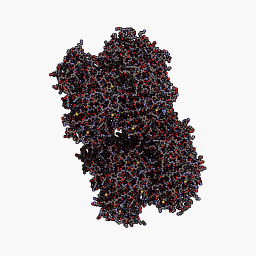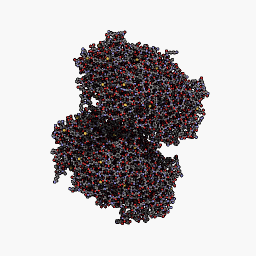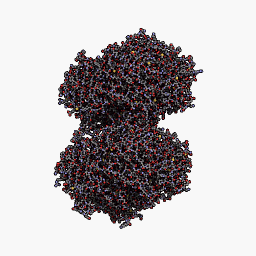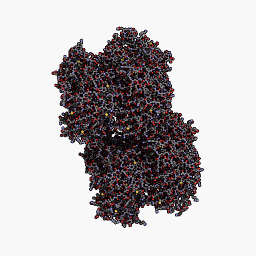Difference between revisions of "Part:BBa K592011"
| Line 417: | Line 417: | ||
<p>2. Benkert P, Biasini M, Schwede T. (2011). "Toward the estimation of the absolute quality of individual protein structure models." Bioinformatics, 27(3):343-50.</p> | <p>2. Benkert P, Biasini M, Schwede T. (2011). "Toward the estimation of the absolute quality of individual protein structure models." Bioinformatics, 27(3):343-50.</p> | ||
<p>3. Arnold K., Bordoli L., Kopp J., and Schwede T. (2006). The SWISS-MODEL Workspace: A web-based environment for protein structure homology modelling. Bioinformatics, 22,195-201.</p> | <p>3. Arnold K., Bordoli L., Kopp J., and Schwede T. (2006). The SWISS-MODEL Workspace: A web-based environment for protein structure homology modelling. Bioinformatics, 22,195-201.</p> | ||
| + | |||
| + | ===Newcastle 2019 Characterisation=== | ||
Revision as of 09:47, 21 October 2019
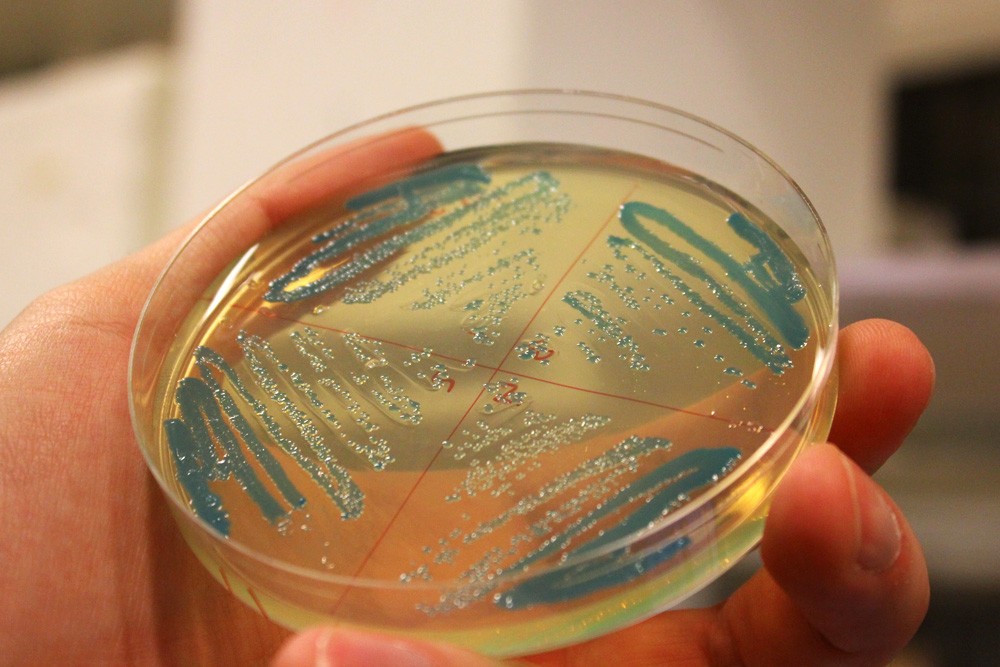
cjBlue, green chromoprotein
This chromoprotein from the Cnidopus japonicus sea anemone, cjBlue, naturally exhibits dark green color when expressed. Compared to some other chromoproteins, such as amilCP (BBa_K592009), amilGFP (BBa_K592010), spisPink (BBa_K1033932), asPink (BBa_K1033933) and aeBlue (BBa_K864401), the color development is slower. The color is readily observed in both LB or on agar plates after 24-48 hours of incubation. The sequence is codon optimized for expression in E coli.
Usage and Biology
This part is useful as a reporter.
iGEM11_Uppsala-Sweden: Expression of green chromoprotein. Escherichia coli constitutively expressing cjBlue (BBa_K592011).
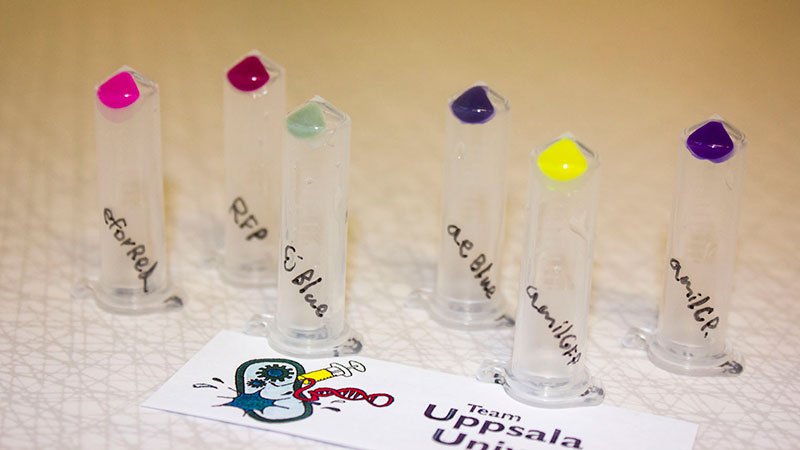
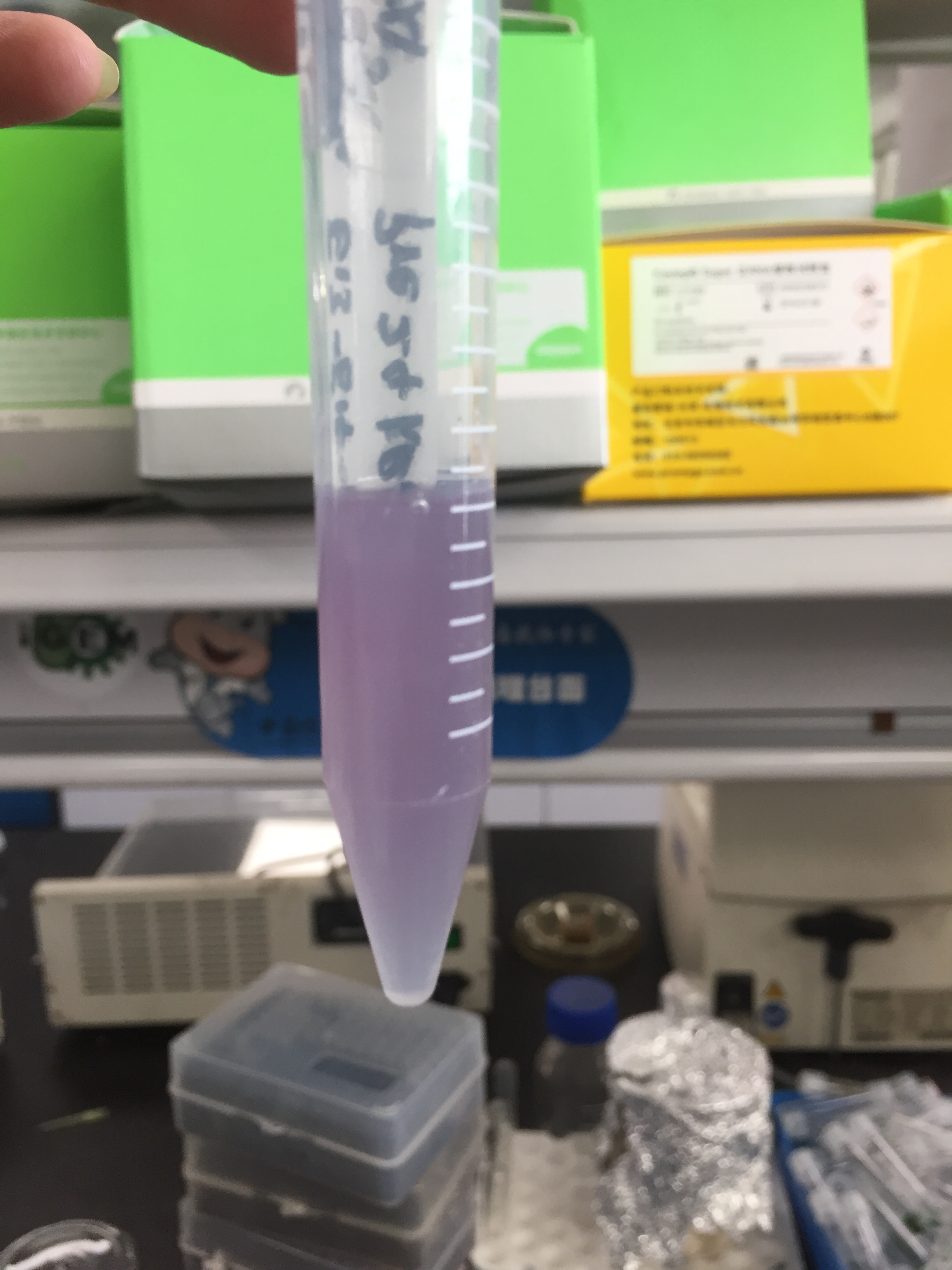
iGEM12_Uppsala_University: The Uppsala chromoprotein collection and RFP. The image shows pellets of E coli expressing chromoproteins eforRed BBa_K592012, RFP BBa_E1010, cjBlue BBa_K592011, aeBlue BBa_K864401, amilGFP BBa_K592010 and amilCP BBa_K592009.
iGEM17_SJTU_BioX_Shanghai:
J23119+Target3+cjBlue BBa_K2285014
J23119+Target1+cjBlue BBa_K2285018
An improved part has been constructed. Since this part is a coding sequence, we added a RBS which is on the upstream of sfGFP BBa_K515005 and terminator BBa_B1006 by PCR. What's more, our device have a stem-loop structure(called Target) following the constitutive promoter J23119, to achieve further control of cjBlue expression.
References
[http://www.jbc.org/content/281/49/37813.abstract] Chan MC, Karasawa S, Mizuno H, Bosanac I, Ho D, Privé GG, Miyawaki A, Ikura M. (2006) Structural characterization of a blue chromoprotein and its yellow mutant from the sea anemone Cnidopus japonicus. J. Biol. Chem. 281(49). 37813-9
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Structure and SWISS MODEL Homology Report
(The following information has been contributed by SVCE_Chennai 2016)
The following sections give a detailed information on this chromoprotein done by its in silico analysis.
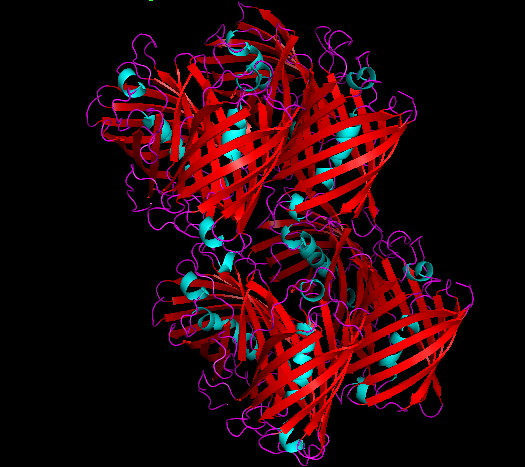
3D Structure
Cartoon representation of cjBlue chromoprotein.
The best match during structure prediction.
| Template | Seq. Identity | Oligo-state | Found by | Method | Resolution | Seq Similarity | Range | Coverage | Description |
|---|---|---|---|---|---|---|---|---|---|
| 2ib5.1.A | 99.57 | homo-octamer | BLAST | X-Ray | 1.80Å | 0.63 | 5 - 232 | 0.99 | Chromo protein |
Ligands -None
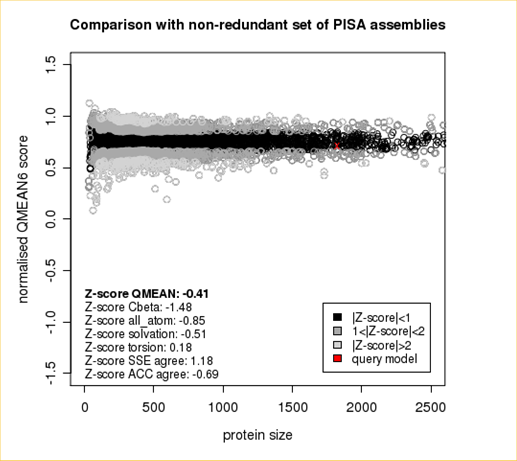
Plot 1
Plot 2
Plot 3
Plot 1
The area built by the circles colored in different shades of grey in the plot on the left hand side represent the QMEAN scores of the reference structures from the PDB. The model's QMEAN score is compared to the scores obtain for experimental structures of similar size (model size +/- 10%) and a Z-score is calculated. A Z-score (or standard score) is a score which is normalised to mean 0 and standard deviation 1. Thus the QMEAN Z-score directly indicates how many standard deviations the model's QMEAN score differs from expected values for experimental structures. In analogy, Z-scores are calculated for all four statistical potential terms as well as the agreement terms being part of the QMEAN score.
Plot 2
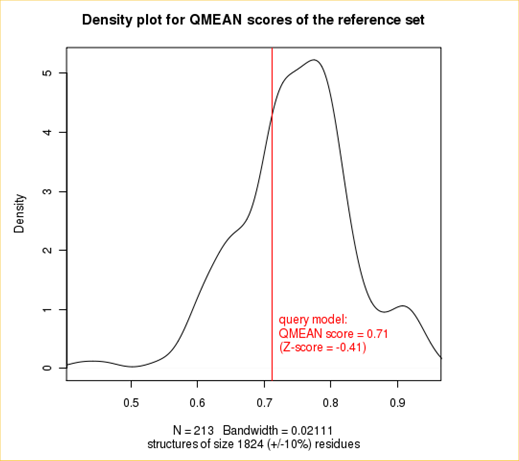
The plot in the middle shows the density plot (based on the QMEAN score) of all reference models used in the Z-score calculation. The location of the query model with respect to the background distribution is marked in red. This plot basically is a "projection" of the first plot for the given protein size. The number of reference models used in the calculation is shown at the bottom of the plot.
Plot 3
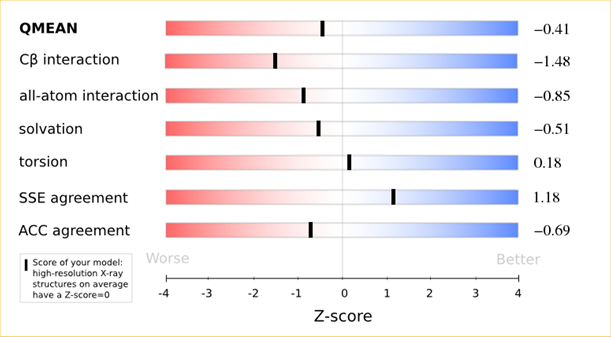
The analysis of these Z-scores of the individual terms can help identifying the geometrical features responsible for an observed large negative QMEAN Z-score. Models of low quality are expected to have strongly negative Z-scores for QMEAN but also for most of the contributing terms. Large negative values correspond to red regions in the color gradient. "Good structures" are expected to have all sliders in the light red to blue region.”
The QMEAN6 score is a composite score consisting of a linear combination of 6 terms (estimated model reliability between 0-1). The pseudo-energies of the contributing terms are given below together with their Z-scores with respect to scores obtained for high-resolution experimental structures of similar size solved by X-ray crystallography:
| Scoring Function Term | Raw Score | Z Score |
|---|---|---|
| Cβ Interaction Energy | -91.57 | -1.48 |
| All Atom Pairwise Energy | -42903.03 | -0.85 |
| Solvation Energy | -145.58 | -0.51 |
| Torsion Angle Energy | -501.77 | 0.18 |
| Secondary Structure Agreement | 88.2% | 1.18 |
| Solvent Accessibility Agreement | 76.0% | -0.69 |
| QMEAN6 score | 0.712 | -0.41 |
Ramachandran Plot- SWISS MODEL Workspace
(The following information has been contributed by SVCE_Chennai 2016)
A score of 94% in the most favoured regions as seen in the Ramachandran plot shows that the above predicted model of cjBlue, green chromoprotein is a very good prediction.
ProtParam Results
(The following information has been contributed by SVCE_Chennai 2016)
Number of amino acids: 232
Molecular weight: 26173.80
Theoretical pI: 7.53
Total number of negatively charged residues (Asp + Glu): 24
Total number of positively charged residues (Arg + Lys): 25
Atomic composition:
Carbon (C) - 1151 Hydrogen (H) - 1776 Nitrogen (N) - 314 Oxygen (O) - 347 Sulfur (S) - 19
Formula: C1151H1776N314O347S19
Total number of atoms: 3607
Extinction coefficients:
Extinction coefficients are in units of M-1 cm-1, at 280 nm measured in water.
Ext. coefficient : 28015
Abs 0.1% (=1 g/l): 1.070, assuming all pairs of Cys residues form cystines
Ext. coefficient : 27390
Abs 0.1% (=1 g/l): 1.046, assuming all Cys residues are reduced
Estimated half-life:
The N-terminal of the sequence considered is M (Met).
The estimated half-life is: 30 hours (mammalian reticulocytes, in vitro).
>20 hours (yeast, in vivo).
>10 hours (Escherichia coli, in vivo).
Instability index:
The instability index (II) is computed to be 29.86.
This classifies the protein as stable.
Aliphatic index: 58.84
Grand average of hydropathicity (GRAVY): -0.558
ExPASy Peptide Cutter Results
(The following information has been contributed by SVCE_Chennai iGEM-2016)
The following enzymes cleave the chromoprotein.
| Name of enzyme | No. of cleavages | Positions of cleavage sites |
|---|---|---|
| Arg-C proteinase | 8 | 10 92 147 150 175 179 198 220 |
| Asp-N endopeptidase | 8 | 6 55 77 97 110 150 165 195 |
| Asp-N endopeptidase + N-terminal Glu | 24 | 6 15 26 28 30 35 40 55 77 85 90 96 97 106 110 140 144 150 165 195 199 205 211 214 |
| BNPS-Skatole | 2 | 90 140 |
| CNBr | 8 | 1 15 25 40 133 146 160 189 |
| Chymotrypsin-high specificity (C-term to [FYW], not before P) | 21 | 14 24 35 53 55 64 72 79 80 88 90 96 101 117 140 148 178 192 194 211 221 |
| Chymotrypsin-low specificity (C-term to [FYWML], not before P) | 50 | 1 13 14 15 22 23 24 25 35 40 53 55 58 64 72 79 80 88 90 96 101 102 110 115 117 122 133 140 146 148 154 159 160 162 169 170 173 174 178 187 192 193 194 197 210 211 214 221 229 231 |
| Clostripain | 8 | 10 92 147 150 175 179 198 220 |
| Formic acid | 8 | 7 56 78 98 111 151 166 196 |
| Glutamyl endopeptidase | 16 | 16 27 29 31 36 41 86 91 97 107 141 145 200 206 212 215 |
| Hydroxylamine | 2 | 129 207 |
| Iodosobenzoic acid | 2 | 90 140 |
| LysC | 17 | 4 12 33 44 47 71 81 109 120 135 136 163 181 182 185 203 228 |
| LysN | 17 | 3 11 32 43 46 70 80 108 119 134 135 162 180 181 184 202 227 |
| NTCB (2-nitro-5-thiocyanobenzoic acid) | 11 | 25 61 113 123 142 143 154 163 171 221 224 |
| Pepsin (pH1.3) | 36 | 13 23 52 54 55 57 79 80 84 87 88 101 102 109 110 114 115 121 153 154 158 159 161 162 169 170 173 174 186 193 194 209 210 211 228 229 |
| Pepsin (pH>2) | 52 | 13 23 34 52 54 55 57 63 64 71 72 79 80 84 87 88 89 90 95 96 100 101 102 109 110 114 115 116 117 121 139 148 153 154 158 159 161 162 169 170 173 174 178 186 193 194 209 210 211 221 228 229 |
| Proline-endopeptidase | 2 | 34 199 |
| Proteinase K | 104 | 2 5 9 11 13 14 16 18 19 24 27 28 29 31 35 36 38 41 43 45 46 51 53 55 57 58 59 64 67 68 69 70 72 73 76 79 80 84 86 88 89 90 91 93 94 95 96 97 100 101 102 103 104 107 108 110 115 116 117 119 121 122 126 132 137 140 141 145 148 149 153 154 158 159 161 162 165 170 171 174 176 177 178 183 184 186 187 192 194 200 201 202 204 206 209 210 211 212 215 218 219 221 224 229 |
| Staphylococcal peptidase I | 16 | 16 27 29 31 36 41 86 91 97 107 141 145 200 206 212 215 |
| Thermolysin | 53 | 1 4 8 10 12 14 18 23 24 39 42 44 52 54 57 66 67 68 79 87 94 99 101 109 114 115 118 120 121 131 132 152 153 158 159 160 161 164 169 173 182 183 185 186 191 193 201 203 209 210 217 218 228 |
| Trypsin | 23 | 4 10 12 44 47 71 81 92 109 120 135 136 147 150 163 175 179 181 182 185 203 220 228 |
These chosen enzymes do not cut:
Caspase1
Caspase10
Caspase2
Caspase3
Caspase4
Caspase5
Caspase6
Caspase7
Caspase8
Caspase9
Enterokinase
Factor Xa
GranzymeB
Thrombin
Tobacco etch virus protease
References
1. Benkert, P., Schwede, T. and Tosatto, S.C.E. (2009). QMEANclust: Estimation of protein model quality by # combining a composite scoring function with structural density information. BMC Struct Biol. 2009 May 20;9:35. If you publish results using DSSP, please cite the following paper: Dictionary of protein secondary structure: Pattern recognition of hydrogen bonded and geometrical features, Biopolymers 22:2577-2637
2. Benkert P, Biasini M, Schwede T. (2011). "Toward the estimation of the absolute quality of individual protein structure models." Bioinformatics, 27(3):343-50.
3. Arnold K., Bordoli L., Kopp J., and Schwede T. (2006). The SWISS-MODEL Workspace: A web-based environment for protein structure homology modelling. Bioinformatics, 22,195-201.
===Newcastle 2019 Characterisation===