Difference between revisions of "Part:BBa K863006"
| Line 1: | Line 1: | ||
| − | |||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K863006 short</partinfo> | <partinfo>BBa_K863006 short</partinfo> | ||
| Line 17: | Line 16: | ||
<partinfo>BBa_K863006 parameters</partinfo> | <partinfo>BBa_K863006 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
| + | First some trials of shaking flask cultivations were made with changing parameters to identify the best conditions for the production of the laccase CueO from ''E. coli'' BL21 (DE3) named ECOL fused to a His tag. Because of no measured activity in the cell lysate a purification method was established (using Ni-NTA His tag resin and Syringe or ÄKTA method). The purified ECOL could be identified by SDS-PAGE (molecular weight of 53.4 kDa) as well as by MALDI-TOF. The fractionated samples were also tested concerning their activity. A maximal activity of X was reached. After measuring activity of ECOL a scale up was made up to 3 L and then also up to 6 L that enables an intense screening afterwards. The scale up will be important for further application. | ||
| + | <html> | ||
| + | </p> | ||
| + | </html> | ||
| + | __TOC__ | ||
| + | |||
| + | ==Cultivation, Purification and SDS-PAGE== | ||
| + | ===Shaking Flask Cultivations=== | ||
| + | <div style="text-align:justify;"> | ||
| + | The first trials to produce ECOL were produced in shaking flask with various designs (from 100 mL<sup>-1</sup> to 1 L flasks, with and without baffles) and under different conditions. The parameters tested during our screening experiments were temperature (27 °C,30 °C and 37 °C), concentrations of chloramphenicol (20-170 µg mL<sup>-1</sup>), various induction strategies ([http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Materials#Autoinduction_medium autoinduction] and manual induction) and cultivation time (6 - 24 h). Furthermore it was cultivated with and without 0.25 mM CuCl<sub>2</sub> to provide a sufficient amount of copper, which is needed for the active center of the laccase. Based on the screening experiments we identified the best conditions under which ECOL was expressed. The addition of CuCl<sub>2</sub> did not increase the activity, so it was omitted. | ||
| + | |||
| + | * flask design: shaking flask without baffles | ||
| + | * medium: [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Materials#Autoinduction_medium autoinduction medium] | ||
| + | * antibiotics: 60 µg mL<sup>-1</sup> chloramphenicol | ||
| + | * temperature: 37 °C | ||
| + | * cultivation time: 12 h | ||
| + | |||
| + | The reproducibility of the measured data and results were investigated for the shaking flask and bioreactor cultivation. | ||
| + | </div> | ||
| + | |||
| + | ===3 L Fermentation ''E. coli'' KRX with <partinfo>BBa_K863005</partinfo>=== | ||
| + | |||
| + | [[Image:Bielefeld2012_ECOL3LFermentation.jpg|450px|thumb|left|'''Figure 1:''' Fermentation of ''E. coli'' KRXwith <partinfo>BBa_K863005</partinfo> (ECOL) in an Infors Labfors Bioreactor, scale: 3 L, [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Materials#Autoinduction_medium autoinduction medium] + 60 µg/mL chloramphenicol, 37 °C, pH 7, agitation on cascade to hold pO<sub>2</sub> at 50 %, OD<sub>600</sub> measured every 30 minutes.]] | ||
| + | |||
| + | <p align="justify"> | ||
| + | After the positive measurement of activity of ECOL we made a scale-up and fermented ''E. coli'' KRX with <partinfo>BBa_K863005</partinfo> in an Infors Labfors fermenter with a total volume of 3 L. Agitation speed, pO<sub>2</sub> and OD<sub>600</sub> were determined and illustrated in Figure 1. The exponential phase started after 1.5 hours of cultivation. The cell growth caused a decrease in pO<sub>2</sub>. After 2 hours of cultivation the agitation speed increased up to 629 rmp (5.9 hours) to hold the minimal pO<sub>2</sub> level of 50 %. Then, after 4 hours there was a break in cell growth due to induction of protein expression. The maximal OD<sub>600</sub> of 2.78 was reached after 5 hours. In comparison to ''E. coli'' KRX (OD<sub>600,max</sub> =4.86 after 8.5 hours) and to ''E. coli'' KRX with <partinfo>BBa_K863000</partinfo> (OD<sub>600,max</sub> =3.53 after 10 hours, time shift due to long lag phase) the OD<sub>600 max</sub> is lower. In the following hours, the OD<sub>600</sub> and the agitation speed decreased and the pO<sub>2</sub> increased, which indicates the death phase of the cells. This is caused by the cell toxicity of ECOL (reference: [http://www.dbu.de/OPAC/ab/DBU-Abschlussbericht-AZ-13191.pdf DBU final report]). Hence, cells were harvested after 12 hours. | ||
| + | </p> | ||
| + | |||
| + | |||
| + | <br style="clear: both" /> | ||
| + | |||
| + | ===Purification of ECOL=== | ||
| + | |||
| + | <p align="justify"> | ||
| + | The harvested cells were resuspended in [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA- equilibration buffer], mechanically disrupted by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Production#Mechanical_lysis_of_the_.28bio-reactor.29_cultivation homogenization] and cell debris were removed by centrifugation. The supernatant of the cell lysate was loaded on the Ni-NTA column (15 mL Ni-NTA resin) with a flow rate of 1 mL min<sup>-1</sup> cm<sup>-2</sup>. Then the column was washed with 10 column volumes (CV) [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA equilibration buffer]. The bound proteins were eluted by an increasing [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA elution buffer] step elution from 5 % (equates to 25 mM imidazol) with a length of 50 mL, to 50 % (equates to 250 mM imidazol) with a length of 60 mL, to 80 % (equates to 400 mM imidazol) with a length of 40 mL and finally to 100 % (equates to 500 mM imidazol) with a length of 80 mL. This strategy was chosen to improve the purification caused by a step by step increasing Ni-NTA-elution buffer concentration. The elution was collected in 10 mL fractions. Due to the high UV-detection signal of the loaded samples and to simplify the illustration of the detected product peak only the UV-detection signal of the wash step and the elution are shown. A typical chromatogram of purified laccases is illustrated [https://static.igem.org/mediawiki/2012/4/49/Bielefeld2012_Chromatogram_examplegrafik.jpg here]. The chromatogram of the ECOL elution is shown in Figure 2: | ||
| + | </p> | ||
| + | |||
| + | [[Image:Bielefeld2012_ECOL3LChromatogramm.jpg|450px|thumb|left|'''Figure 2:''' Chromatogram of wash and elution fractions from FLPC Ni-NTA His tag Purification of ECOL produced by 3 L fermentation of ''E. coli'' KRX with <partinfo>BBa_K863005</partinfo>. ECOL was eluted by a concentration of 50 % (equates to 250 mM imidazol) with a maximal UV-detection signal of 292 mAU. ]] | ||
| + | |||
| + | <p align="justify"> | ||
| + | The chromatogram shows two distinguished peaks. The first peak was detected at a [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA-equilibration buffer] concentration of 5 % (equates to 25 mM imidazol) and resulted from the elution of weakly bound proteins. After increasing the Ni-NTA elution buffer concentration to 50 % (equates to 250 mM imidazol), an UV-detection signal peak of 292 mAU was measured. The area of this peak indicates that a high amount of protein was eluted. The corresponding fractions were analyzed by SDS-PAGE to detect ECOL. There were no further peaks detectable. The following increasing UV detection signal results from the rising imidazol concentration of the Ni-NTA elution buffer. The corresponding SDS-PAGES are shown in Figure 3. | ||
| + | </p> | ||
| + | |||
| + | <br style="clear: both" /> | ||
| + | |||
| + | ===SDS-PAGE of ECOL purification=== | ||
| + | |||
| + | [[Image:Bielefeld2012_SDS_ECOL3L.jpg|450px|thumb|left|'''Figure 3:''' SDS-Pages of purified ''E. coli'' KRX containing [https://parts.igem.org/wiki/index.php?title=Part:BBa_K863005 BBa_K863005] lysate (fermented in 3 L an Infors Labfors fermenter). The flow-through and elution fraction 2-9 are shown. The arrow marks the ECOL band with a molecular weight of 53.4 kDa.]] | ||
| + | <p align="justify"> | ||
| + | In Figure 3 the SDS-PAGE of the Ni-NTA His tag purification of the lysed culture (''E. coli'' KRX containing [https://parts.igem.org/wiki/index.php?title=Part:BBa_K863005 BBa_K863005]) is shown including the flow-through and the fractions 2 to 9. The red arrow indicates the band of ECOL with a molecular weight of 53.4 kDa, which appears in all fractions. The strongest bands appear in fractions 6 and 7. These were the first two fractions (each 10 mL) eluted with 50 % Ni-NTA elution buffer (equates to 250 mM imidazol), in which the distinguished peak appeared. | ||
| + | |||
| + | These bands were analyzed by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#MALDI MALDI-TOF] and identified as CueO (ECOL). In contrast, the second, faint band with a lower molecular weight could not be identified. | ||
| + | <br style="clear: both" /> | ||
| + | </p> | ||
| + | |||
| + | ===6 L Fermentation of ''E. coli'' KRX with <partinfo>BBa_K863005</partinfo>=== | ||
| + | |||
| + | [[Image:Bielefeld2012_ECOL6LFermentation.jpg|450px|thumb|left|'''Figure 4:''' Fermentation of ''E. coli'' KRX with <partinfo>BBa_K863005</partinfo> (ECOL) in a Bioengineering NFL22 fermenter, scale: 6 L, [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Materials#Autoinduction_medium autoinduction medium] + 60 µg/mL chloramphenicol, 37 °C, pH 7, agitation increased when pO<sub>2</sub> was below 30 %, OD<sub>600</sub> taken every hour.]] | ||
| + | |||
| + | |||
| + | <p align="justify"> | ||
| + | Another scale-up of the fermentation of E. coli KRX with <partinfo>BBa_K863005</partinfo> was made up to a final working volume of 6 L in a Bioengineering NFL 22 fermenter. Agitation speed, pO<sub>2</sub> and OD<sub>600</sub> were determined and illustrated in Figure 3. There was no noticeable lag phase and the cells immediately began to grow. The cells were in an exponential phase between 2 and 4 hours of cultivation, which results in a decrease of pO<sub>2</sub> value and therefore in an increase of agitation speed. After 4 hours of cultivation the maximal OD<sub>600</sub> of 2.76 was reached, which is comparable to the 3 L fermentation of ''E. coli'' KRX with <partinfo>BBa_K863005</partinfo>. Due to induction of protein expression there is a break in cell growth. The death phase started, which is indicated by an increasing pO<sub>2</sub> and a decreasing OD<sub>600</sub>. This demonstrates the cytotoxicity of the laccase for ''E. coli'', which was reported by the [http://www.dbu.de/OPAC/ab/DBU-Abschlussbericht-AZ-13191.pdf DBU]. In comparison to the fermentation of ''E. coli'' KRX with <partinfo>BBa_K863000</partinfo> under the same conditions (OD<sub>600,max</sub>= 3.53), the OD<sub>600,max</sub> was lower. Cells were harvested after 12 hours. | ||
| + | </p> | ||
| + | |||
| + | <br style="clear: both" /> | ||
| + | |||
| + | ===Purification of ECOL=== | ||
| + | |||
| + | <p align="justify"> | ||
| + | The harvested cells were resuspended in [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA-equilibration buffer], mechanically disrupted by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Production#Mechanical_lysis_of_the_.28bio-reactor.29_cultivation homogenization] and cell debris were removed by centrifugation. The supernatant of the cell lysate was loaded on the Ni-NTA column (15 mL Ni-NTA resin) with a flow rate of 1 mL min<sup>-1</sup> cm<sup>-2</sup>. The column was washed by 10 column volumes (CV) [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA- equilibration buffer]. The bound proteins were eluted by an increasing [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA- elution buffer] gradient from 0 % to 100 % with a length of 200 mL and the elution was collected in 10 mL fractions. Due to the high UV-detection signal of the loaded samples and to simplify the illustration of the detected product peak only the UV-detection signal of the wash step and the eluate are shown. A typical chromatogram of purified laccases is shown [https://static.igem.org/mediawiki/2012/4/49/Bielefeld2012_Chromatogram_examplegrafik.jpg here]. The chromatogram of the ECOL elution is shown in Figure 5: | ||
| + | </p> | ||
| + | |||
| + | [[Image:Bielefeld2012_ECOL6LChromatogramm.jpg|450px|thumb|left|'''Figure 5:''' Chromatogram of wash and elution from FLPC Ni-NTA His tag purification of ECOL produced by 3 L fermentation of ''E. coli'' KRX with <partinfo>BBa_K863005</partinfo>. ECOL was eluted between a process volume 670 mL to 750 mL with a maximal UV-detection signal of 189 mAU.]] | ||
| + | |||
| + | |||
| + | <p align="justify"> | ||
| + | After washing the column with 10 CV [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA-elution buffer] the elution process was started. At a process volume of 670 mL to 750 mL the chromatogram shows a remarkable widespread peak (UV-detection signal 189 mAU) caused by the elution of a high amount of proteins. The run of the curve show a fronting. This can be explained by the elution of weakly bound proteins, which elutes at low imidazol concentrations. A better result could be achieved with a step elution strategy ([http://2012.igem.org/Team:Bielefeld-Germany/Results/Summary#Purification_of_ECOL see purification of the 3 L Fermentation above]). To detect ECOL the corresponding fractions were analyzed by SDS-PAGE. | ||
| + | </p> | ||
| + | <br style="clear: both" /> | ||
| + | |||
| + | ===SDS-PAGES of ECOL purification=== | ||
| + | |||
| + | [[Image:Bielefeld2012_coli0910.jpg|450px|thumb|left|'''Figure 6:''' SDS-Pages of lysed ''E. coli'' KRX culture containing [https://parts.igem.org/wiki/index.php?title=Part:BBa_K863005 BBa_K863005] (fermented in a 6 L Bioengineering NFL22) after purification. The flow-through, wash and the elution fraction 1 to 15 are shown (except from fraction 11/12). The arrow marks the ECOL band with a molecular weight of 53.4 kDa.]] | ||
| + | |||
| + | <p align="justify"> | ||
| + | In Figure 6 the SDS-PAGE of the Ni-NTA His tag purification of the lysed culture ''E. coli'' KRX containing [https://parts.igem.org/wiki/index.php?title=Part:BBa_K863005 BBa_K863005] (6 L fermentation) including the flow-through, wash and the fractions 1 to 15 (except from fraction 11/12) is shown. The red arrow indicates the band of ECOL with a molecular weight of 53.4 kDa, which appears in all fractions. The strongest bands appear from fractions 3 and 8 with a decreasing amount of other non-specific bands. In summary, the scale up was successful, improving protein production and purification once again. | ||
Revision as of 02:55, 27 September 2012
ecol laccase from E. coli
E.coli laccase ORF
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 225
- 1000COMPATIBLE WITH RFC[1000]
First some trials of shaking flask cultivations were made with changing parameters to identify the best conditions for the production of the laccase CueO from E. coli BL21 (DE3) named ECOL fused to a His tag. Because of no measured activity in the cell lysate a purification method was established (using Ni-NTA His tag resin and Syringe or ÄKTA method). The purified ECOL could be identified by SDS-PAGE (molecular weight of 53.4 kDa) as well as by MALDI-TOF. The fractionated samples were also tested concerning their activity. A maximal activity of X was reached. After measuring activity of ECOL a scale up was made up to 3 L and then also up to 6 L that enables an intense screening afterwards. The scale up will be important for further application.
Contents
Cultivation, Purification and SDS-PAGE
Shaking Flask Cultivations
The first trials to produce ECOL were produced in shaking flask with various designs (from 100 mL-1 to 1 L flasks, with and without baffles) and under different conditions. The parameters tested during our screening experiments were temperature (27 °C,30 °C and 37 °C), concentrations of chloramphenicol (20-170 µg mL-1), various induction strategies ([http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Materials#Autoinduction_medium autoinduction] and manual induction) and cultivation time (6 - 24 h). Furthermore it was cultivated with and without 0.25 mM CuCl2 to provide a sufficient amount of copper, which is needed for the active center of the laccase. Based on the screening experiments we identified the best conditions under which ECOL was expressed. The addition of CuCl2 did not increase the activity, so it was omitted.
- flask design: shaking flask without baffles
- medium: [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Materials#Autoinduction_medium autoinduction medium]
- antibiotics: 60 µg mL-1 chloramphenicol
- temperature: 37 °C
- cultivation time: 12 h
The reproducibility of the measured data and results were investigated for the shaking flask and bioreactor cultivation.
3 L Fermentation E. coli KRX with BBa_K863005
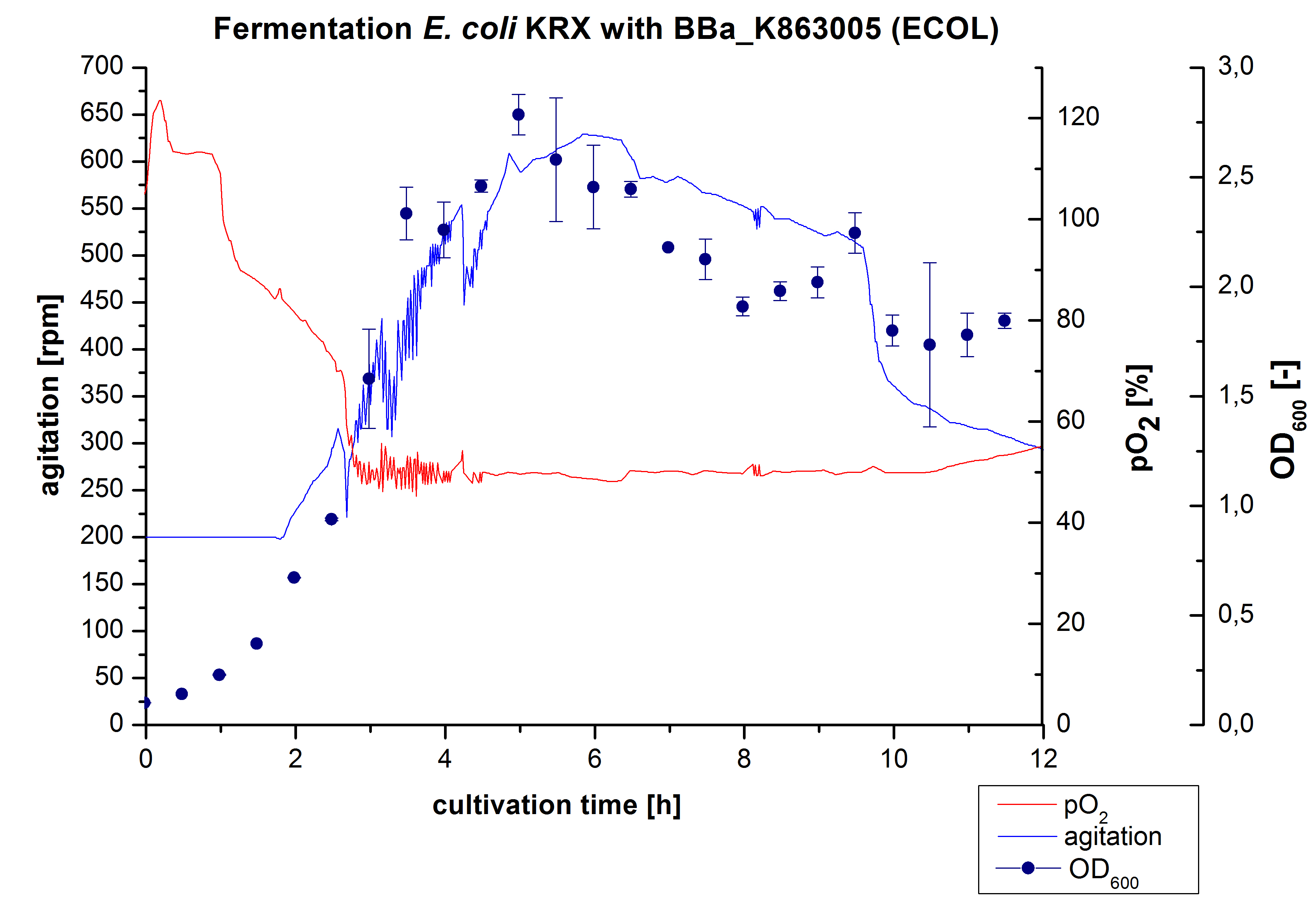
After the positive measurement of activity of ECOL we made a scale-up and fermented E. coli KRX with BBa_K863005 in an Infors Labfors fermenter with a total volume of 3 L. Agitation speed, pO2 and OD600 were determined and illustrated in Figure 1. The exponential phase started after 1.5 hours of cultivation. The cell growth caused a decrease in pO2. After 2 hours of cultivation the agitation speed increased up to 629 rmp (5.9 hours) to hold the minimal pO2 level of 50 %. Then, after 4 hours there was a break in cell growth due to induction of protein expression. The maximal OD600 of 2.78 was reached after 5 hours. In comparison to E. coli KRX (OD600,max =4.86 after 8.5 hours) and to E. coli KRX with BBa_K863000 (OD600,max =3.53 after 10 hours, time shift due to long lag phase) the OD600 max is lower. In the following hours, the OD600 and the agitation speed decreased and the pO2 increased, which indicates the death phase of the cells. This is caused by the cell toxicity of ECOL (reference: [http://www.dbu.de/OPAC/ab/DBU-Abschlussbericht-AZ-13191.pdf DBU final report]). Hence, cells were harvested after 12 hours.
Purification of ECOL
The harvested cells were resuspended in [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA- equilibration buffer], mechanically disrupted by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Production#Mechanical_lysis_of_the_.28bio-reactor.29_cultivation homogenization] and cell debris were removed by centrifugation. The supernatant of the cell lysate was loaded on the Ni-NTA column (15 mL Ni-NTA resin) with a flow rate of 1 mL min-1 cm-2. Then the column was washed with 10 column volumes (CV) [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA equilibration buffer]. The bound proteins were eluted by an increasing [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA elution buffer] step elution from 5 % (equates to 25 mM imidazol) with a length of 50 mL, to 50 % (equates to 250 mM imidazol) with a length of 60 mL, to 80 % (equates to 400 mM imidazol) with a length of 40 mL and finally to 100 % (equates to 500 mM imidazol) with a length of 80 mL. This strategy was chosen to improve the purification caused by a step by step increasing Ni-NTA-elution buffer concentration. The elution was collected in 10 mL fractions. Due to the high UV-detection signal of the loaded samples and to simplify the illustration of the detected product peak only the UV-detection signal of the wash step and the elution are shown. A typical chromatogram of purified laccases is illustrated here. The chromatogram of the ECOL elution is shown in Figure 2:
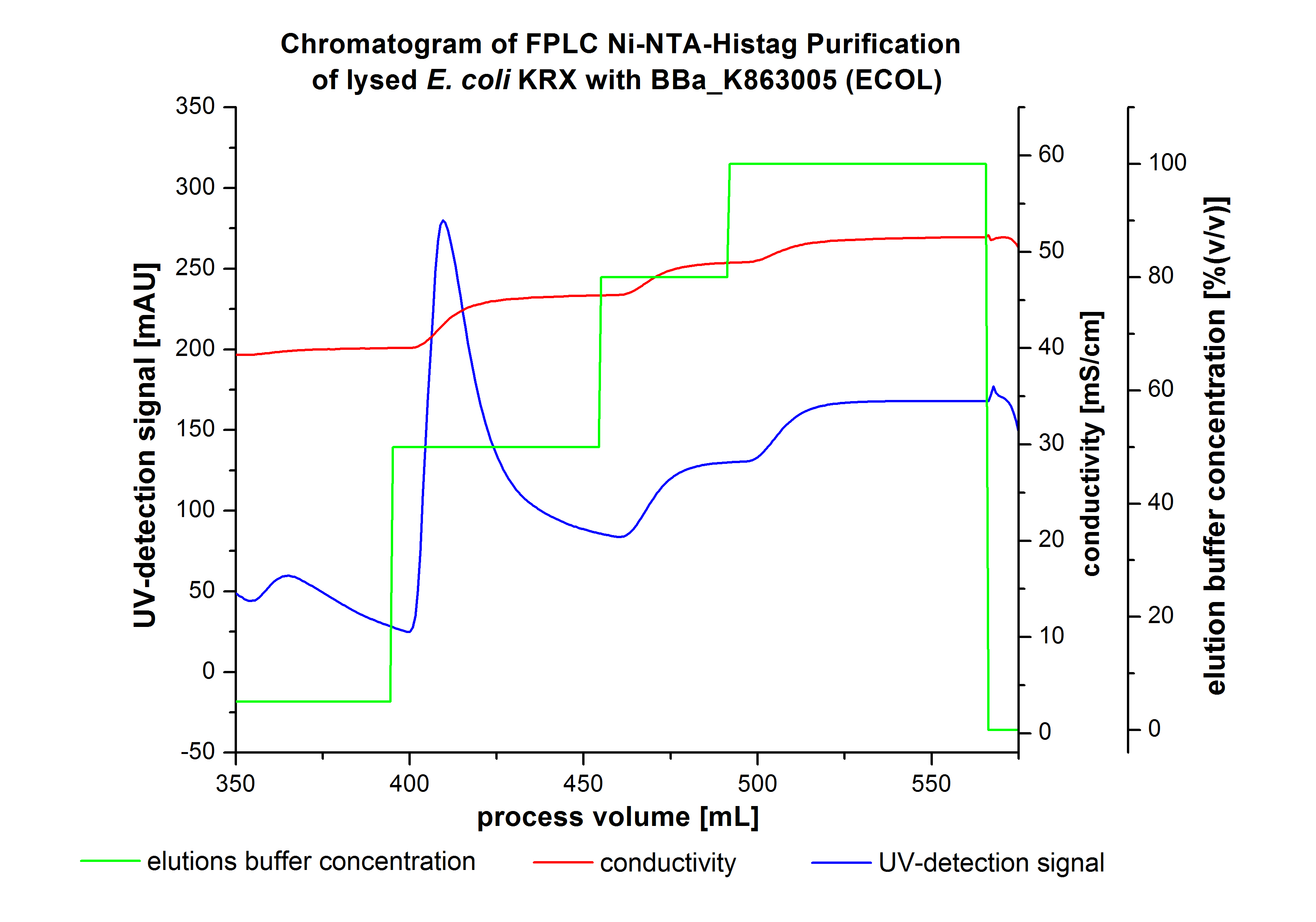
The chromatogram shows two distinguished peaks. The first peak was detected at a [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA-equilibration buffer] concentration of 5 % (equates to 25 mM imidazol) and resulted from the elution of weakly bound proteins. After increasing the Ni-NTA elution buffer concentration to 50 % (equates to 250 mM imidazol), an UV-detection signal peak of 292 mAU was measured. The area of this peak indicates that a high amount of protein was eluted. The corresponding fractions were analyzed by SDS-PAGE to detect ECOL. There were no further peaks detectable. The following increasing UV detection signal results from the rising imidazol concentration of the Ni-NTA elution buffer. The corresponding SDS-PAGES are shown in Figure 3.
SDS-PAGE of ECOL purification
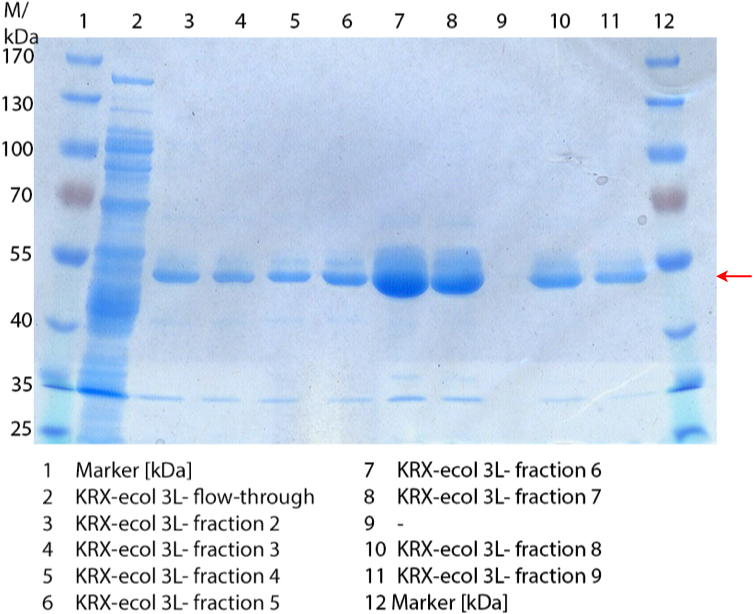
In Figure 3 the SDS-PAGE of the Ni-NTA His tag purification of the lysed culture (E. coli KRX containing BBa_K863005) is shown including the flow-through and the fractions 2 to 9. The red arrow indicates the band of ECOL with a molecular weight of 53.4 kDa, which appears in all fractions. The strongest bands appear in fractions 6 and 7. These were the first two fractions (each 10 mL) eluted with 50 % Ni-NTA elution buffer (equates to 250 mM imidazol), in which the distinguished peak appeared.
These bands were analyzed by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#MALDI MALDI-TOF] and identified as CueO (ECOL). In contrast, the second, faint band with a lower molecular weight could not be identified.
6 L Fermentation of E. coli KRX with BBa_K863005
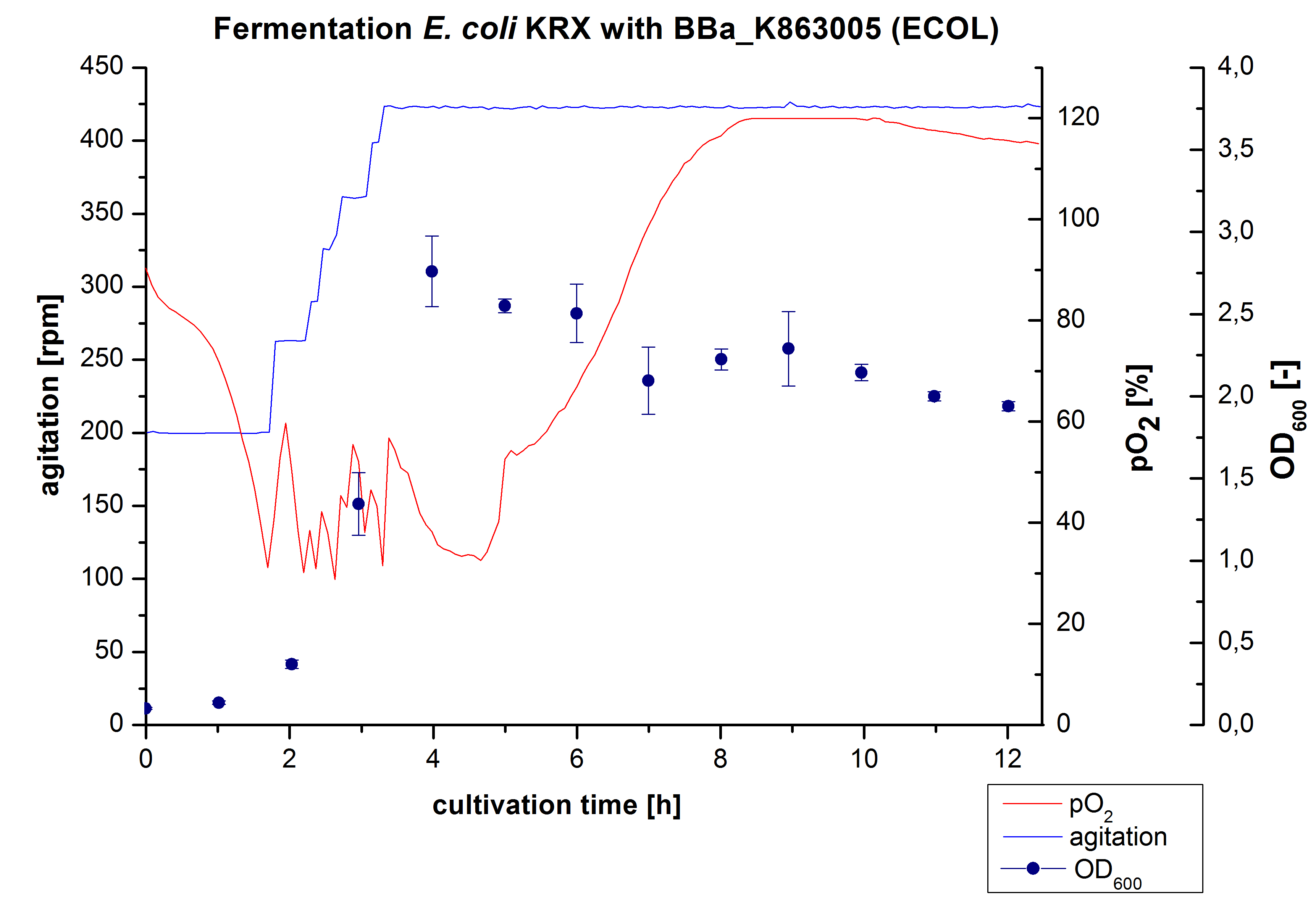
Another scale-up of the fermentation of E. coli KRX with BBa_K863005 was made up to a final working volume of 6 L in a Bioengineering NFL 22 fermenter. Agitation speed, pO2 and OD600 were determined and illustrated in Figure 3. There was no noticeable lag phase and the cells immediately began to grow. The cells were in an exponential phase between 2 and 4 hours of cultivation, which results in a decrease of pO2 value and therefore in an increase of agitation speed. After 4 hours of cultivation the maximal OD600 of 2.76 was reached, which is comparable to the 3 L fermentation of E. coli KRX with BBa_K863005. Due to induction of protein expression there is a break in cell growth. The death phase started, which is indicated by an increasing pO2 and a decreasing OD600. This demonstrates the cytotoxicity of the laccase for E. coli, which was reported by the [http://www.dbu.de/OPAC/ab/DBU-Abschlussbericht-AZ-13191.pdf DBU]. In comparison to the fermentation of E. coli KRX with BBa_K863000 under the same conditions (OD600,max= 3.53), the OD600,max was lower. Cells were harvested after 12 hours.
Purification of ECOL
The harvested cells were resuspended in [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA-equilibration buffer], mechanically disrupted by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Production#Mechanical_lysis_of_the_.28bio-reactor.29_cultivation homogenization] and cell debris were removed by centrifugation. The supernatant of the cell lysate was loaded on the Ni-NTA column (15 mL Ni-NTA resin) with a flow rate of 1 mL min-1 cm-2. The column was washed by 10 column volumes (CV) [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA- equilibration buffer]. The bound proteins were eluted by an increasing [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA- elution buffer] gradient from 0 % to 100 % with a length of 200 mL and the elution was collected in 10 mL fractions. Due to the high UV-detection signal of the loaded samples and to simplify the illustration of the detected product peak only the UV-detection signal of the wash step and the eluate are shown. A typical chromatogram of purified laccases is shown here. The chromatogram of the ECOL elution is shown in Figure 5:
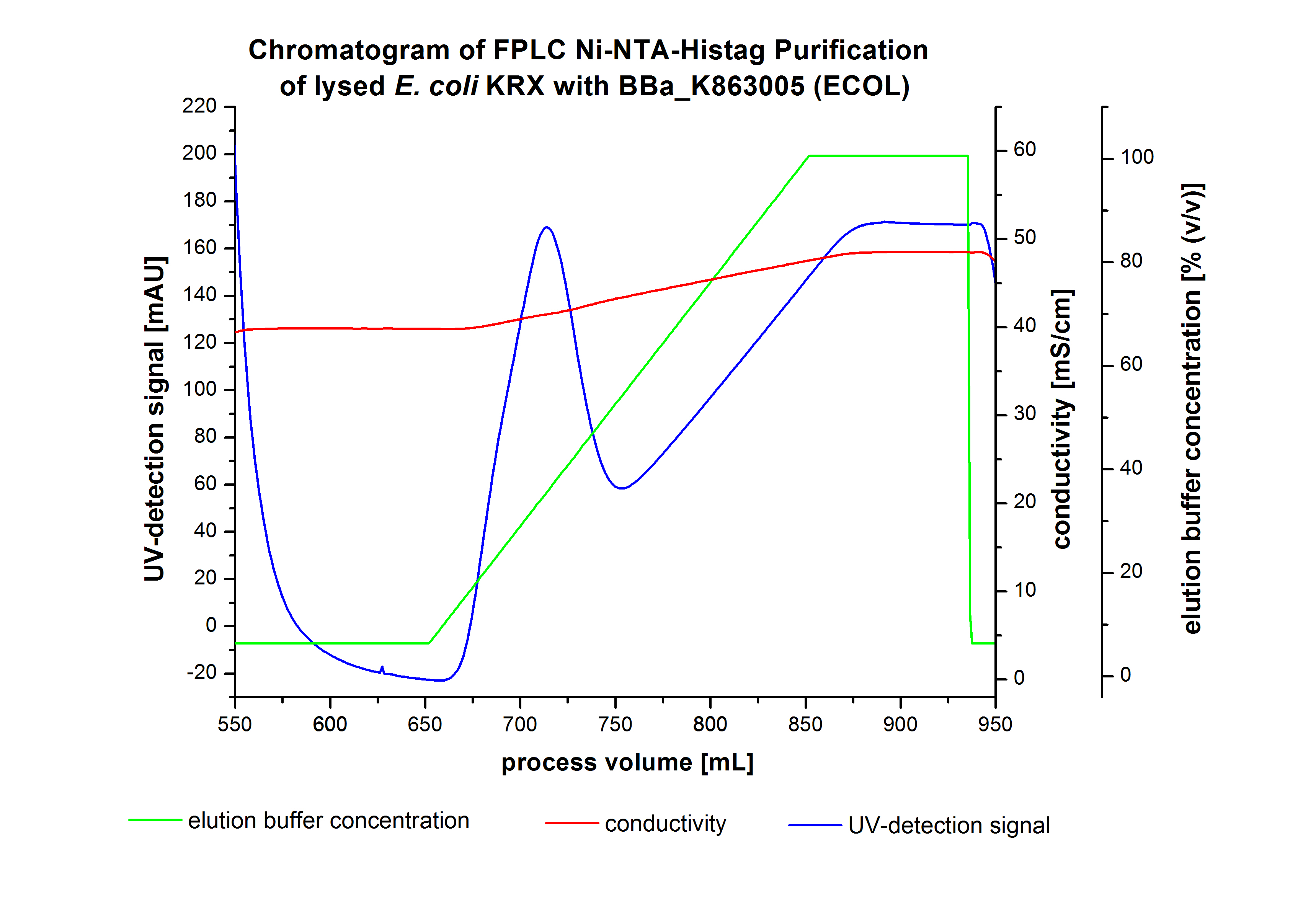
After washing the column with 10 CV [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA-elution buffer] the elution process was started. At a process volume of 670 mL to 750 mL the chromatogram shows a remarkable widespread peak (UV-detection signal 189 mAU) caused by the elution of a high amount of proteins. The run of the curve show a fronting. This can be explained by the elution of weakly bound proteins, which elutes at low imidazol concentrations. A better result could be achieved with a step elution strategy ([http://2012.igem.org/Team:Bielefeld-Germany/Results/Summary#Purification_of_ECOL see purification of the 3 L Fermentation above]). To detect ECOL the corresponding fractions were analyzed by SDS-PAGE.
SDS-PAGES of ECOL purification
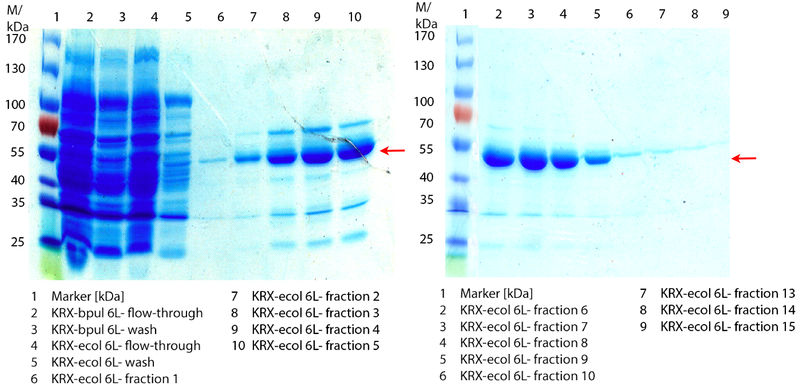
In Figure 6 the SDS-PAGE of the Ni-NTA His tag purification of the lysed culture E. coli KRX containing BBa_K863005 (6 L fermentation) including the flow-through, wash and the fractions 1 to 15 (except from fraction 11/12) is shown. The red arrow indicates the band of ECOL with a molecular weight of 53.4 kDa, which appears in all fractions. The strongest bands appear from fractions 3 and 8 with a decreasing amount of other non-specific bands. In summary, the scale up was successful, improving protein production and purification once again.
