Part:BBa_K525121
S-layer cspB from Corynebacterium glutamicum with TAT-Sequence and lipid anchor, PT7 and RBS
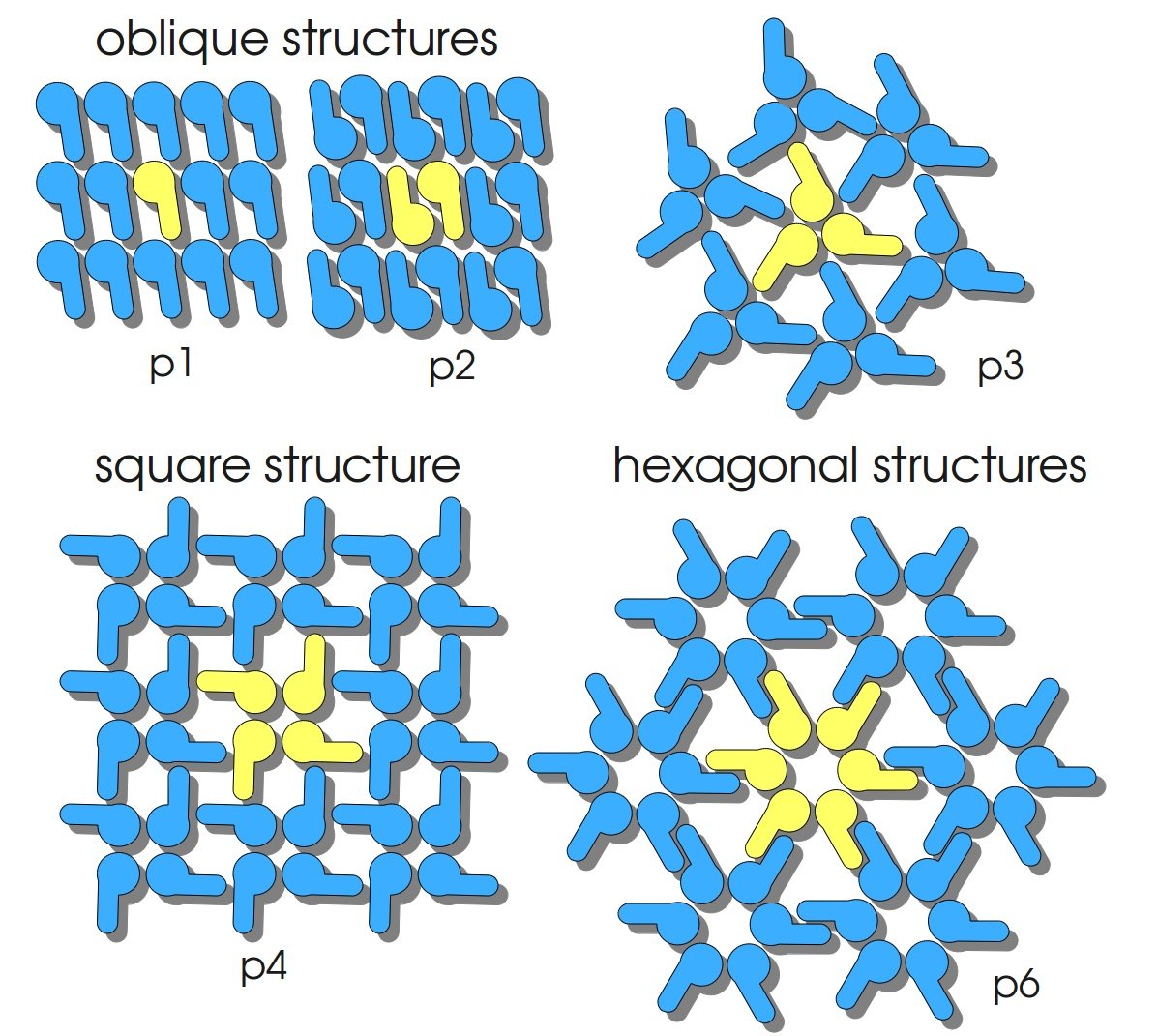
S-layers (crystalline bacterial surface layer) are crystal-like layers consisting of multiple protein monomers and can be found in various (archae-)bacteria. They constitute the outermost part of the cell wall. Especially their ability for self-assembly into distinct geometries is of scientific interest. At phase boundaries, in solutions and on a variety of surfaces they form different lattice structures. The geometry and arrangement is determined by the C-terminal self assembly-domain, which is specific for each S-layer protein. The most common lattice geometries are oblique, square and hexagonal. By modifying the characteristics of the S-layer through combination with functional groups and protein domains as well as their defined position and orientation to eachother (determined by the S-layer geometry) it is possible to realize various practical applications ([http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.2006.00573.x/full Sleytr et al., 2007]).
Usage and Biology
S-layer proteins can be used as scaffold for nanobiotechnological applications and devices by e.g. fusing the S-layer's self-assembly domain to other functional protein domains. It is possible to coat surfaces and liposomes with S-layers. A big advantage of S-layers: after expressing in E. coli and purification, the nanobiotechnological system is cell-free. This enhances the biological security of a device.
The S-layer of C. glutamicum is characterized by a hexagonal lattice symmetry. Attachment between S-layer and cell wall was found to be due to the hydrophobic carboxy-terminus of the PS2 protein [http://www.sciencedirect.com/science/article/pii/S016816560400241X Hansmeier et al., 2004]).
Important parameters
| Experiment | Characteristic | Result |
|---|---|---|
| Expression (E. coli) | ||
| Compatibility | E. coli KRX | |
| Inductor | L-rhamnose | |
| Optimal temperature | 37 °C | |
| Specific growth rate (un-/induced) | 0.260 h-1 / 0.106 h-1 | |
| Doubling time (un-/induced) | 2,67 h / 6.52 h | |
| Characterisation | ||
| Number of amino acids | 461 | |
| Molecular weight | 50.6 | |
| Theoretical pI | 4.21 | |
| Localisation | mainly cell membrane |
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 1421
Illegal XhoI site found at 248
Illegal XhoI site found at 866 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 1400
Illegal SapI site found at 647
Illegal SapI site found at 859
Illegal SapI site found at 1407
Expression in E. coli
For characterizations, the cspB gene was fused to a monomeric RFP (BBa_E1010) using Gibson assembly.
The CspB|mRFP fusion protein was overexpressed in E. coli KRX after induction of a T7 polymerase gene in the KRX's genome by supplementation of 0,1 % L-rhamnose using the autinduction protocol developed by Promega.
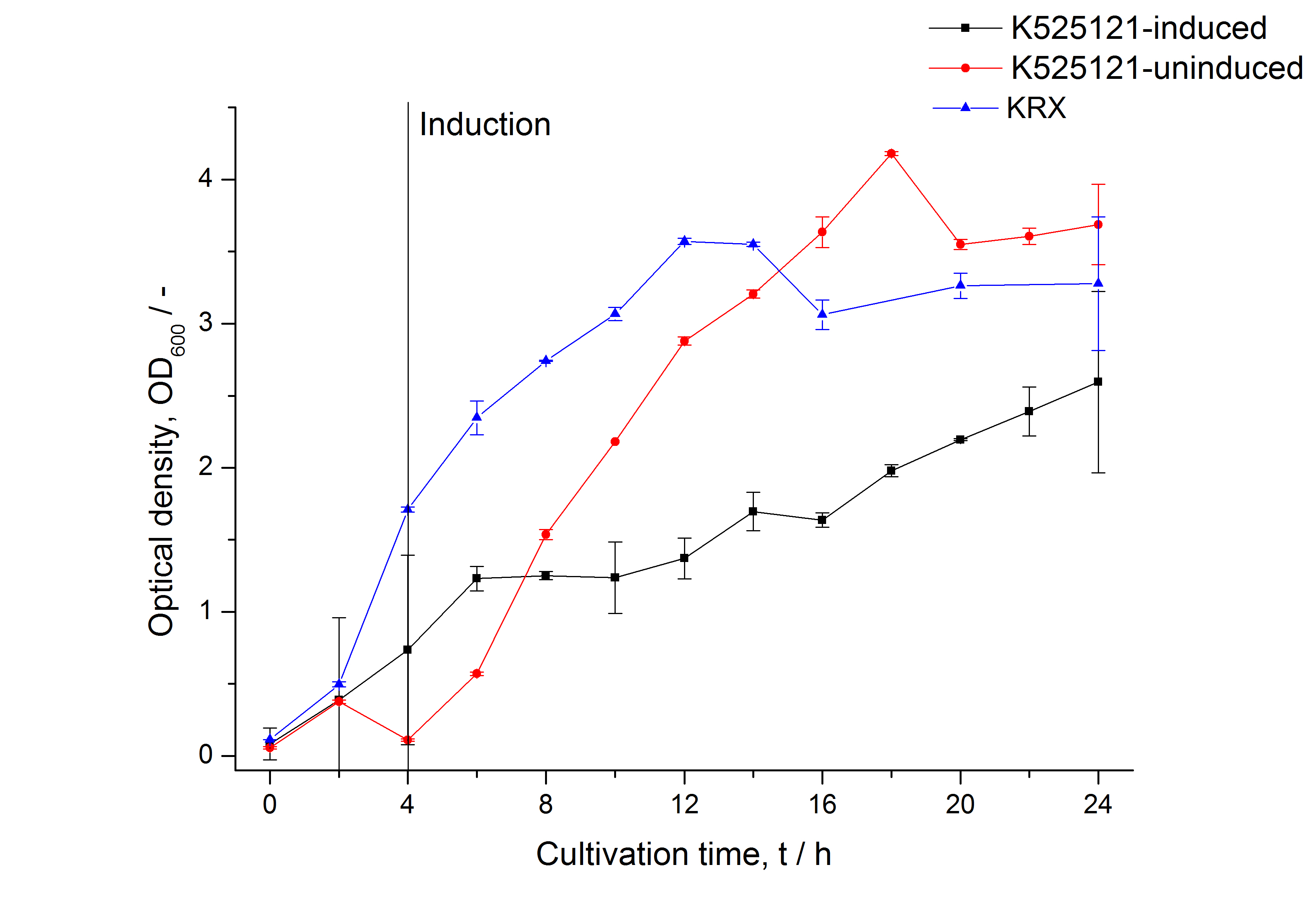
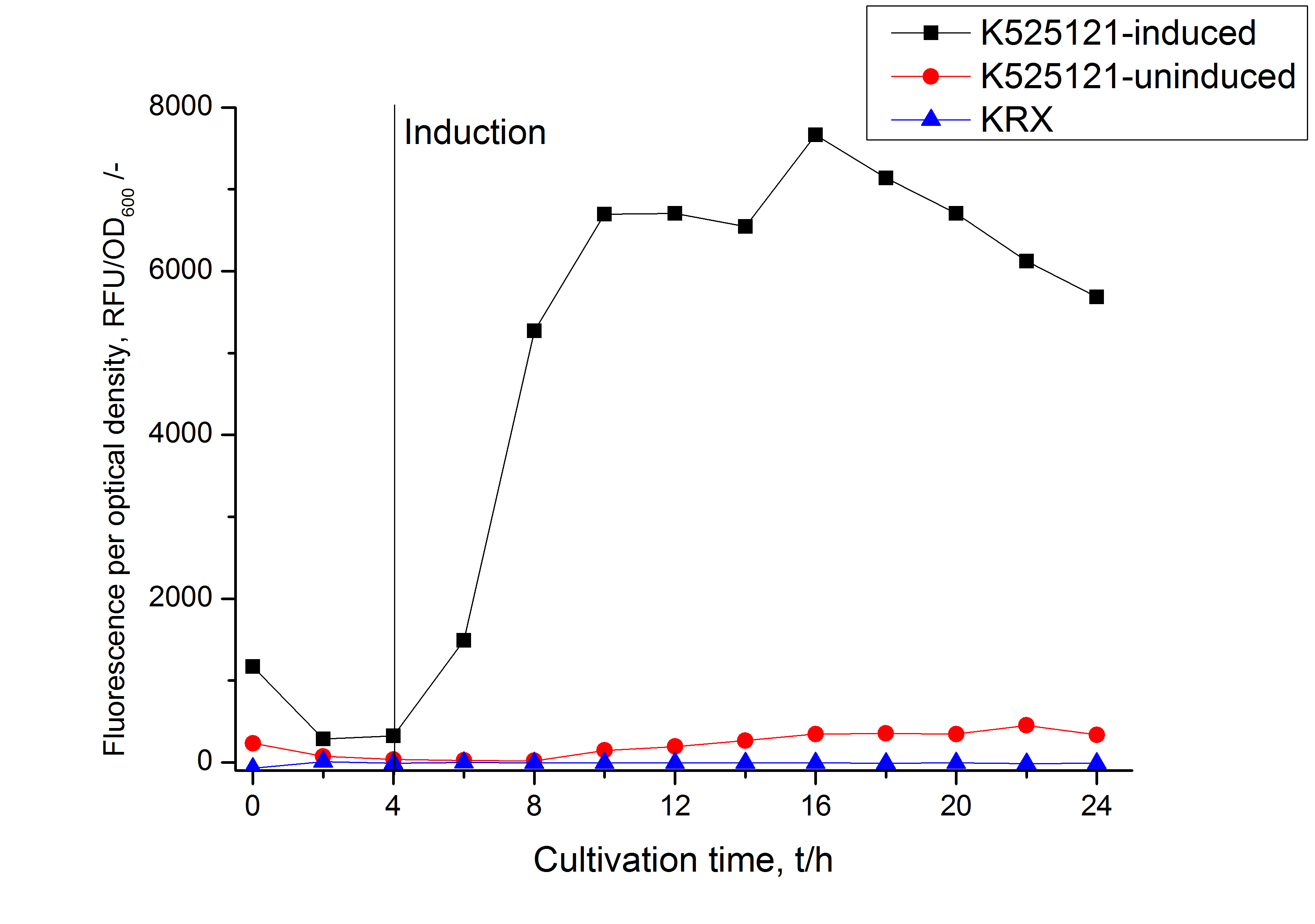
Identification and localisation
After a cultivation time of 18 h the CspB|mRFP fusion protein was localized in E. coli KRX. Therefore a part of the produced biomass was mechanically disrupted and the resulting lysate was washed with ddH2O. The periplasm was detached by using an osmotic shock from other parts of the cells. The existance of fluorescene in the periplasm fraction, showed in fig. 3, indicates that C. glutamicum TAT-signal sequence is at least in part functional in E. coli KRX.
The S-layer fusion protein could not be found in the polyacrylamide gel after a SDS-PAGE of the lysate and the cell debris were still red. This indicates that the fusion protein integrates into the cell membrane with its lipid anchor. For testing this assumption the washed lysate was treated with ionic, nonionic and zwitterionic detergents to release the CspB|mRFP out of the membranes.
The existance of flourescence in the detergent fractions and the proportionally low fluorescence in the wash fraction confirm the hypothesis of an insertion into the cell membrane (fig. 3). An insertion of these S-layer proteins might stabilize the membrane structure and increase the stability of cells against mechanical and chemical treatment. A stabilization of E. coli expressing S-layer proteins was discribed by Lederer et al., (2010).
An other important fact is, that there is actually mRFP fluorescence measurable in such high concentrated detergent solutions. The S-layer seems to stabilize the biologically active conformation of mRFP. The MALDI-TOF analysis of the relevant size range in the polyacrylamid gel approved the existance of the intact fusion protein in all detergent fractions (fig 4).
MALDI-TOF analysis was first used to identify the location of the fusion protein in different fractions. Fractions of medium supernatant after cultivation, periplasmatic isolation, cell lysis, denaturation in 6 M urea and the following wash with 2 % (v/v) Triton X-100, 2 % SDS (w/v) were loaded onto a SDS-PAGE and fragments of the gel were measured with MALDI TOF.
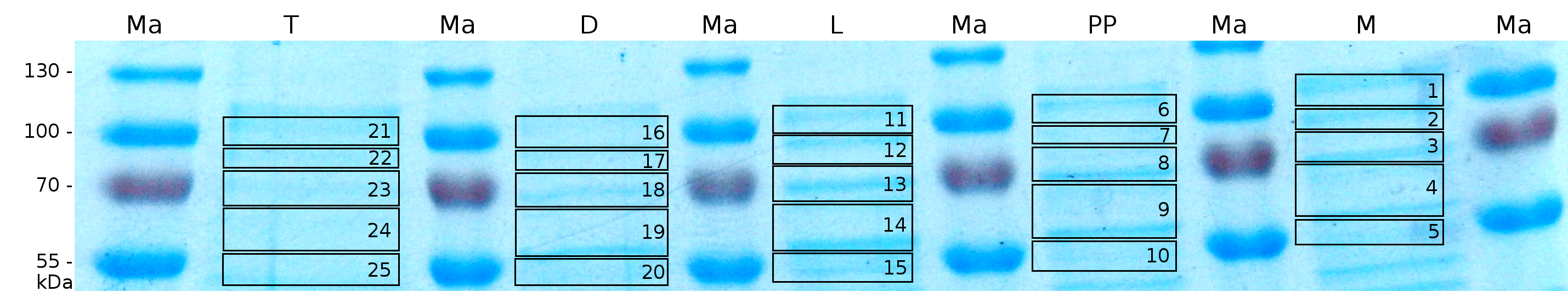
The following table shows the sequence coverage (in %) of our measurable gel samples with the amino acid sequence of fusion protein CspB/mRFP (BBa_E1010).
| number of gel sample | sequence coverage (%) |
|---|---|
| 1 | 1.9 |
| 2 | 11.5 |
| 3 | 8.0 |
| 4 | 2.6 |
| 5 | 0.0 |
| 6 | 0.0 |
| 7 | 2.6 |
| 8 | 0.0 |
| 9 | 0.0 |
| 10 | 0.0 |
| 11 | 9.4 |
| 12 | 2.6 |
| 13 | 2.8 |
| 14 | 0.0 |
| 15 | 0.0 |
| 16 | 0.0 |
| 17 | 8.0 |
| 18 | 0.0 |
| 19 | 0.0 |
| 20 | 0.0 |
| 21 | 12.2 |
| 22 | 12.2 |
| 23 | 0.0 |
| 24 | 0.0 |
| 25 | 0.0 |
Fig. 5 shows these data. The gel samples were arranged after estimated molecular mass cut out from the gel.
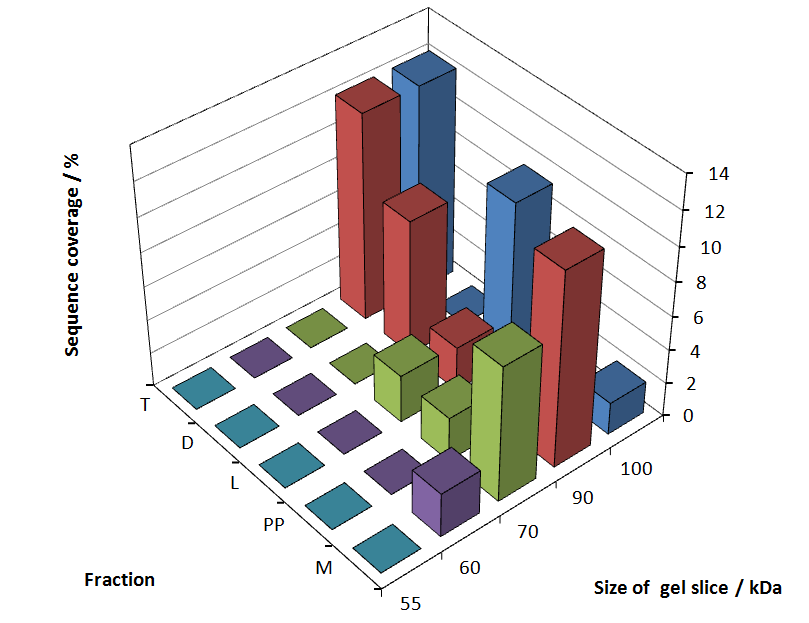
As expected, only minor sequence coverage was found in the periplasmatic fraction, due to the lipid anchor located at the carboxy-terminus. This hydrophobic region inhibits the transport of the protein to the periplasm, mediated by the amino-terminal TAT-sequence. Little fluorescence was also found in the lysis fraction, verifying our assumtion, that the protein integrates or strongly binds to the cell membrane. Using urea to disintegrate the S-layer fusion protein from the cell membrane resulted only in a slightly higher sequence coverage. However, washing the pellet with 2 % Triton X-100 (v/v), 2 % SDS (w/v), previously treated with urea, resulted in a higher sequence coverage and can therefore be expected as more applicable to desintegrate the S-layer fusion protein. Sequence coverage in the supernatant of the cultivation medium can be explained with the late phase of cultivation where some cells are lysed.
To obtain more specific informations about the location of the S-layer fusion protein, after comparison with same treated fraction of E. coli KRX all gel bands in a defined size area were cut out of the gel and analysed with MALDI-TOF. Results are shown in fig. 7.
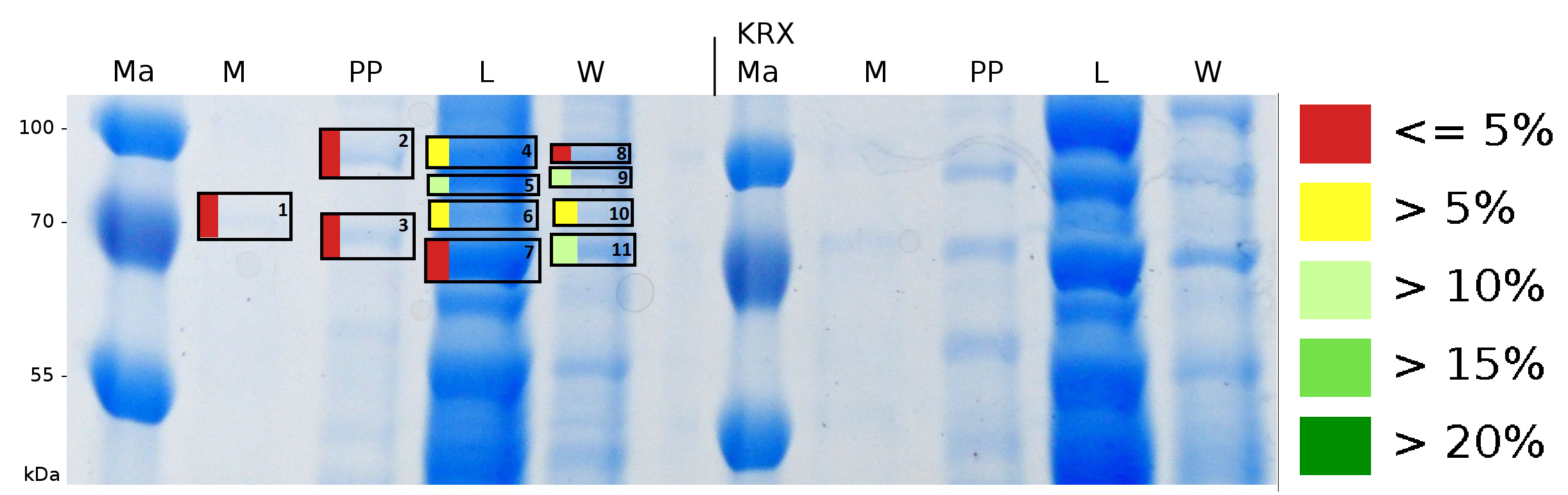
Sequence coverage was only found in the wash and the lysis fraction, again indicating that the S-layer protein is integrating in the cell membrane. Thus transport to the periplasm mediated through the amino-terminal TAT-sequence cannot take place or after transport to the periplasm binds to the inner cell membrane.
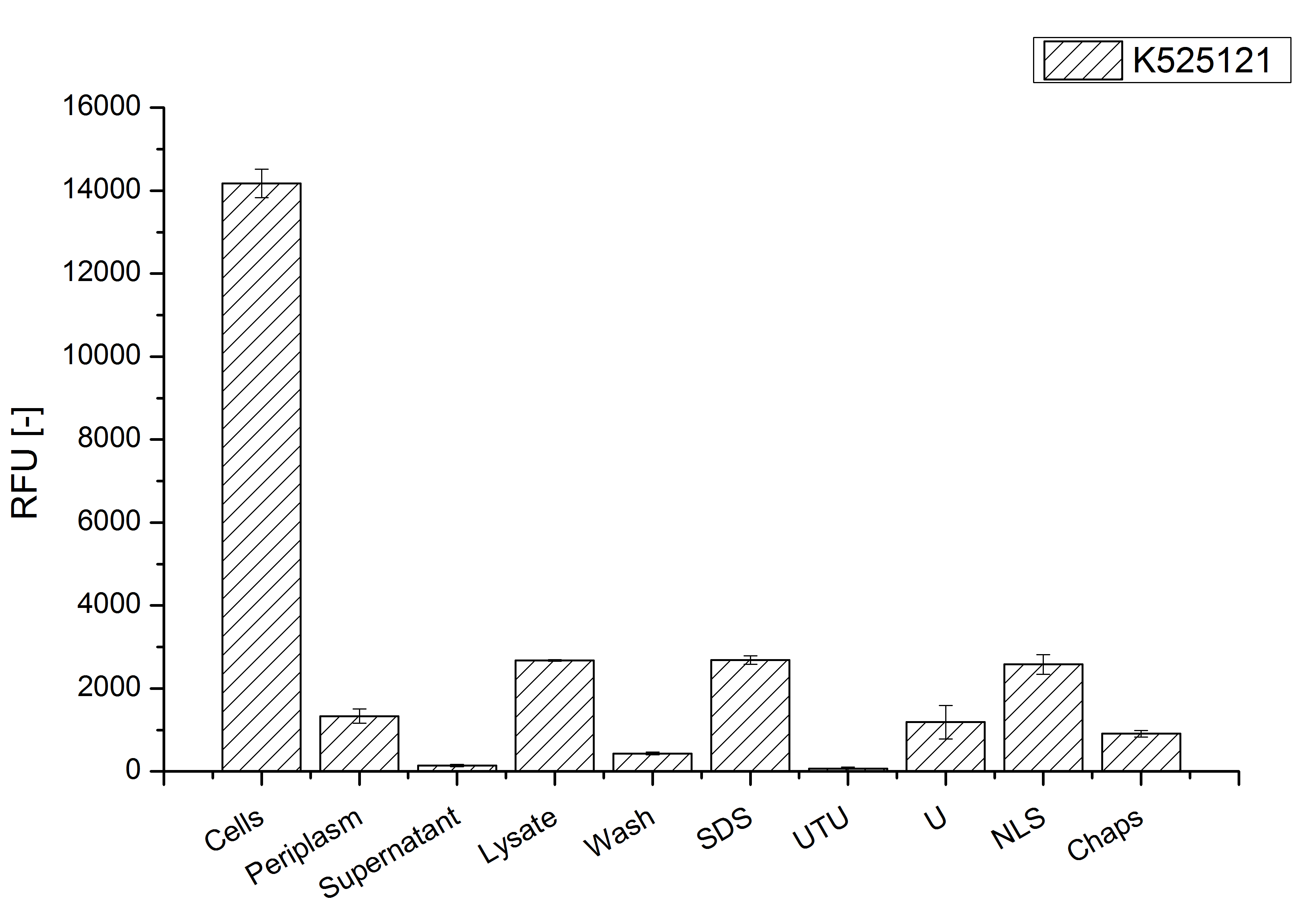
The influence of other detergents to disintegrate the S-layer fusion protein was tested after disrupting the cells with a ribolyser. The cell pellet was incubated in 10 % (v/v) Sodium dodecyl sulfate (SDS), in 7 M urea and 3 M thiourea (UTU), in 10 M urea (U) in 10 % (v/v) n-lauroyl sarcosine (NLS) and in 2 % CHAPS (C). Samples of the incubations with these detergents were loaded onto a SDS-PAGE prior to measurement with MALDI TOF.
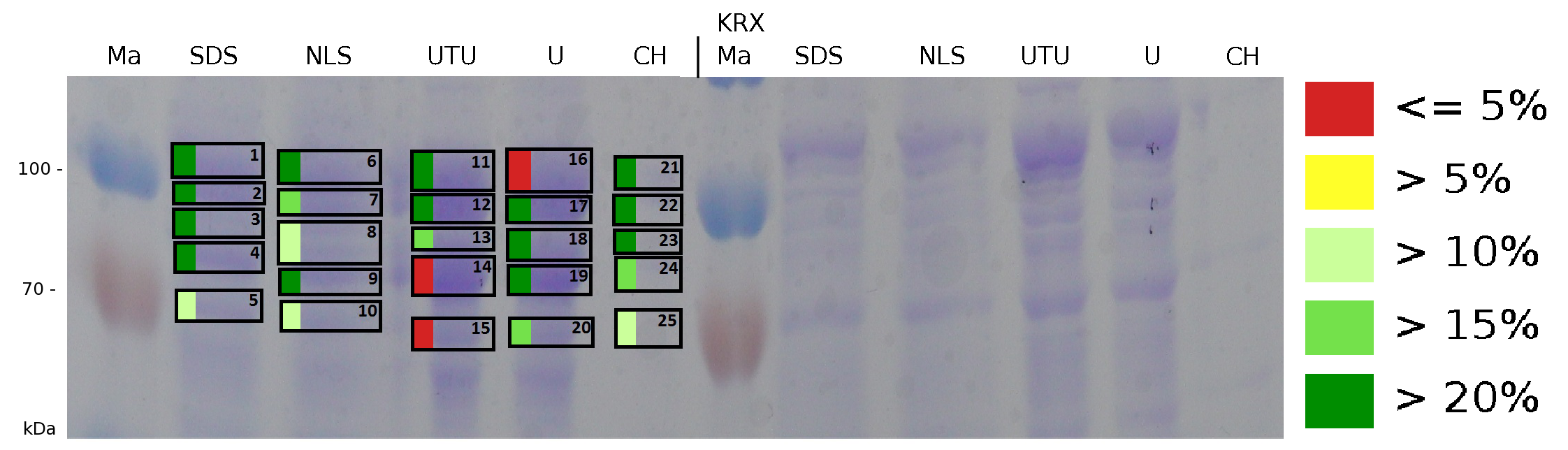
The result of the MALDI-TOF measurement clearly demonstrates that all used detergents are applicable to disintegrate the S-layer fusion proteins from the bacterial cell membrane of E. coli. Fluorescence measurement of fractions, treated with the detergents, show significantly different values, indicating that some of the detergents (e.g. 3 M thiourea, 7 M urea) have a strong effect on protein folding.
//proteindomain/internal
//rnap/bacteriophage/t7
| None |