Difference between revisions of "Part:BBa E1010"
| Line 20: | Line 20: | ||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
<partinfo>BBa_E1010 SequenceAndFeatures</partinfo> | <partinfo>BBa_E1010 SequenceAndFeatures</partinfo> | ||
| + | |||
| + | ===Parts table=== | ||
| + | <html> | ||
| + | <html><!--- Please copy this table containing parameters for BBa_E1010 at the end of the parametrs section ahead of the references. ---><style type="text/css">table#AutoAnnotator {border:1px solid black; width:100%; border-collapse:collapse;} th#AutoAnnotatorHeader { border:1px solid black; width:100%; background-color: rgb(221, 221, 221);} td.AutoAnnotator1col { width:100%; border:1px solid black; } span.AutoAnnotatorSequence { font-family:'Courier New', Arial; } td.AutoAnnotatorSeqNum { text-align:right; width:2%; } td.AutoAnnotatorSeqSeq { width:98% } td.AutoAnnotatorSeqFeat1 { width:3% } td.AutoAnnotatorSeqFeat2a { width:27% } td.AutoAnnotatorSeqFeat2b { width:97% } td.AutoAnnotatorSeqFeat3 { width:70% } table.AutoAnnotatorNoBorder { border:0px; width:100%; border-collapse:collapse; } table.AutoAnnotatorWithBorder { border:1px solid black; width:100%; border-collapse:collapse; } td.AutoAnnotatorOuterAmino { border:0px solid black; width:20% } td.AutoAnnotatorInnerAmino { border:1px solid black; width:50% } td.AutoAnnotatorAminoCountingOuter { border:1px solid black; width:40%; } td.AutoAnnotatorBiochemParOuter { border:1px solid black; width:60%; } td.AutoAnnotatorAminoCountingInner1 { width: 7.5% } td.AutoAnnotatorAminoCountingInner2 { width:62.5% } td.AutoAnnotatorAminoCountingInner3 { width:30% } td.AutoAnnotatorBiochemParInner1 { width: 5% } td.AutoAnnotatorBiochemParInner2 { width:55% } td.AutoAnnotatorBiochemParInner3 { width:40% } td.AutoAnnotatorCodonUsage1 { width: 3% } td.AutoAnnotatorCodonUsage2 { width:14.2% } td.AutoAnnotatorCodonUsage3 { width:13.8% } td.AutoAnnotatorAlignment1 { width: 3% } td.AutoAnnotatorAlignment2 { width: 10% } td.AutoAnnotatorAlignment3 { width: 87% } td.AutoAnnotatorLocalizationOuter {border:1px solid black; width:40%} td.AutoAnnotatorGOOuter {border:1px solid black; width:60%} td.AutoAnnotatorLocalization1 { width: 7.5% } td.AutoAnnotatorLocalization2 { width: 22.5% } td.AutoAnnotatorLocalization3 { width: 70% } td.AutoAnnotatorGO1 { width: 5% } td.AutoAnnotatorGO2 { width: 35% } td.AutoAnnotatorGO3 { width: 60% } td.AutoAnnotatorPredFeat1 { width:3% } td.AutoAnnotatorPredFeat2a { width:27% } td.AutoAnnotatorPredFeat3 { width:70% } div.AutoAnnotator_trans { position:absolute; background:rgb(11,140,143); background-color:rgba(11,140,143, 0.8); height:5px; top:100px; } div.AutoAnnotator_sec_helix { position:absolute; background:rgb(102,0,102); background-color:rgba(102,0,102, 0.8); height:5px; top:110px; } div.AutoAnnotator_sec_strand { position:absolute; background:rgb(245,170,26); background-color:rgba(245,170,26, 1); height:5px; top:110px; } div.AutoAnnotator_acc_buried { position:absolute; background:rgb(89,168,15); background-color:rgba(89,168,15, 0.8); height:5px; top:120px; } div.AutoAnnotator_acc_exposed { position:absolute; background:rgb(0, 0, 255); background-color:rgba(0, 0, 255, 0.8); height:5px; top:120px; } div.AutoAnnotator_dis { position:absolute; text-align:center; font-family:Arial,Helvetica,sans-serif; background:rgb(255, 200, 0); background-color:rgba(255, 200, 0, 1); height:16px; width:16px; top:80px; border-radius:50%; } </style><div id='AutoAnnotator_container_1396983808569'><table id="AutoAnnotator"><tr><!-- Time stamp in ms since 1/1/1970 1396983808569 --><th id="AutoAnnotatorHeader" colspan="2">Protein data table for BioBrick <a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_E1010">BBa_E1010</a> automatically created by the <a href="http://2013.igem.org/Team:TU-Munich/Results/AutoAnnotator">BioBrick-AutoAnnotator</a> version 1.0</th></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Nucleotide sequence</strong> in <strong>RFC 10</strong>: (underlined part encodes the protein)<br><span class="AutoAnnotatorSequence"> <u>ATGGCTTCC ... ACCGGTGCT</u>TAATAACGCTGATAGTGCTAGTGTAGATCGC</span><br> <strong>ORF</strong> from nucleotide position 1 to 675 (excluding stop-codon)</td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Amino acid sequence:</strong> (RFC 25 scars in shown in bold, other sequence features underlined; both given below)<br><span class="AutoAnnotatorSequence"><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorSeqNum">1 <br>101 <br>201 </td><td class="AutoAnnotatorSeqSeq">MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTAKLKVTKGGPLPFAWDILSPQFQYGSKAYVKHPADIPDYLKLSFPEGFKWERVMNFE<br>DGGVVTVTQDSSLQDGEFIYKVKLRGTNFPSDGPVMQKKTMGWEASTERMYPEDGALKGEIKMRLKLKDGGHYDAEVKTTYMAKKPVQLPGAYKTDIKLD<br>ITSHNEDYTIVEQYERAEGRHSTGA*</td></tr></table></span></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Sequence features:</strong> (with their position in the amino acid sequence, see the <a href="http://2013.igem.org/Team:TU-Munich/Results/Software/FeatureList">list of supported features</a>)<table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorSeqFeat1"></td><td class="AutoAnnotatorSeqFeat2b">None of the supported features appeared in the sequence</td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Amino acid composition:</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Ala (A)</td><td class="AutoAnnotatorInnerAmino">12 (5.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Arg (R)</td><td class="AutoAnnotatorInnerAmino">9 (4.0%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Asn (N)</td><td class="AutoAnnotatorInnerAmino">4 (1.8%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Asp (D)</td><td class="AutoAnnotatorInnerAmino">14 (6.2%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Cys (C)</td><td class="AutoAnnotatorInnerAmino">0 (0.0%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Gln (Q)</td><td class="AutoAnnotatorInnerAmino">8 (3.6%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Glu (E)</td><td class="AutoAnnotatorInnerAmino">22 (9.8%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Gly (G)</td><td class="AutoAnnotatorInnerAmino">23 (10.2%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">His (H)</td><td class="AutoAnnotatorInnerAmino">5 (2.2%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Ile (I)</td><td class="AutoAnnotatorInnerAmino">9 (4.0%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Leu (L)</td><td class="AutoAnnotatorInnerAmino">12 (5.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Lys (K)</td><td class="AutoAnnotatorInnerAmino">22 (9.8%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Met (M)</td><td class="AutoAnnotatorInnerAmino">9 (4.0%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Phe (F)</td><td class="AutoAnnotatorInnerAmino">10 (4.4%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Pro (P)</td><td class="AutoAnnotatorInnerAmino">12 (5.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Ser (S)</td><td class="AutoAnnotatorInnerAmino">12 (5.3%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Thr (T)</td><td class="AutoAnnotatorInnerAmino">14 (6.2%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Trp (W)</td><td class="AutoAnnotatorInnerAmino">3 (1.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Tyr (Y)</td><td class="AutoAnnotatorInnerAmino">11 (4.9%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Val (V)</td><td class="AutoAnnotatorInnerAmino">14 (6.2%)</td></tr></table></td></tr></table></td></tr><tr><td class="AutoAnnotatorAminoCountingOuter"><strong>Amino acid counting</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Total number:</td><td class="AutoAnnotatorAminoCountingInner3">225</td></tr><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Positively charged (Arg+Lys):</td><td class="AutoAnnotatorAminoCountingInner3">31 (13.8%)</td></tr><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Negatively charged (Asp+Glu):</td><td class="AutoAnnotatorAminoCountingInner3">36 (16.0%)</td></tr><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Aromatic (Phe+His+Try+Tyr):</td><td class="AutoAnnotatorAminoCountingInner3">29 (12.9%)</td></tr></table></td><td class="AutoAnnotatorBiochemParOuter"><strong>Biochemical parameters</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Atomic composition:</td><td class="AutoAnnotatorBiochemParInner3">C<sub>1135</sub>H<sub>1749</sub>N<sub>299</sub>O<sub>347</sub>S<sub>9</sub></td></tr><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Molecular mass [Da]:</td><td class="AutoAnnotatorBiochemParInner3">25423.7</td></tr><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Theoretical pI:</td><td class="AutoAnnotatorBiochemParInner3">5.65</td></tr><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Extinction coefficient at 280 nm [M<sup>-1</sup> cm<sup>-1</sup>]:</td><td class="AutoAnnotatorBiochemParInner3">32890 / 32890 (all Cys red/ox)</td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Plot for hydrophobicity, charge, predicted secondary structure, solvent accessability, transmembrane helices and disulfid bridges</strong> <input type='button' id='hydrophobicity_charge_button' onclick='show_or_hide_plot_1396983808569()' value='Show'><span id="hydrophobicity_charge_explanation"></span><div id="hydrophobicity_charge_container" style='display:none'><div id="hydrophobicity_charge_placeholder0" style="width:100%;height:150px"></div><div id="hydrophobicity_charge_placeholder1" style="width:100%;height:150px"></div></div></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Codon usage</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorCodonUsage1"></td><td class="AutoAnnotatorCodonUsage2">Organism:</td><td class="AutoAnnotatorCodonUsage3"><i>E. coli</i></td><td class="AutoAnnotatorCodonUsage3"><i>B. subtilis</i></td><td class="AutoAnnotatorCodonUsage3"><i>S. cerevisiae</i></td><td class="AutoAnnotatorCodonUsage3"><i>A. thaliana</i></td><td class="AutoAnnotatorCodonUsage3"><i>P. patens</i></td><td class="AutoAnnotatorCodonUsage3">Mammals</td></tr><tr><td class="AutoAnnotatorCodonUsage1"></td><td class="AutoAnnotatorCodonUsage2">Codon quality (<a href="http://en.wikipedia.org/wiki/Codon_Adaptation_Index">CAI</a>):</td><td class="AutoAnnotatorCodonUsage3">excellent (0.84)</td><td class="AutoAnnotatorCodonUsage3">good (0.72)</td><td class="AutoAnnotatorCodonUsage3">good (0.68)</td><td class="AutoAnnotatorCodonUsage3">good (0.74)</td><td class="AutoAnnotatorCodonUsage3">good (0.78)</td><td class="AutoAnnotatorCodonUsage3">good (0.71)</td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Alignments</strong> (obtained from <a href='http://predictprotein.org'>PredictProtein.org</a>)<table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorAlignment1"></td><td class="AutoAnnotatorAlignment2">SwissProt:</td><td class="AutoAnnotatorAlignment3"><a href='http://www.uniprot.org/uniprot/Q9U6Y8'>Q9U6Y8</a> (86% identity on 221 AAs), <a href='http://www.uniprot.org/uniprot/P83690'>P83690</a> (63% identity on 215 AAs)</td></tr><tr><td class="AutoAnnotatorAlignment1"></td><td class="AutoAnnotatorAlignment2">TrEML:</td><td class="AutoAnnotatorAlignment3"><a href='http://www.uniprot.org/uniprot/Q5S3G8'>Q5S3G8</a> (95% identity on 225 AAs), <a href='http://www.uniprot.org/uniprot/D0VWW2'>D0VWW2</a> (94% identity on 220 AAs)</td></tr><tr><td class="AutoAnnotatorAlignment1"></td><td class="AutoAnnotatorAlignment2">PDB:</td><td class="AutoAnnotatorAlignment3"><a href='http://www.rcsb.org/pdb/explore/explore.do?structureId=2h5q'>2h5q</a> (94% identity on 216 AAs), <a href='http://www.rcsb.org/pdb/explore/explore.do?structureId=2qlg'>2qlg</a> (94% identity on 215 AAs)</td></tr></table></td></tr><tr><th id='AutoAnnotatorHeader' colspan="2"><strong>Predictions</strong> (obtained from <a href='http://predictprotein.org'>PredictProtein.org</a>)</th></tr><tr><td class="AutoAnnotatorLocalizationOuter"><strong>Subcellular Localization</strong> (reliability in brackets)<table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorLocalization1"></td><td class="AutoAnnotatorLocalization2">Archaea:</td><td class="AutoAnnotatorLocalization3">secreted (100%)</td></tr><tr><td class="AutoAnnotatorLocalization1"></td><td class="AutoAnnotatorLocalization2">Bacteria:</td><td class="AutoAnnotatorLocalization3">cytosol (52%)</td></tr><tr><td class="AutoAnnotatorLocalization1"></td><td class="AutoAnnotatorLocalization2">Eukarya:</td><td class="AutoAnnotatorLocalization3">cytosol (20%)</td></tr></table></td><td class="AutoAnnotatorGOOuter"><strong>Gene Ontology</strong> (reliability in brackets)<br><table class="AutoAnnotatorNoBorder"><tr><td class='AutoAnnotatorGO1'></td><td class='AutoAnnotatorGO2'>Molecular Function Ontology:</td><td class='AutoAnnotatorGO3'> - </td></tr><tr><td class='AutoAnnotatorGO1'></td><td class='AutoAnnotatorGO2'>Biological Process Ontology:</td><td class='AutoAnnotatorGO3'><a href='http://amigo.geneontology.org/cgi-bin/amigo/term_details?term=GO:0018298'>GO:0018298</a> (40%), <a href='http://amigo.geneontology.org/cgi-bin/amigo/term_details?term=GO:0008218'>GO:0008218</a> (27%)</td></tr><tr><td class='AutoAnnotatorGO1'> </td><td class='AutoAnnotatorGO2'> </td><td class='AutoAnnotatorGO3'> </td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Predicted features:</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorPredFeat1"></td><td class="AutoAnnotatorPredFeat2a">Disulfid bridges:</td><td class="AutoAnnotatorPredFeat3"> - </td></tr><tr><td class="AutoAnnotatorPredFeat1"></td><td class="AutoAnnotatorPredFeat2a">Transmembrane helices:</td><td class="AutoAnnotatorPredFeat3"> - </td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"> The BioBrick-AutoAnnotator was created by <a href="http://2013.igem.org/Team:TU-Munich">TU-Munich 2013</a> iGEM team. For more information please see the <a href="http://2013.igem.org/Team:TU-Munich/Results/Software">documentation</a>.<br>If you have any questions, comments or suggestions, please leave us a <a href="http://2013.igem.org/Team:TU-Munich/Results/AutoAnnotator">comment</a>.</td></tr></table></div><br><!-- IMPORTANT: DON'T REMOVE THIS LINE, OTHERWISE NOT SUPPORTED FOR IE BEFORE 9 --><!--[if lte IE 8]><script language="javascript" type="text/javascript" src="http://2013.igem.org/Team:TU-Munich/excanvas.js"></script><![endif]--><script type='text/javascript' src='http://code.jquery.com/jquery-1.10.0.min.js'></script><script type='text/javascript' src='http://2013.igem.org/Team:TU-Munich/Flot.js?action=raw&ctype=text/js'></script><script>function show_or_hide_plot_1396983808569(){hydrophobicity_datapoints = [[2.5,-0.28],[3.5,-1.36],[4.5,-0.88],[5.5,0.18],[6.5,-0.44],[7.5,-0.44],[8.5,0.82],[9.5,0.36],[10.5,-1.44],[11.5,-0.10],[12.5,-0.18],[13.5,0.10],[14.5,-1.18],[15.5,0.10],[16.5,-1.16],[17.5,-0.46],[18.5,-1.46],[19.5,0.28],[20.5,-0.80],[21.5,-0.18],[22.5,-0.74],[23.5,-1.28],[24.5,-1.56],[25.5,-1.56],[26.5,-0.58],[27.5,-0.64],[28.5,-0.02],[29.5,-1.28],[30.5,-0.66],[31.5,-2.26],[32.5,-1.64],[33.5,-2.46],[34.5,-2.08],[35.5,-2.26],[36.5,-2.26],[37.5,-2.26],[38.5,-1.50],[39.5,-1.88],[40.5,-1.76],[41.5,-0.70],[42.5,-1.40],[43.5,-0.50],[44.5,-0.58],[45.5,0.40],[46.5,-0.10],[47.5,-0.10],[48.5,-0.94],[49.5,-0.24],[50.5,-1.40],[51.5,-0.50],[52.5,-0.04],[53.5,0.60],[54.5,1.04],[55.5,1.18],[56.5,-0.28],[57.5,0.94],[58.5,1.14],[59.5,0.62],[60.5,0.48],[61.5,0.48],[62.5,0.14],[63.5,-1.32],[64.5,-1.42],[65.5,-1.18],[66.5,-0.64],[67.5,-1.98],[68.5,-0.92],[69.5,-0.92],[70.5,-0.00],[71.5,-0.62],[72.5,-0.48],[73.5,-1.16],[74.5,-0.54],[75.5,-2.08],[76.5,-0.40],[77.5,-0.08],[78.5,-0.46],[79.5,-1.08],[80.5,0.38],[81.5,-1.30],[82.5,-0.22],[83.5,0.32],[84.5,1.14],[85.5,0.06],[86.5,0.14],[87.5,-0.70],[88.5,0.02],[89.5,-1.32],[90.5,-1.18],[91.5,-1.18],[92.5,-2.00],[93.5,-1.72],[94.5,-0.56],[95.5,-1.08],[96.5,0.18],[97.5,0.38],[98.5,-1.16],[99.5,-1.62],[100.5,-1.00],[101.5,-0.72],[102.5,0.82],[103.5,1.38],[104.5,2.30],[105.5,2.24],[106.5,0.70],[107.5,-0.84],[108.5,-0.86],[109.5,-1.86],[110.5,-0.96],[111.5,-0.96],[112.5,-0.96],[113.5,-0.88],[114.5,-1.42],[115.5,-1.62],[116.5,-0.02],[117.5,0.42],[118.5,-0.28],[119.5,1.26],[120.5,-0.08],[121.5,-0.22],[122.5,-0.86],[123.5,-0.16],[124.5,-1.14],[125.5,-1.06],[126.5,-1.26],[127.5,-0.68],[128.5,-0.76],[129.5,-1.32],[130.5,-0.70],[131.5,-1.58],[132.5,-0.42],[133.5,0.12],[134.5,0.12],[135.5,-0.58],[136.5,-1.04],[137.5,-2.02],[138.5,-2.02],[139.5,-1.40],[140.5,-0.80],[141.5,-0.72],[142.5,-0.22],[143.5,-0.76],[144.5,-0.82],[145.5,-1.34],[146.5,-1.54],[147.5,-1.52],[148.5,-1.62],[149.5,-1.80],[150.5,-1.80],[151.5,-1.60],[152.5,-2.06],[153.5,-1.44],[154.5,-0.36],[155.5,-0.44],[156.5,0.18],[157.5,-0.44],[158.5,0.10],[159.5,-1.44],[160.5,-0.28],[161.5,-1.10],[162.5,0.36],[163.5,-1.32],[164.5,0.22],[165.5,-0.94],[166.5,-0.74],[167.5,-1.58],[168.5,-0.88],[169.5,-2.28],[170.5,-1.76],[171.5,-1.76],[172.5,-1.32],[173.5,-1.94],[174.5,-0.46],[175.5,-0.98],[176.5,-0.42],[177.5,-0.92],[178.5,-0.48],[179.5,-0.94],[180.5,0.20],[181.5,-0.44],[182.5,-1.08],[183.5,-1.14],[184.5,-0.68],[185.5,-1.74],[186.5,-0.20],[187.5,0.26],[188.5,0.50],[189.5,0.02],[190.5,0.46],[191.5,-1.08],[192.5,-0.90],[193.5,-1.52],[194.5,-0.98],[195.5,-1.50],[196.5,0.04],[197.5,-0.52],[198.5,1.08],[199.5,0.04],[200.5,0.66],[201.5,-0.74],[202.5,-0.74],[203.5,-2.34],[204.5,-2.90],[205.5,-3.00],[206.5,-2.50],[207.5,-0.90],[208.5,0.64],[209.5,0.64],[210.5,0.20],[211.5,0.08],[212.5,-1.52],[213.5,-3.26],[214.5,-2.20],[215.5,-2.20],[216.5,-2.02],[217.5,-2.22],[218.5,-1.96],[219.5,-2.48],[220.5,-1.92],[221.5,-1.92],[222.5,-0.66]];charge_datapoints = [[2.5,-0.20],[3.5,-0.40],[4.5,-0.40],[5.5,-0.40],[6.5,-0.20],[7.5,-0.20],[8.5,0.00],[9.5,0.00],[10.5,0.20],[11.5,0.00],[12.5,0.40],[13.5,0.40],[14.5,0.60],[15.5,0.40],[16.5,0.20],[17.5,0.00],[18.5,0.00],[19.5,-0.20],[20.5,-0.20],[21.5,0.00],[22.5,0.10],[23.5,-0.10],[24.5,-0.10],[25.5,-0.30],[26.5,-0.30],[27.5,-0.60],[28.5,-0.40],[29.5,-0.60],[30.5,-0.40],[31.5,-0.60],[32.5,-0.40],[33.5,-0.20],[34.5,0.00],[35.5,0.00],[36.5,0.00],[37.5,0.00],[38.5,-0.20],[39.5,-0.20],[40.5,-0.20],[41.5,0.00],[42.5,0.20],[43.5,0.20],[44.5,0.40],[45.5,0.40],[46.5,0.40],[47.5,0.40],[48.5,0.40],[49.5,0.20],[50.5,0.20],[51.5,0.20],[52.5,0.00],[53.5,0.00],[54.5,0.00],[55.5,0.00],[56.5,-0.20],[57.5,-0.20],[58.5,-0.20],[59.5,-0.20],[60.5,-0.20],[61.5,0.00],[62.5,0.00],[63.5,0.00],[64.5,0.00],[65.5,0.00],[66.5,0.00],[67.5,0.20],[68.5,0.20],[69.5,0.20],[70.5,0.20],[71.5,0.40],[72.5,0.30],[73.5,0.30],[74.5,0.30],[75.5,0.10],[76.5,-0.10],[77.5,-0.20],[78.5,-0.40],[79.5,-0.40],[80.5,-0.20],[81.5,0.00],[82.5,0.00],[83.5,0.20],[84.5,0.20],[85.5,0.20],[86.5,-0.20],[87.5,-0.20],[88.5,-0.20],[89.5,0.00],[90.5,0.00],[91.5,0.00],[92.5,0.20],[93.5,0.20],[94.5,0.00],[95.5,0.00],[96.5,0.20],[97.5,-0.20],[98.5,-0.40],[99.5,-0.40],[100.5,-0.40],[101.5,-0.40],[102.5,-0.20],[103.5,0.00],[104.5,0.00],[105.5,0.00],[106.5,0.00],[107.5,-0.20],[108.5,-0.20],[109.5,-0.20],[110.5,-0.20],[111.5,-0.20],[112.5,-0.20],[113.5,-0.20],[114.5,-0.40],[115.5,-0.40],[116.5,-0.40],[117.5,-0.20],[118.5,0.00],[119.5,0.20],[120.5,0.40],[121.5,0.40],[122.5,0.60],[123.5,0.40],[124.5,0.40],[125.5,0.20],[126.5,0.20],[127.5,0.00],[128.5,0.00],[129.5,-0.20],[130.5,-0.20],[131.5,-0.20],[132.5,-0.20],[133.5,-0.20],[134.5,0.00],[135.5,0.20],[136.5,0.40],[137.5,0.40],[138.5,0.40],[139.5,0.40],[140.5,0.20],[141.5,-0.20],[142.5,-0.20],[143.5,-0.20],[144.5,-0.20],[145.5,-0.40],[146.5,0.00],[147.5,0.00],[148.5,0.00],[149.5,0.00],[150.5,0.00],[151.5,-0.40],[152.5,-0.40],[153.5,-0.40],[154.5,-0.40],[155.5,0.00],[156.5,0.20],[157.5,0.00],[158.5,0.00],[159.5,0.20],[160.5,0.00],[161.5,0.20],[162.5,0.40],[163.5,0.60],[164.5,0.40],[165.5,0.60],[166.5,0.20],[167.5,0.20],[168.5,0.00],[169.5,0.10],[170.5,-0.10],[171.5,-0.10],[172.5,-0.10],[173.5,-0.30],[174.5,-0.40],[175.5,-0.20],[176.5,0.00],[177.5,0.00],[178.5,0.20],[179.5,0.20],[180.5,0.00],[181.5,0.20],[182.5,0.40],[183.5,0.40],[184.5,0.40],[185.5,0.40],[186.5,0.20],[187.5,0.00],[188.5,0.00],[189.5,0.00],[190.5,0.00],[191.5,0.20],[192.5,0.20],[193.5,0.00],[194.5,0.00],[195.5,0.20],[196.5,0.00],[197.5,-0.20],[198.5,0.00],[199.5,0.00],[200.5,-0.20],[201.5,-0.10],[202.5,0.10],[203.5,-0.10],[204.5,-0.30],[205.5,-0.30],[206.5,-0.40],[207.5,-0.40],[208.5,-0.20],[209.5,-0.20],[210.5,-0.20],[211.5,-0.20],[212.5,-0.40],[213.5,-0.20],[214.5,0.00],[215.5,-0.20],[216.5,-0.20],[217.5,0.20],[218.5,0.10],[219.5,0.10],[220.5,0.30],[221.5,0.30],[222.5,0.10]];dis_datapoints = [];trans_datapoints = [];sec_helix_datapoints = [[8,10],[57,66],[79,85]];sec_strand_datapoints = [[13,22],[25,31],[41,49],[93,100],[104,113],[117,126],[148,152],[156,168],[172,183],[195,200],[201,203],[210,221]];acc_exposed_datapoints = [[1,6],[9,10],[13,13],[15,15],[19,19],[23,23],[26,26],[32,32],[34,35],[37,37],[39,39],[41,41],[50,51],[53,53],[70,70],[74,74],[77,78],[81,81],[84,84],[88,90],[92,92],[110,110],[114,117],[128,128],[131,132],[134,134],[137,140],[142,142],[144,145],[148,148],[154,155],[158,158],[168,169],[171,171],[178,178],[182,182],[184,186],[188,188],[191,191],[200,200],[203,203],[205,207],[209,209],[212,212],[215,216],[218,218],[223,225]];acc_buried_datapoints = [[7,8],[12,12],[14,14],[16,16],[18,18],[20,20],[22,22],[24,24],[27,27],[29,29],[31,31],[33,33],[40,40],[44,44],[46,46],[48,48],[54,58],[60,65],[67,68],[71,72],[79,79],[82,83],[87,87],[91,91],[93,93],[95,97],[99,99],[103,107],[111,111],[113,113],[118,118],[120,120],[122,122],[124,124],[126,126],[129,129],[133,133],[135,136],[146,146],[150,151],[156,157],[159,159],[161,161],[163,163],[165,165],[167,167],[173,173],[175,175],[177,177],[179,179],[183,183],[187,187],[189,190],[194,195],[197,197],[199,199],[211,211],[217,217],[219,219]];flot_plot_options = []; flot_plot_options[0] = {grid: {borderWidth: {top: 0,right: 0,bottom: 0,left: 0}},legend: {show: false},xaxes: [{show: true,min: 0,max: 200,ticks: [[0.5, '1'], [24.5, '25'], [49.5, '50'], [74.5, '75'], [99.5, '100'], [124.5, '125'], [149.5, '150'], [174.5, '175'], [199.5, '200']],tickLength: -5}],yaxes: [{show: true,ticks: [[0, '0'], [4.5,'hydro-<br>phobic '], [-4.5,'hydro-<br>philic ']],min: -4.5,max: +4.5,font: {size: 12,lineHeight: 14,style: 'italic',weight: 'bold',family: 'sans-serif',variant: 'small-caps',color: 'rgba(100,149,237,1)'}},{show: true,ticks: [[0, ''], [1,'positive<br> charge'], [-1,'negative<br> charge']],position: 'right',min: -1,max: 1,font: {size: 12,lineHeight: 14,style: 'italic',weight: 'bold',family: 'sans-serif',variant: 'small-caps',color: 'rgba(255,99,71,1)'}}]};number_of_plots = 2;for ( plot_num = 1 ; plot_num < number_of_plots ; plot_num ++){flot_plot_options[plot_num] = $.extend(true, {} ,flot_plot_options[0]);flot_plot_options[plot_num].xaxes = [{min: plot_num*200,max: (plot_num + 1)*200,ticks: [ [plot_num*200 + 0.5, (plot_num*200 + 1).toString()], [plot_num*200 + 24.5, (plot_num*200 + 25).toString()], [plot_num*200 + 49.5, (plot_num*200 + 50).toString()], [plot_num*200 + 74.5, (plot_num*200 + 75).toString()], [plot_num*200 + 99.5, (plot_num*200 + 100).toString()], [plot_num*200 + 124.5, (plot_num*200 + 125).toString()], [plot_num*200 + 149.5, (plot_num*200 + 150).toString()], [plot_num*200 + 174.5, (plot_num*200 + 175).toString()], [plot_num*200 + 199.5, (plot_num*200 + 200).toString()] ],tickLength: -5}];};try {if( $('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_button').val() =='Show' ){$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_container').css('display','block');$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_button').val('Hide');var description_html = '<div id=\'AutoAnnotator_plot_selectors\'>';description_html = description_html + '<br> <input type=\'checkbox\' id=\'hydrophobicity_checkbox\' checked=\'checked\'> Moving average over 5 amino acids for hydrophobicity (<img src=\'https://static.igem.org/mediawiki/2013/e/e9/TUM13_hydrophobicity_icon.png\' alt=\'blue graph\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'charge_checkbox\' checked=\'checked\'> Moving average over 5 amino acids for charge (<img src=\'https://static.igem.org/mediawiki/2013/3/3e/TUM13_charge_icon.png\' alt=\'red graph\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'dis_checkbox\' checked=\'checked\'> Predicted disulfid bridges (<img src=\'https://static.igem.org/mediawiki/2013/2/28/TUM13_dis_icon.png\' alt=\'yellow circle\' height=\'10\'></img>) with the number of the bridge in the center';description_html = description_html + '<br> <input type=\'checkbox\' id=\'trans_checkbox\' checked=\'checked\'> Predicted transmembrane helices (<img src=\'https://static.igem.org/mediawiki/2013/7/78/TUM13_trans_icon.png\' alt=\'turquois bars\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'sec_checkbox\' checked=\'checked\'> Predicted secondary structure: Helices (<img src=\'https://static.igem.org/mediawiki/2013/b/bf/TUM13_helix_icon.png\' alt=\'violet bars\' height=\'10\'></img>) and beta-strands (<img src=\'https://static.igem.org/mediawiki/2013/b/bf/TUM13_strand_icon.png\' alt=\'yellow bars\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'acc_checkbox\' checked=\'checked\'> Predicted solvent accessability: Exposed (<img src=\'https://static.igem.org/mediawiki/2013/1/16/TUM13_exposed_icon.png\' alt=\'blue bars\' height=\'10\'></img>) and buried (<img src=\'https://static.igem.org/mediawiki/2013/0/0b/TUM13_buried_icon.png\' alt=\'green bars\' height=\'10\'></img>) residues';description_html = description_html + '<br></div>';$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_explanation').html(description_html);plot_according_to_selectors_1396983808569();$('#AutoAnnotator_container_1396983808569 #AutoAnnotator_plot_selectors').find('input').click(plot_according_to_selectors_1396983808569);}else{$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_container').css('display','none');$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_button').val('Show');$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_explanation').html('');}}catch(err){txt='There was an error with the button controlling the visibility of the plot.\n';txt=txt+'The originating error is:\n' + err + '\n\n';alert(txt);}};function plot_according_to_selectors_1396983808569(){try{var plot_datasets = [[],[]];if($('#AutoAnnotator_container_1396983808569 #hydrophobicity_checkbox').prop('checked') == true){plot_datasets[0] = { color: 'rgba(100,149,237,1)',data: hydrophobicity_datapoints,label: 'Hydrophobicity',lines: { show: true, fill: true, fillColor: 'rgba(100,149,237,0.1)' },yaxis: 1};}if($('#AutoAnnotator_container_1396983808569 #charge_checkbox').prop('checked') == true){plot_datasets[1] = {color: 'rgba(255,99,71,1)',data: charge_datapoints,label: 'Charge',lines: { show: true, fill: true, fillColor: 'rgba(255,99,71,0.1)' },yaxis: 2};}for (plot_num = 0 ; plot_num < number_of_plots ; plot_num ++){$.plot('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_placeholder'+ plot_num.toString(), plot_datasets, flot_plot_options[plot_num] );}var screen_width = $('canvas.flot-base').width(); var pos_of_first_tick = 46;var pos_of_last_tick = screen_width - 51;var tick_diff = (screen_width - 97)/199;if($('#AutoAnnotator_container_1396983808569 #dis_checkbox').prop('checked') == true){for ( j = 0 ; j < dis_datapoints.length ; j++ ){$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_placeholder' + Math.floor((dis_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_dis\' style=\'left:' + ((pos_of_first_tick - 8 + (dis_datapoints[j][0] - 1)*tick_diff - Math.floor((dis_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px;\'><b>' + (j+1) + '</b></div>');$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_placeholder' + Math.floor((dis_datapoints[j][1] - 1)/200) ).append('<div class=\'AutoAnnotator_dis\' style=\'left:' + ((pos_of_first_tick - 8 + (dis_datapoints[j][1] - 1)*tick_diff - Math.floor((dis_datapoints[j][1] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px;\'><b>' + (j+1) + '</b></div>');}}if($('#AutoAnnotator_container_1396983808569 #trans_checkbox').prop('checked') == true){for ( j = 0 ; j < trans_datapoints.length ; j++ ){$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_placeholder' + Math.floor((trans_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_trans\' style=\'width:' + (((trans_datapoints[j][1] - trans_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px ;left:' + ((pos_of_first_tick + (trans_datapoints[j][0] - 1.5)*tick_diff - Math.floor((trans_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}}if($('#AutoAnnotator_container_1396983808569 #sec_checkbox').prop('checked') == true){for ( j = 0 ; j < sec_helix_datapoints.length ; j++ ){$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_placeholder' + Math.floor((sec_helix_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_sec_helix\' style=\'width:' + (((sec_helix_datapoints[j][1] - sec_helix_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (sec_helix_datapoints[j][0] - 1.5)*tick_diff - Math.floor((sec_helix_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}for ( j = 0 ; j < sec_strand_datapoints.length ; j++ ){$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_placeholder' + Math.floor((sec_strand_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_sec_strand\' style=\'width:' + (((sec_strand_datapoints[j][1] - sec_strand_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (sec_strand_datapoints[j][0] - 1.5)*tick_diff - Math.floor((sec_strand_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}}if($('#AutoAnnotator_container_1396983808569 #acc_checkbox').prop('checked') == true){for ( j = 0 ; j < acc_buried_datapoints.length ; j++ ){$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_placeholder' + Math.floor((acc_buried_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_acc_buried\' style=\'width:' + (((acc_buried_datapoints[j][1] - acc_buried_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (acc_buried_datapoints[j][0] - 1.5)*tick_diff - Math.floor((acc_buried_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}for ( j = 0 ; j < acc_exposed_datapoints.length ; j++ ){$('#AutoAnnotator_container_1396983808569 #hydrophobicity_charge_placeholder' + Math.floor((acc_exposed_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_acc_exposed\' style=\'width:' + (((acc_exposed_datapoints[j][1] - acc_exposed_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (acc_exposed_datapoints[j][0] - 1.5)*tick_diff - Math.floor((acc_exposed_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}}}catch(err){txt='There was an error while drawing the selected elements for the plot.\n';txt=txt+'The originating error is:\n' + err + '\n\n';throw(txt);}}var jqAutoAnnotator = jQuery.noConflict(true);</script></html> | ||
Revision as of 19:05, 8 April 2014
**highly** engineered mutant of red fluorescent protein from Discosoma striata (coral)
monomeric RFP: Red Fluorescent Protein. Excitation peak: 584 nm Emission peak: 607 nm
Usage and Biology
Robert E. Campbell started with Discosoma RFP (DsRed) and evolved a faster folding, monomeric variant. See paper listed in source. Codon optimized for expression in bacteria (?? DE)
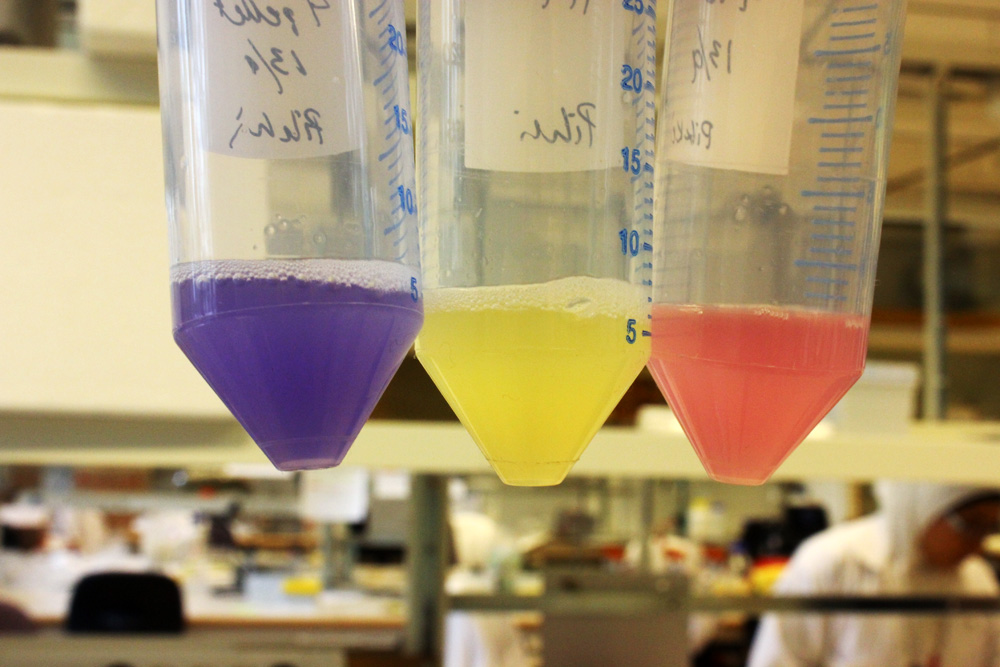
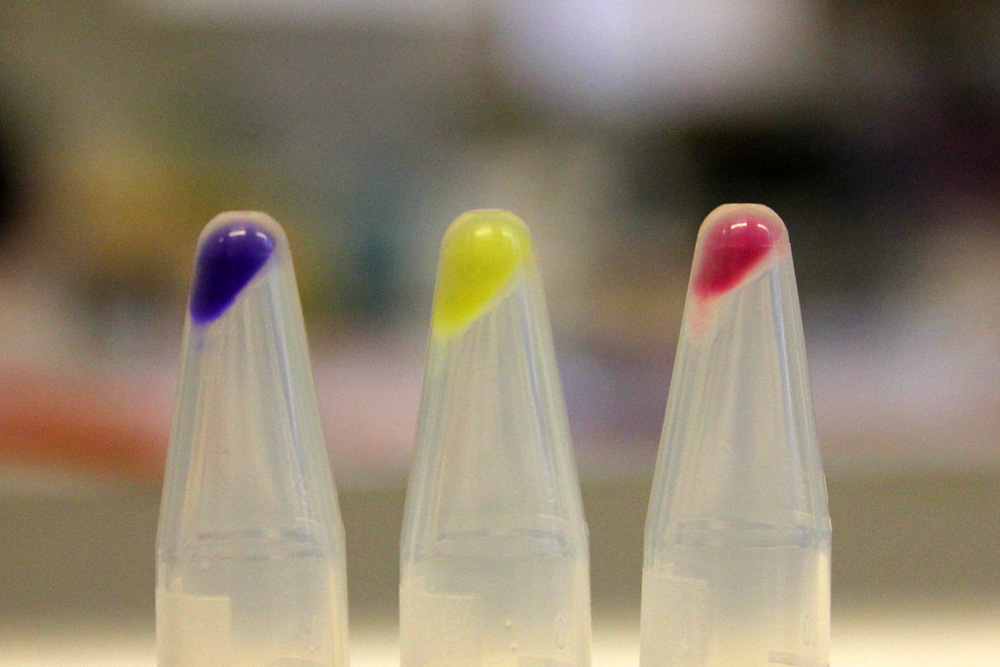
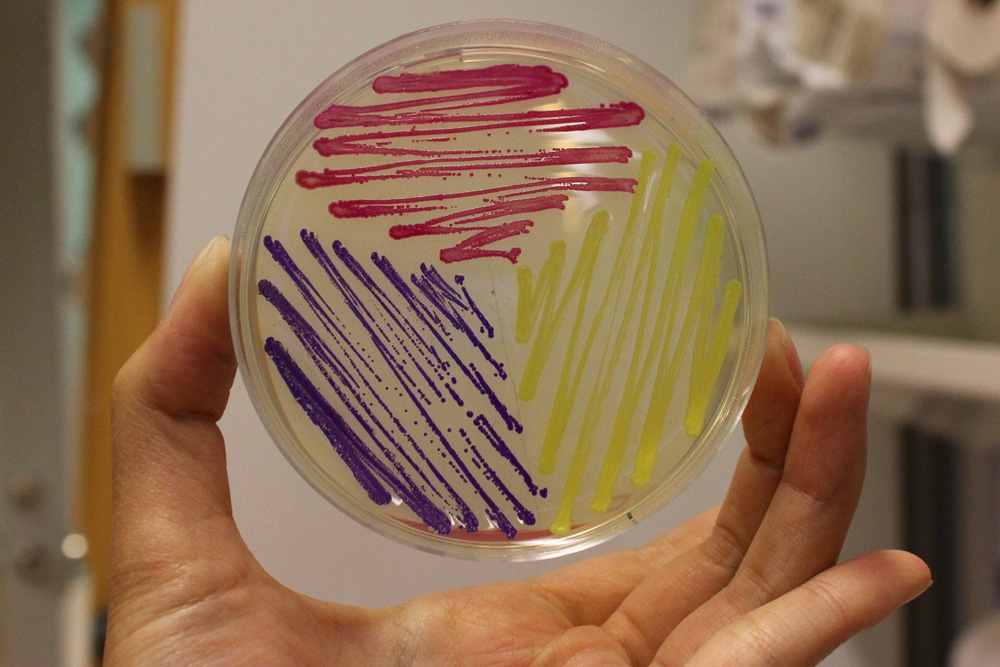
iGEM11_Uppsala-Sweden: Expression of chromoproteins. The images above show E coli constitutively expressing amilCP BBa_K592009 (blue), amilGFP BBa_K592010 (yellow) and RFP BBa_E1010 (red).
Sequence and Features
Barcodes are discontinued, but one was appended to the sequence of this part. Composite parts using this part will include the barcode. More ...
Assembly Compatibility:
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 555
Illegal AgeI site found at 667 - 1000COMPATIBLE WITH RFC[1000]
Parts table
| Protein data table for BioBrick BBa_E1010 automatically created by the BioBrick-AutoAnnotator version 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide sequence in RFC 10: (underlined part encodes the protein) ATGGCTTCC ... ACCGGTGCTTAATAACGCTGATAGTGCTAGTGTAGATCGC ORF from nucleotide position 1 to 675 (excluding stop-codon) | ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid sequence: (RFC 25 scars in shown in bold, other sequence features underlined; both given below)
| ||||||||||||||||||||||||||||||||||||||||||||||
Sequence features: (with their position in the amino acid sequence, see the list of supported features)
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid composition:
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid counting
| Biochemical parameters
| |||||||||||||||||||||||||||||||||||||||||||||
| Plot for hydrophobicity, charge, predicted secondary structure, solvent accessability, transmembrane helices and disulfid bridges | ||||||||||||||||||||||||||||||||||||||||||||||
Codon usage
| ||||||||||||||||||||||||||||||||||||||||||||||
Alignments (obtained from PredictProtein.org)
| ||||||||||||||||||||||||||||||||||||||||||||||
| Predictions (obtained from PredictProtein.org) | ||||||||||||||||||||||||||||||||||||||||||||||
Subcellular Localization (reliability in brackets)
| Gene Ontology (reliability in brackets)
| |||||||||||||||||||||||||||||||||||||||||||||
Predicted features:
| ||||||||||||||||||||||||||||||||||||||||||||||
| The BioBrick-AutoAnnotator was created by TU-Munich 2013 iGEM team. For more information please see the documentation. If you have any questions, comments or suggestions, please leave us a comment. | ||||||||||||||||||||||||||||||||||||||||||||||