Difference between revisions of "Part:BBa E1010"
Tochingyuet (Talk | contribs) |
|||
| Line 18: | Line 18: | ||
Peking iGEM 2016 has fused this part with triple spytag. The fused protein is participate in Peking’s polymer network. By adding this protein, the whole polymer network become visible in most conditions. If you want to learn more about Peking’s polymer network and the role of mRFP in this network, please click here https://parts.igem.org/Part:BBa_K1989004". | Peking iGEM 2016 has fused this part with triple spytag. The fused protein is participate in Peking’s polymer network. By adding this protein, the whole polymer network become visible in most conditions. If you want to learn more about Peking’s polymer network and the role of mRFP in this network, please click here https://parts.igem.org/Part:BBa_K1989004". | ||
| + | |||
| + | |||
| + | ===Contribution=== | ||
| + | |||
| + | <partinfo>BBa_E1010 AddReview 5</partinfo> | ||
| + | <I>Hong Kong-CUHK iGEM 2017</I> | ||
| + | |width='60%' valign='top'| | ||
| + | |||
| + | <b>Charaterization of mRFP pH stabillity</b> | ||
| + | <p>We grew C41 bacteria with parts BBa_J61002 in 2XYT for 24 hours. Purifying the mRFP by Ion Exchange Chromatography and Hydrophobic Interaction Chromatography, we measured the fluoresece (ex ,em ) of purified mRFP. which is diluted to 10µg/100µl (total 200µl) in triplicates, into different buffers (ranges from pH2 to pH12). The result shows that the stability drops dramatically in pH condition below 6 and relatively stable in pH 6-10. </p> | ||
| + | <div align="center"> | ||
| + | |||
| + | [[File:Mrfp.PNG|center|thumb|350px|''<b>Fig.1</b> Vary pH attributed to different fluorescent intensity of RFP.]] | ||
| + | <table cellpadding="2" border="1px" cellspacing="0" align="center" width="70%"> | ||
| + | <caption><p align="justify"><b>Table 1</b> Plate reader setting of fluorescent measurement</p></caption> | ||
| + | <td><b>Measurement Type</b></td> | ||
| + | <td>Fluorescence</td> | ||
| + | |||
| + | <tr> | ||
| + | <td><b>Microplate name<</b>/td> | ||
| + | <td>COSTAR 96</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td><b>Scan mode</b></td> | ||
| + | <td>orbital averaging</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td><b>Scan diameter [nm]</b></td> | ||
| + | <td>3</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td><b>Excitation</b></td> | ||
| + | <td>550-20</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td><b>Emission</b></td> | ||
| + | <td>605-40</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td><b>Dichronic filter</b> </td> | ||
| + | <td>auto 572.5</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td><b>Gain </b></td> | ||
| + | <td>500</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td><b>Focal height [nm]</b></td> | ||
| + | <td>9</td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | |||
| + | |}; | ||
| + | |||
| + | {|width='80%' style='border:1px solid gray' | ||
| + | |- | ||
| + | |width='10%'| | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
Revision as of 09:32, 29 October 2017
**highly** engineered mutant of red fluorescent protein from Discosoma striata (coral)
monomeric RFP: Red Fluorescent Protein. Excitation peak: 584 nm Emission peak: 607 nm
Usage and Biology
Robert E. Campbell started with Discosoma RFP (DsRed) and evolved a faster folding, monomeric variant. See paper listed in source. Codon optimized for expression in bacteria (?? DE)
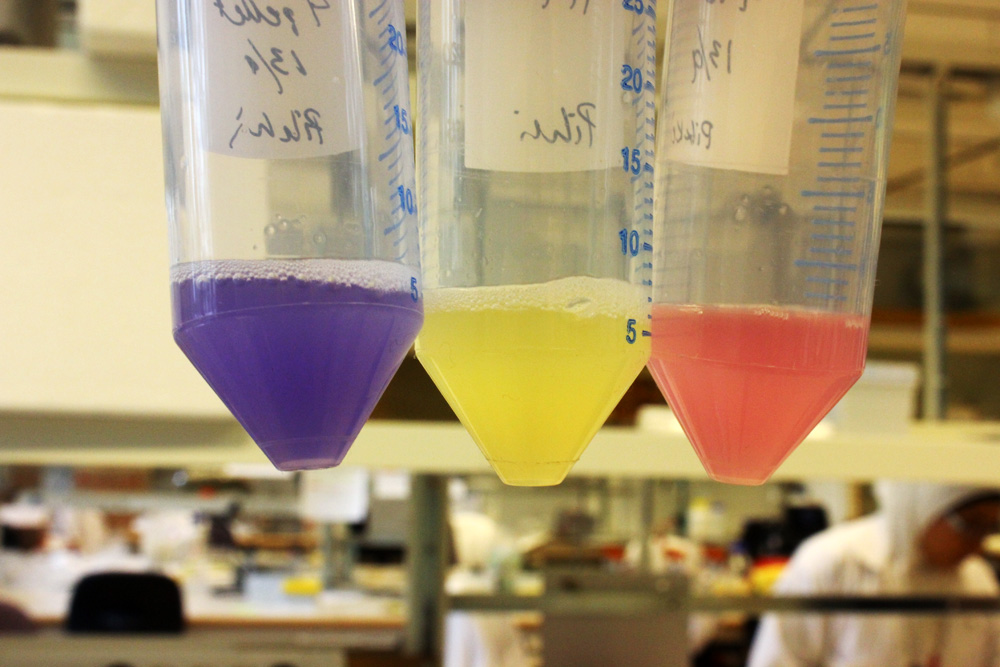
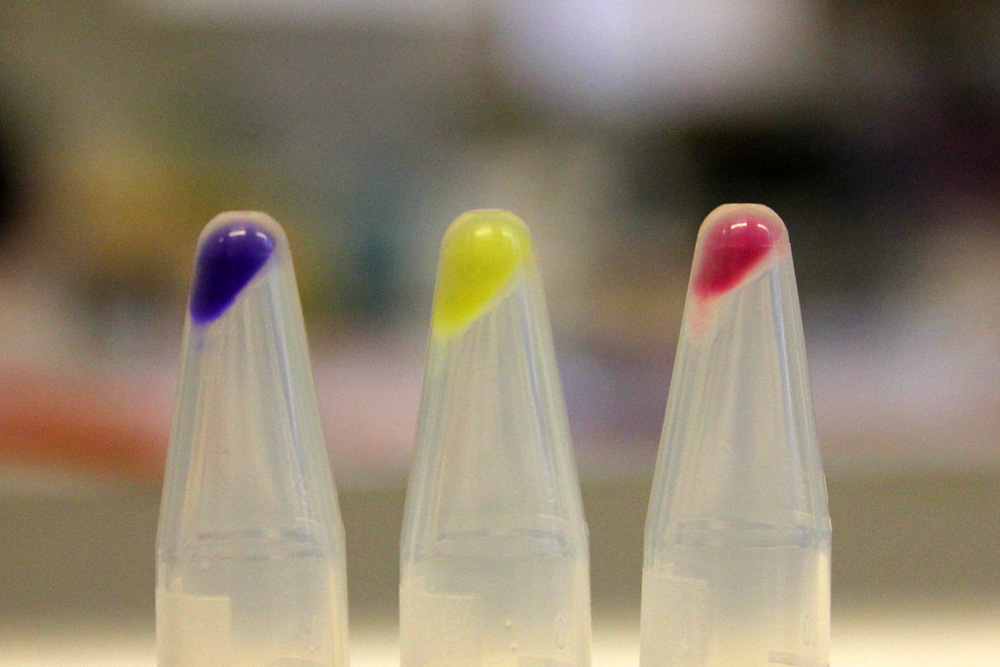
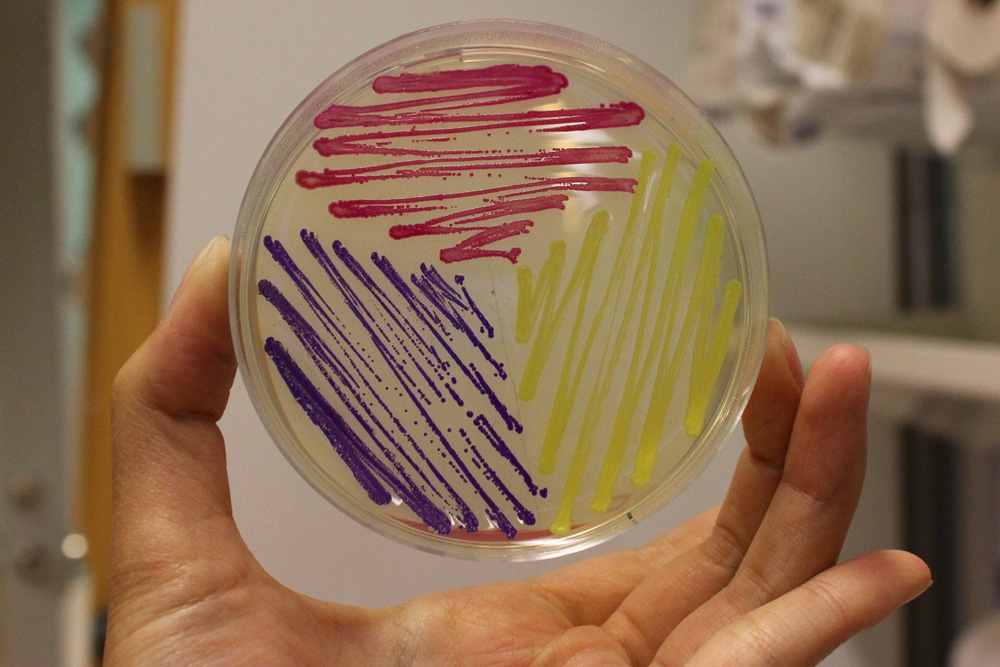
iGEM11_Uppsala-Sweden: Expression of chromoproteins. The images above show E coli constitutively expressing amilCP BBa_K592009 (blue), amilGFP BBa_K592010 (yellow) and RFP BBa_E1010 (red).
Peking iGEM 2016 has fused this part with triple spytag. The fused protein is participate in Peking’s polymer network. By adding this protein, the whole polymer network become visible in most conditions. If you want to learn more about Peking’s polymer network and the role of mRFP in this network, please click here https://parts.igem.org/Part:BBa_K1989004".
Contribution
Hong Kong-CUHK iGEM 2017 |width='60%' valign='top'|
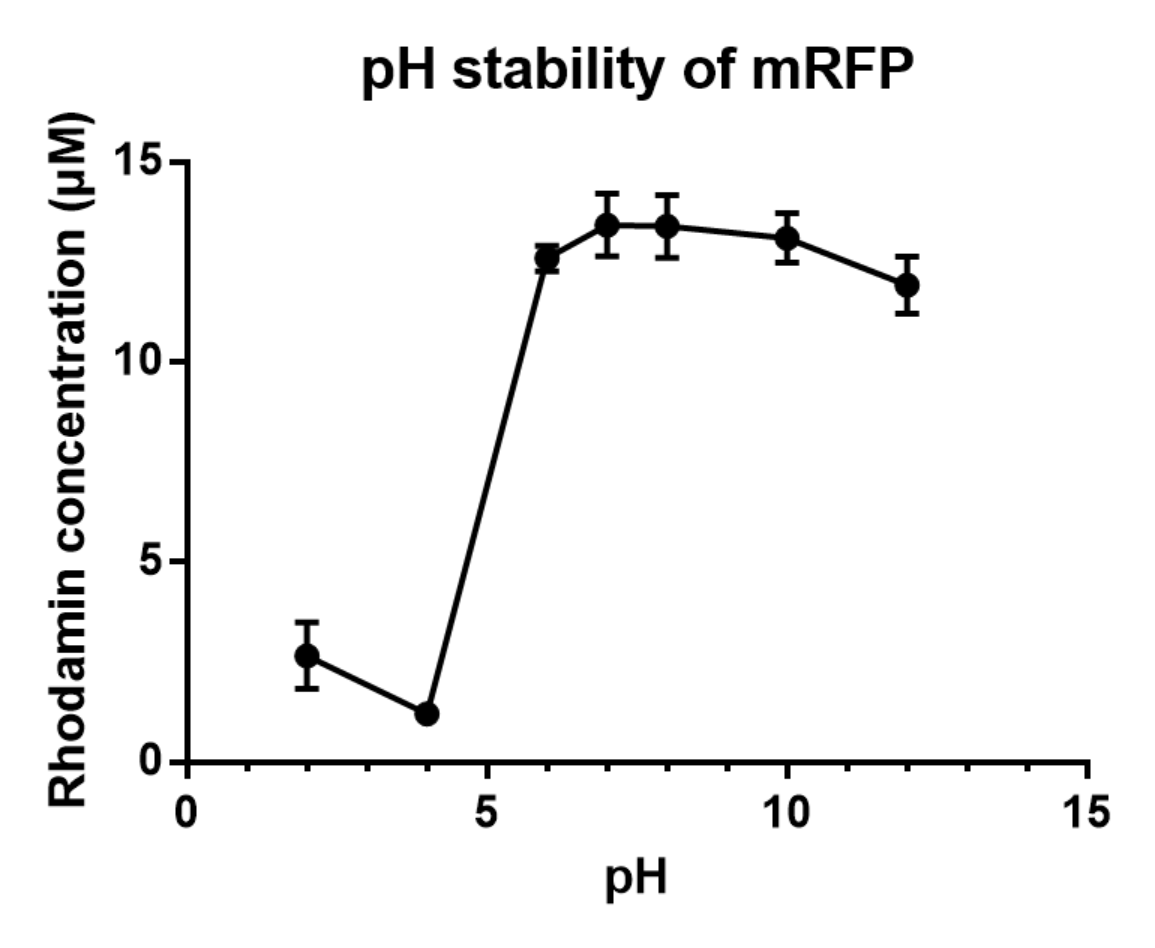
Charaterization of mRFP pH stabillity
We grew C41 bacteria with parts BBa_J61002 in 2XYT for 24 hours. Purifying the mRFP by Ion Exchange Chromatography and Hydrophobic Interaction Chromatography, we measured the fluoresece (ex ,em ) of purified mRFP. which is diluted to 10µg/100µl (total 200µl) in triplicates, into different buffers (ranges from pH2 to pH12). The result shows that the stability drops dramatically in pH condition below 6 and relatively stable in pH 6-10.
| Measurement Type | Fluorescence |
| Microplate name</td> | COSTAR 96 |
| Scan mode | orbital averaging |
| Scan diameter [nm] | 3 |
| Excitation | 550-20 |
| Emission | 605-40 |
| Dichronic filter | auto 572.5 |
| Gain | 500 |
| Focal height [nm] | 9 |
|};
|
Sequence and Features
Barcodes are discontinued, but one was appended to the sequence of this part. Composite parts using this part will include the barcode. More ...
Assembly Compatibility:
Parts table
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||