Part:BBa_E2050:Experience
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_E2050
Part:BBa_E2050:Experience
Aberdeen iGEM 2010 Bio-brick Experience
The yeast mOrange open reading frame (Bio-brick E2050) was tested as part of a yeast expression construct, in which E2050 was translationally fused downstream of a tandem N-peptide open reading frame (Biobrick BBa_K385004). The gene fusion was placed under control of the GAL1 promoter (BioBrick K106699). The whole construct was cloned into a yeast shuttle vector and transformed into yeast. After growth of the transformant culture in medium containing galactose to induce the promoter, expression of mOrange was assessed using flow cytometry, (excitation wavelength 488nm, fluorescence wavelength 585nm). No mOrange fluorence was detectable by this method. However, a similar, related construct, in which the mOrange sequence was replaced by GFP, did express significant quantities of green fluorescent protein (see Aberdeen 2010 wiki for details ). This led us to conclude the mOrange sequence was non-functional in this particular expression construct.
iGEM14_Carnegie_Mellon. We characterized a set of fluorescent proteins consisting of BFP. GFP, YFP, OFP, and RFP. We calculated the signal-to-noise ratio of all the proteins in two different cell lines (MACH and Top10). mOrange (OFP) had a high signal-to-noise ratio in MACH cells and a lower signal-to-noise ratio in Top10 cells. In both cases it had a high signal-to-noise ratio relative to other fluorescent proteins. OFP was measured at (ex/em = 548nm/562nm).


SCU_China-2017
Based on the part BBa_E2050 which containing coding sequence of mOrange only, we added pTetR promotor(BBa_R0040), RBS(BBa_B0034) and double terminators(BBa_B0010 and BBa_B0012) to it.
In order to observe the fluorescence of mOrange easily and obviously, we transformed plasmid containing this part into E. coli BL21(DE3) strain which doesn’t have tetR gene. In this circumstance, mOrange can be expressed in BL21(DE3) cells constitutively.
After transformation, BL21(DE3) cells were cultured in LB solid media containing chloramphenicol for about 18 hours. We used two controls in the whole process, BL21(DE3) competent cells transformed with recombinant plasmid pCI-luxI-pSB1C3 and BL21(DE3) competent cells transformed with nothing.
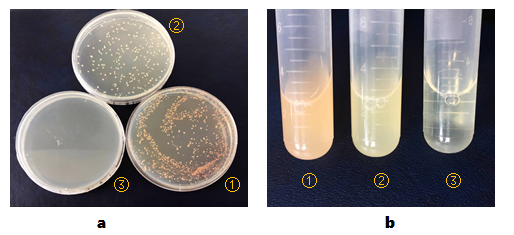
BL21(DE3) colonies transformed with part BBa_K2276009 are orange. BL21(DE3) colonies transformed with pCI-luxI-pSB1C3 are in normal color. BL21(DE3) cells without any other plasmids can’t grow on plate containing chloramphenicol(Figure 1, a).
And then, we isolated the single colonies from selective plates, and inoculated a culture of about 3 mL LB medium containing chloramphenicol. The cultures were incubated at 37℃ for about 24 hours. Culture of BL21(DE3) cells transformed with part BBa_K2276009 are orange obviously. And BL21(DE3) cells transformed with pCI-luxI-pSB1C3 are light yellow(Figure 1, b).
User Reviews
UNIQa3b7870ddafbf1f4-partinfo-00000001-QINU
|
•
Slam |
We were unable to detect mOrange fluorescence in our GAL1 regulated construct using this Bio-brick part. See above for more details. |
|
Antiquity |
This review comes from the old result system and indicates that this part did not work in some test. |
UNIQa3b7870ddafbf1f4-partinfo-00000004-QINU
User Reviews
|
•
Tianjin iGEM |
We used this part to confirm our yeast surface display system. Our result confirmed that this part is good. UNIQa3b7870ddafbf1f4-partinfo-00000006-QINU |

 1 Registry Star
1 Registry Star