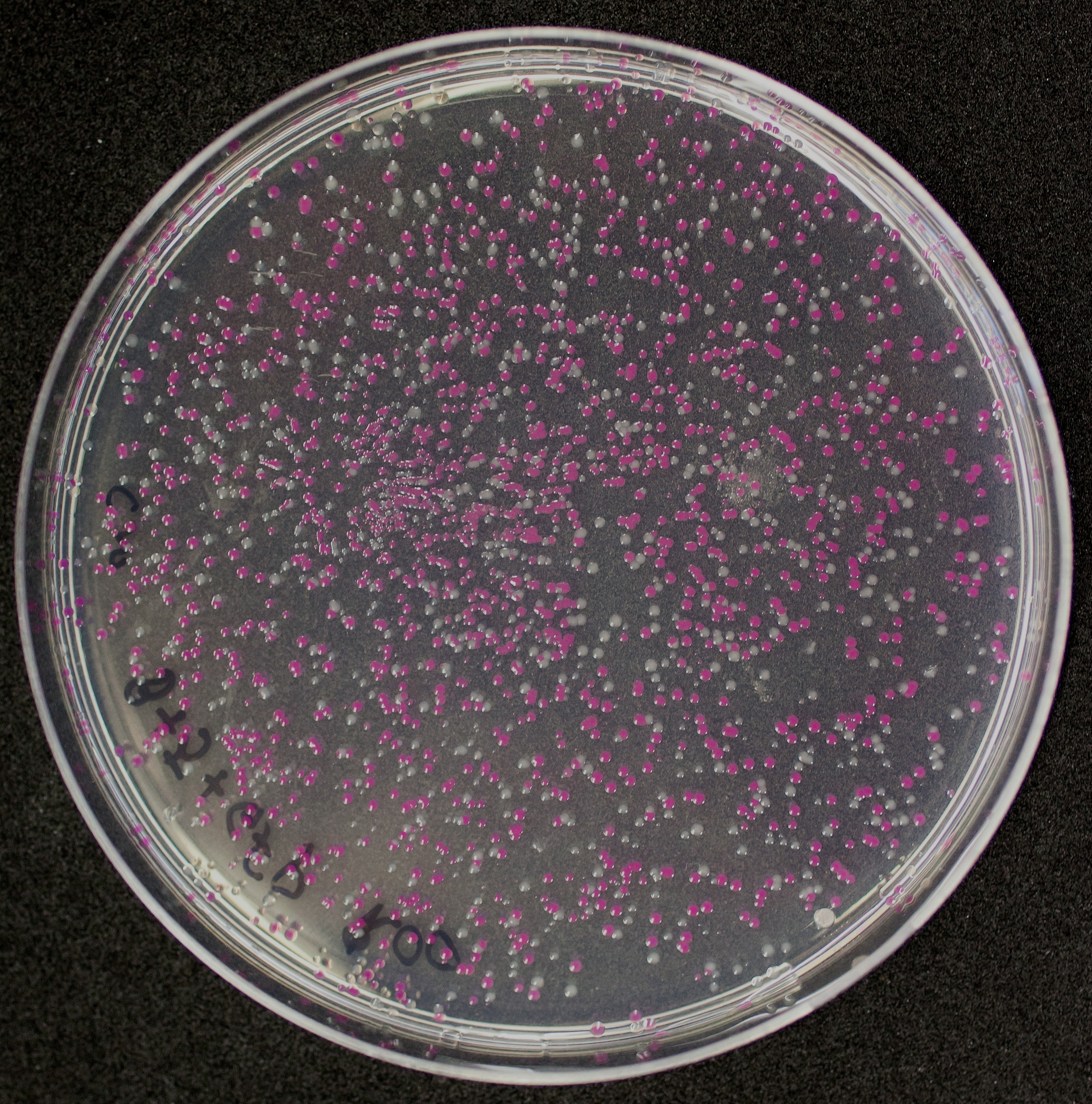
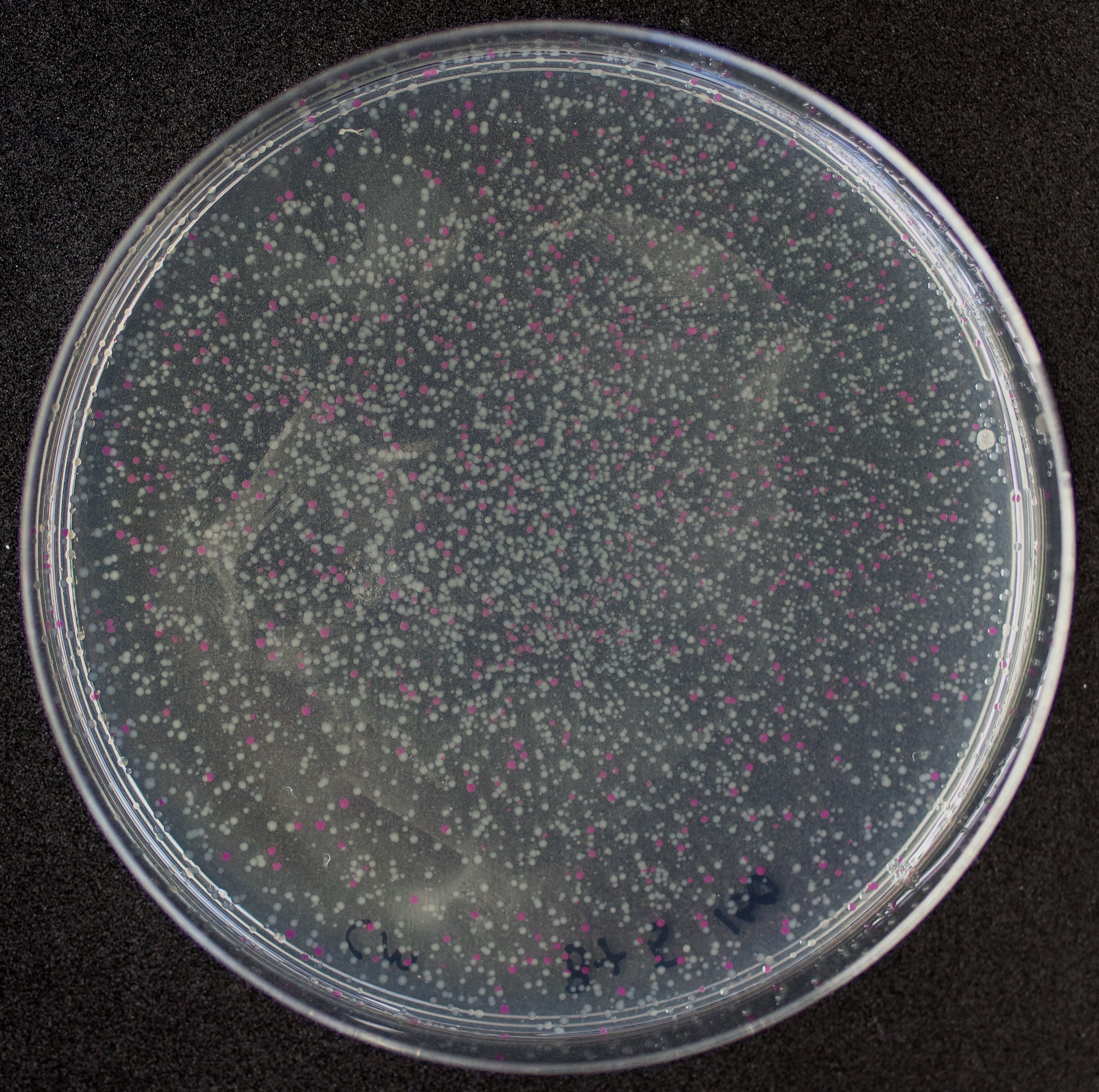
[http://2010.igem.org/Team:Groningen Team Groningen 2010] reports the usage of this part as a cloning tool. When ligating any part, or part assembly, into any standard backbone that contains this part, the non-restricted and single-restricted backbones that self-circularize will produce red colonies on rich media plates (we use TY). These undesired transformants can than be avoided in the screening for the correct construct. With this method, the backbone desired for a new construct does not need to be purified from agarose gel to decrease the amount of undesired tranformants caused by ligation of the original part present in the backbone. The amount of incorrect transformants depends, of course, on the ratio of backbone (mixed with J04450) vs. BioBrick insert, the size of the BioBrick insert, and whether the insert is an assembly of two BioBricks. The images below show two ligations with different efficiencies.
Usage in Chromobacterium Violaceum
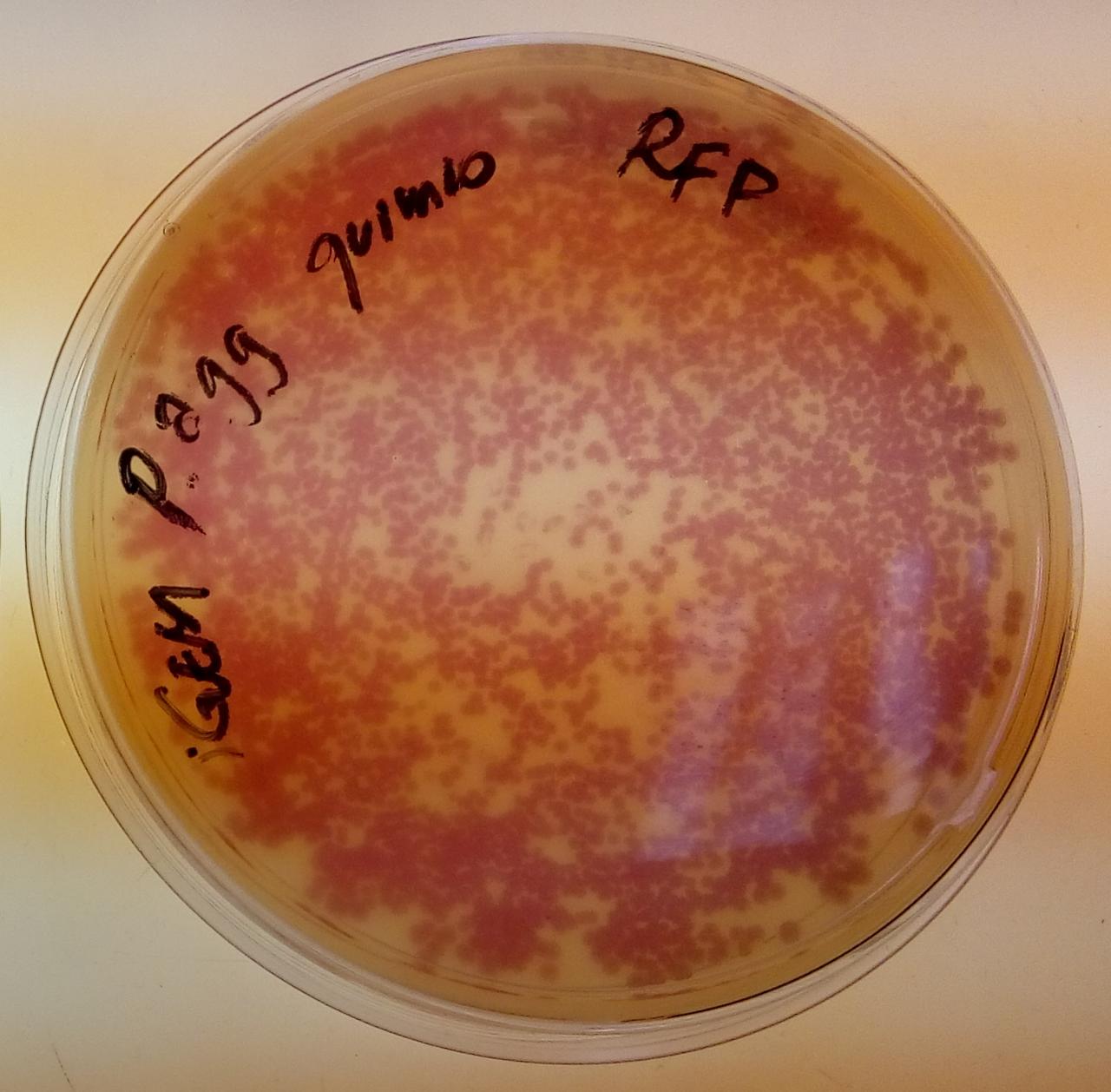
[http://2016.igem.org/Team:Tec-Monterrey Team Tec-Monterrey 2016] characterized the output of the part BBa_J04450 in a novel chassis, Chromobacterium Violaceum, as it produces a native purple pigment Violacein, we were curious whether RFP would be useful as a reporter gene. Furthermore, we characterized its expression under lac promoter. We did the transformation of C. Violaceum by a method that has not been reported yet, we made C. Violaceum competent cells with the protocol that is in our wiki, we concluded that the best O.D. for the heat shock transformation is 0.5 since it showed clearly better results than 0.4 or 0.6, we will continue to work in the transformation efficiency.
[Image: ]
]
Team Grenoble-Alpes 2018: Delay before fluorescence apparition and leaky expression after transformation in three Escherichia coli strains
The main goal of this characterisation was to find out which E. Coli strain was the most effective among three different strains (TOP10, DH5α and BL21), to obtain a rapid and high red fluorescence with pSB1C3-BBa_J04450.
We also studied the influence of adding 0.3mM of IPTG on the RFP expression in these three strains to see if the leak was different between host strains.
Indeed, former iGEM teams had described a leaky expression of fluorescence with this biobrick, which means there was no need to induce expression with iPTG to obtain a high rate of fluorescence.
As we raised the question in the context of our project, we thought it might be useful to know which E. Coli host strain is the better to use depending on the conditions you want. For instance, you might want to strictly control RFP expression and so you will need to reduce the leaky expression at most. On the contrary, you may want to have the highest leak as possible to avoid using IPTG, a need our team encountered for the using of our automated system.
EXPERIMENT
We wanted to test three E. Coli strains usually used in laboratories : DH5α, TOP10 and BL21, to see, in the context of our system, which one would be the host with the fastest and highest production of RFP.
We used a bacterial stock of our own, coming from commercial bacteria cultured and made competent using our protocol (see competency protocol here) a few weeks before the experiment and conserved at -80°C.
We made an heat-shock transformation (see transformation protocol here) of 23ng pSB1C3-BBa_J04450 in 25μL of each bacterial strains. We then cultured transformed bacteria in 250μL LB broth medium for one hour until they expressed chloramphenicol resistance. Then, 750μL of LB broth was added with chloramphenicol and incubated for one hour. After centrifuging the bacteria for 10 minutes at 5000 rpm, we replaced the old medium by resuspending the bacterial pellet. We also split the samples in half, and put each new samples in 30mL LB broth with chloramphenicol, completed or not with 0.3mM of IPTG.
We then measured red fluorescence (620nm) and O.D.600nm every hour for 27 hours.
NB: We had to do O.D.600nm manually as the TriStar microplate reader wasn’t able to do such measurements. Due to the experiment duration and the limited access to the laboratory, we were not able to make O.D.600nm measurements between 9 hours and 18 hours in the experiment. For the night, we thus launched automatic measurements for fluorescence, but we do not have any O.D.600nm linked to these measurements.
RESULTS
The results are represented in the following figures.
The data presented here start 17 hours or 18 hours after transformation because we didn’t observed any fluorescence until this point. This might be due to the huge culture volume in which we put bacteria (30mL). A huge volumed allowed us to make a long experiment with lots of measurements, but it also diluted a lot the bacteria and their fluorescence.

Figure 1
The first figure shows the fluorescence obtained for each sample, without taking into account the number of bacteria in the sample. These results were useful as a part of our project, as we wanted the highest fluorescence rate with the lowest time of apparition after transformation to improve the sensibility and speed of our detection system, regardless of the corresponding amount of bacteria.
The figure shows that fluorescence obtained with TOP10 E. Coli was a lot higher than the ones obtained with either BL21 or DH5α. 620nm measured fluorescence for TOP10 was twice as high as the one obtained with BL21 and more than 7 times as high as the one for DH5α. We also observed fluorescence a lot sooner with the TOP10 bacteria : 18 hours after transformation versus 23-24 hours with the other strains.
As for the DH5α and BL21 comparison, we can see that DH5α had a much lower fluorescence rate, whereas BL21 gave pretty good results when induced with ITPG.

Figure 2
The second figure represents the fluorescence rate rectified with O.D.600nm measurements. This allows to compare in a more quantitative way the fluorescence obtained, as if for each point, we had the same number of bacteria. It must be noticed that the differences in O.D.600nm were not clearly significative between the samples, which means that we can still compare them as they probably were in the same growing phase.
On this figure, we can see that there is no significant difference between the experiment with fluorescence only, so we can make the same observations as in figure 1.

Figure 3
The third and last figure represents the same results as in figure 1, but we made close-ups to better study each strain and the differences between induced and non induced promoter.
For the DH5α, we can see that there is a higher fluorescence rate for 0.3mM IPTG than without induction. However, we can still observe fluorescence without induction, which corresponds to the promoter leak : it represents almost half of fluorescence obtained with IPTG.
For the TOP10, we can see that there is almost no difference between the patterns obtained with or without induction. This means that the promoter leak is so important that IPTG induction is not significantly useful at all : the biobrick may be transformed and the RFP expressed in this host without using IPTG.
For BL21, the difference between induced and non-induced expression is clearly apparent : there is almost no fluorescence without IPTG, whereas expression with induction began to increase exponentially from 23 hours after transformation. This strain appears to be the most effective if you need to reduce the promoter leak as much as possible.
Results showed the differences between E. Coli host strains, which allow variable RFP expression rates, different delays before fluorescence apparition, and also particular responses to induction. The results are summarized in the following table :
| E. Coli strain | DH5α | TOP10 | BL21 |
|---|---|---|---|
| Delay before fluorescence observation | ±22 hours | ±18 hours | ±22 hours |
| Fluorescence rate with IPTG | Low | Medium | High |
| Promoter leak | Medium | Extremely high | Extremely low |
The delay before fluorescence must be analyzed with caution, as it might have been affected by the huge culture volume compared to the start amount of bacteria.
The fluorescence rate and promoter leak must be further studied in these three strains with higher concentrations of IPTG to see if we would obtain different results.
//classic/reporter/pret
//function/reporter/color
//function/reporter/pigment
| emission | RFP |
| excitation | |
| tag | None |

 1 Registry Star
1 Registry Star