Difference between revisions of "Part:BBa K3182100"
| Line 26: | Line 26: | ||
[[File:T--Linkoping_Sweden--pinkwhite1.jpeg|300px|thumb|left|<b>Figure 2.</b> Black arrow indicates a unsuccessful ligation where the colony turns pink, the white colonies was successfully ligated and the pink color was removed. Host in this picture was DH5-alpha and were incubated for 48 hours in 37 degrees Celsius.]] | [[File:T--Linkoping_Sweden--pinkwhite1.jpeg|300px|thumb|left|<b>Figure 2.</b> Black arrow indicates a unsuccessful ligation where the colony turns pink, the white colonies was successfully ligated and the pink color was removed. Host in this picture was DH5-alpha and were incubated for 48 hours in 37 degrees Celsius.]] | ||
| − | [[File:T--Linkoping_Sweden--pinkwhitescreeningphoto.jpeg|560px|thumb|right|<b>Figure 2.</b> | + | [[File:T--Linkoping_Sweden--pinkwhitescreeningphoto.jpeg|560px|thumb|right|<b>Figure 2.</b> DH5-alpha cells used for pink-white screening. BBa K3182100 was cut with BamHI and PstI to remove pCons-AsPink and BBa K3182006 (magainin 2) was ligated into the plasmid. The white colonies indicate a successfull ligation (or vector re-ligation). All colonies were later colony screened with PCR amplification of the insert and the ampified strand was run on an agarose gel. The gel implied that all screened colonies was successfull, i.e. contained BBa K3182100 with magainin 2 instead of pCons-AsPink. ]] |
<br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br> | <br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br> | ||
Revision as of 19:20, 23 July 2019
pT7-CBDcipA-pCons-AsPink
A cellulose binding domain (CBDcipA) from Clostridium thermocellum Cellulose scaffolding protein (CipA) which can be used to purify or attach proteins to cellulose, this part has a sfGFP fused to the CBDcipA. The part also has a flexible GS-linker (-GGGGSGGGGS-) with a thrombin site (-LVPRGS-, thrombin RS) added at the end, clevage with thrombin will add one glycine and one serine to the N-terminal of the C-terminal fusion protein of the CBDcipA. Also added a BamHI recognition sequence (BamHI RS) to enable changeable fusion protein to the CBDcipA. BamHI was chosen because its RS codes for one glycine and one serine, fitting it to the end of the thrombin site, BamHI can also be used with methylated DNA.
This part utilizes a pCons-AsPink dropout enabling colour-screening for positive colonies. Using BamHI and PstI or SpeI on both this part assembled in pSB1C3 and the insert of choice will yield a fusion protein between CBDcipA and the insert (if designed to correctly). The fusion protein can later be cleaved with thrombin to yield two separate proteins. The C-terminal fusion will have one glycine and one serine added to the N-terminal of the protein.
For usage and information of CBDcipA please see: BBa_K3182000 and BBa_K3182108.
This method was used to assemble: .
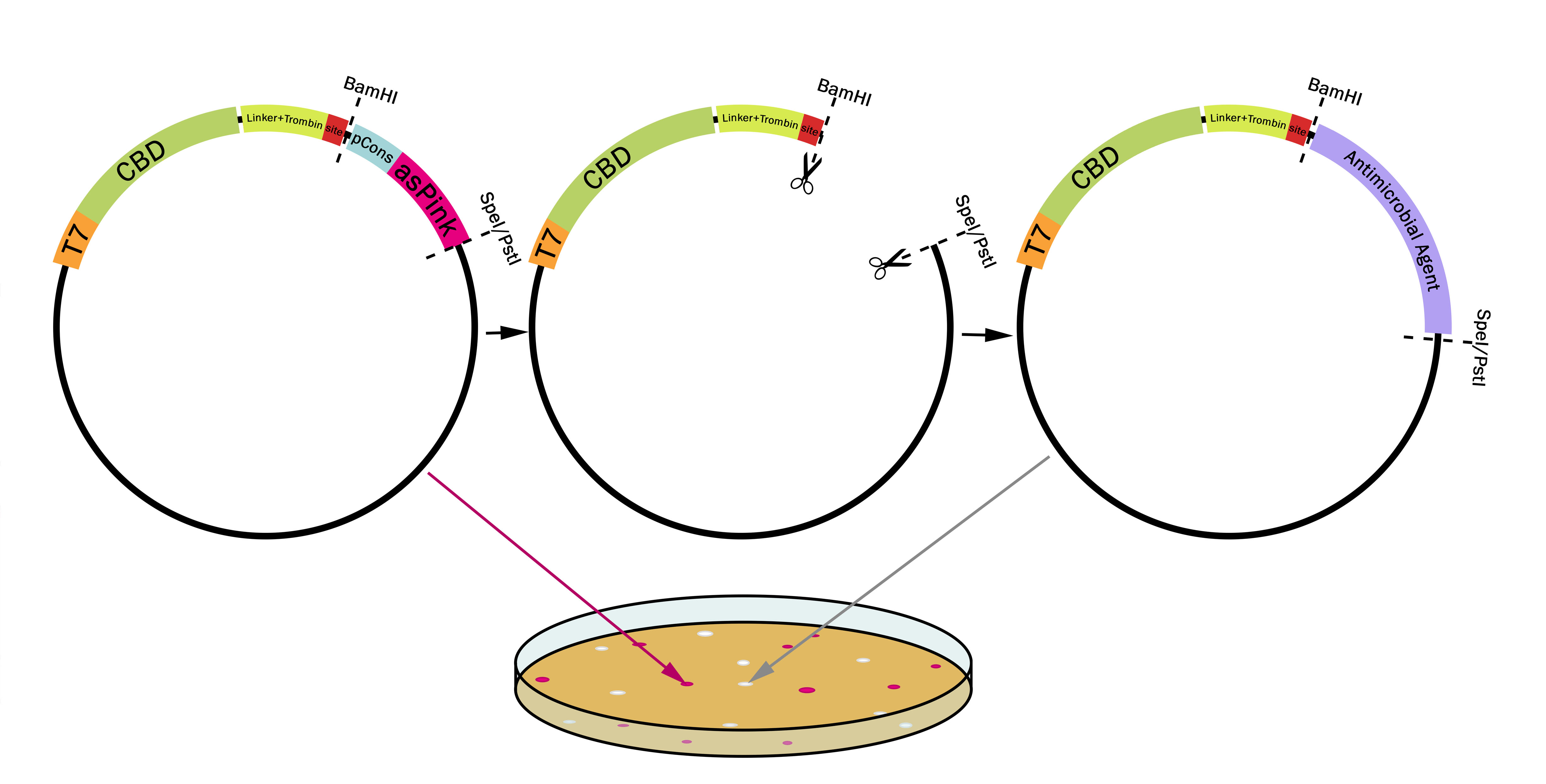
Expression system
The part has a very strong expression with a T7 promotor (BBa_I719005) as well as a 5'-UTR (BBa_K1758100) region which has been shown to further increase expression in E. coli (BBa_K1758106), ([http://www.ncbi.nlm.nih.gov/pubmed/2676996 Olins et al. 1989]), ([http://www.ncbi.nlm.nih.gov/pubmed/23927491 Takahashi et al. 2013]). Both this part and the part were sfGFP was changed for AsPink (BBa_K3182000) showed great expression.
Usage and Biology
Design any gene which you want to be fused to the CBDcipA with a BamHI recognition sequence in the 5'-end. The biobrick suffix can be used in the 3'-end. Cut the vector and insert with BamHI and PstI (SpeI also works), remove enzymes and mix, no need for gel purification. Transform the host (BL21 (DE3) for quickest results) and incubate at 37 degrees Celsius overnight, if the color is weak or can't be seen, incubate in 24-37 degrees for an additional 16-24 hours.

INSERT AGAROSE GEL HERE
Incubating the bacteria, first in 37 degrees Celsius for 16 hours and then 16 degrees for 24 hours gave a very weak color. A protein production host, such as BL21 (DE3) gave the best results, where color development could be seen after the first incubation (toxic fusions should be assembled in hosts without the T7-RNA-polymerase).
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 592
Illegal NheI site found at 615 - 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 580
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]


