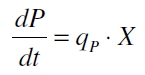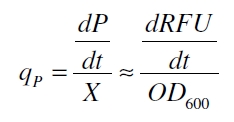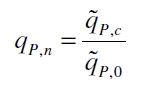Difference between revisions of "Part:BBa K389016:Experience"
(→Applications of BBa_K389016) |
|||
| Line 5: | Line 5: | ||
===Applications of BBa_K389016=== | ===Applications of BBa_K389016=== | ||
| − | ====Transfer function of <partinfo>K389016<partinfo>==== | + | ====Transfer function of <partinfo>K389016</partinfo>==== |
| − | + | ||
The data for the transfer function was measured and analyzed as described below. The data was fitted with a dose response function of the form | The data for the transfer function was measured and analyzed as described below. The data was fitted with a dose response function of the form | ||
| Line 13: | Line 12: | ||
| − | with the hill coefficient p, the bottom asymptote A1, the top asymptote A2 and the inflection point log(x<sub>0</sub>). | + | with the hill coefficient p, the bottom asymptote A1, the top asymptote A2 and the inflection point log(x<sub>0</sub>). Figure 1 shows the measured normalized specific production rates q<sub>P,n</sub> plotted against the logarithm of the concentration of the inductor acetosyringone in µM. The fit has an R<sup>2</sup> = 0.99. |
| − | [[Image:Bielefeld_Final_RFP_fit.jpg|600px|thumb|center|Transfer function for the part <partinfo>K389016</partinfo> (R<sup>2</sup> = 0.99).]] | + | [[Image:Bielefeld_Final_RFP_fit.jpg|600px|thumb|center|'''Fig. 1: Transfer function for the part <partinfo>K389016</partinfo> (R<sup>2</sup> = 0.99).''']] |
| − | The important data from the transfer function is summarized in | + | The important data from the transfer function is summarized in table 1: |
<center> | <center> | ||
| Line 58: | Line 57: | ||
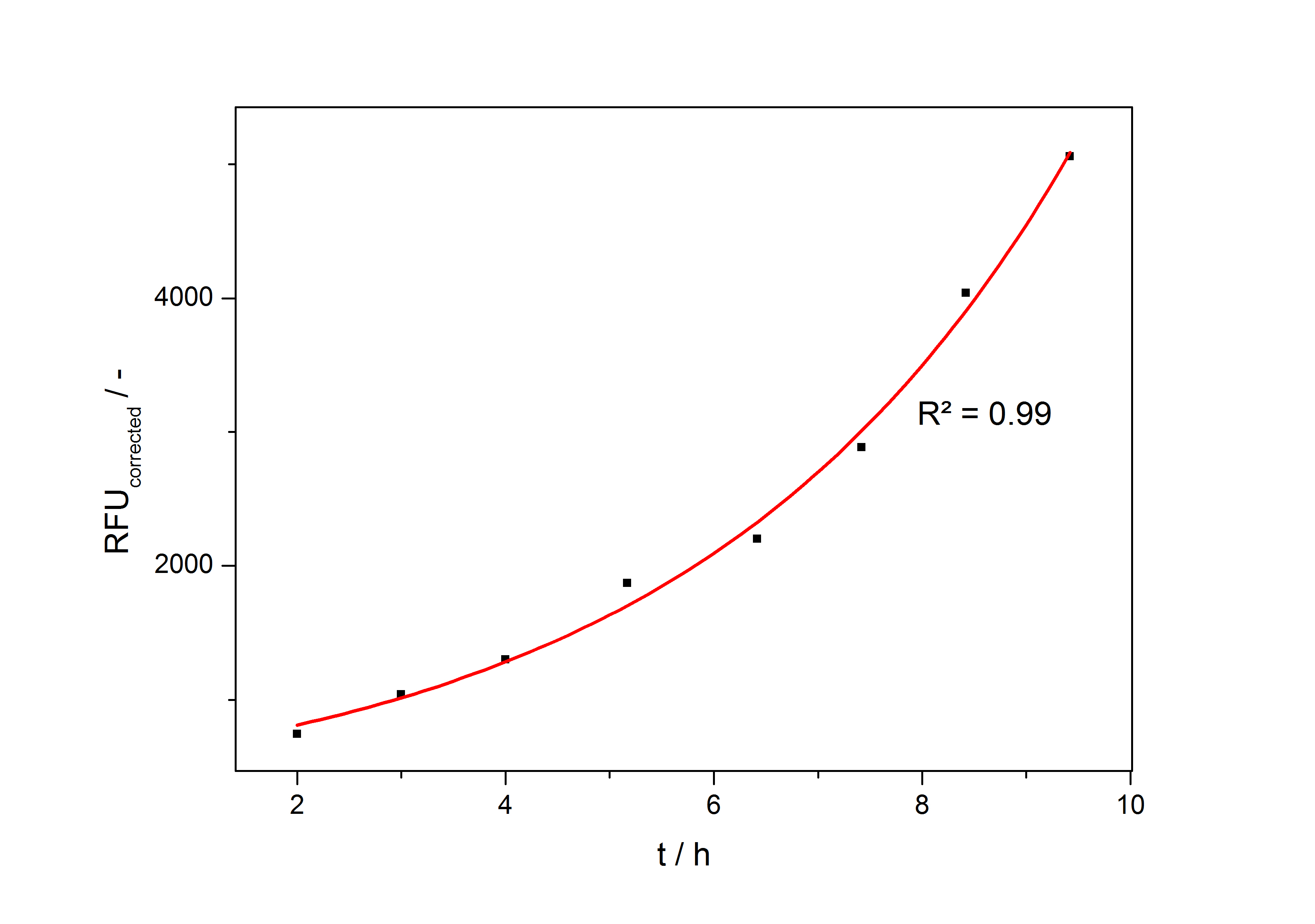
The accumulation of mRFP in the cells is always exponential. A typical fitted product accumulation curve is shown below: | The accumulation of mRFP in the cells is always exponential. A typical fitted product accumulation curve is shown below: | ||
| − | [[Image:Bielefeld_ExpFit_auf_RFU.jpg|500px|thumb|center|'''Exponential fit on the measured RFU plotted against cultivation time of a cultivation of <partinfo>K389016</partinfo> in ''Escherichia coli'' DB3.1 in LB medium with 10 mg ml<sup>-1</sup> chloramphenicol and 150 µM acetosyringone. ''']] | + | [[Image:Bielefeld_ExpFit_auf_RFU.jpg|500px|thumb|center|'''Fig. 2: Exponential fit on the measured RFU plotted against cultivation time of a cultivation of <partinfo>K389016</partinfo> in ''Escherichia coli'' DB3.1 in LB medium with 10 mg ml<sup>-1</sup> chloramphenicol and 150 µM acetosyringone. ''']] |
The product accumulation in a cultivation can be described as: | The product accumulation in a cultivation can be described as: | ||
Revision as of 15:23, 25 October 2010
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_K389016
Transfer function of BBa_K389016
The data for the transfer function was measured and analyzed as described below. The data was fitted with a dose response function of the form
with the hill coefficient p, the bottom asymptote A1, the top asymptote A2 and the inflection point log(x0). Figure 1 shows the measured normalized specific production rates qP,n plotted against the logarithm of the concentration of the inductor acetosyringone in µM. The fit has an R2 = 0.99.
The important data from the transfer function is summarized in table 1:
Table 1: Data from the transfer function for the part BBa_K389016.
| Parameter | Value |
|---|---|
| Hill coefficient | 1.673 |
| Inflection point | 26.5 µM |
| Top asymptote | 2.62 |
Data analysis for BBa_K389016
The data analysis is made in three steps. First step is the processing of the fluorescence raw data gained by the fluorescence plate reader for every sample:
In the second step the RFUcorrected of every sample is plotted against the cultivation time it was drawn. The data is fitted by an exponential fit of the following style:
The accumulation of mRFP in the cells is always exponential. A typical fitted product accumulation curve is shown below:
The product accumulation in a cultivation can be described as:
with the amount of product P, the cell count X and the specific production rate qP.
RFU is commensurate to the concentration of mRFP (P) and the OD600 is commensurate to the cell count (X) ([http:/parts.igem.org/Part:BBa_F2620:Experience/Endy/Data_analysis Canton and Labno, 2004]):
With these assumptions it is possible to calculate the specific production rate of mRFP qP in the third step: the specific production rate for every sample of a cultivation is calculated by the derivation of the exponential fit line which describes the accumulation of product in the culture (dRFU/dt) and the measured OD600 data:
The specific production rates qP of all samples of all cultivations made with a specific inductor concentration c are averaged and normalized against the specific production rate of the uninduced system qP,0:
This normalized specific production rate we calculated is commensurate to relative promotor units (RPU) which is commensurate to PoPS (polymerase per seconds) (Canton and Labno, 2004; Pasotti et al., 2009):
References
Canton B and Labno A (2004) Data processing of Part BBa_F2620, https://parts.igem.org/Part:BBa_F2620:Experience/Endy/Data_analysis.
Pasotti L, Zucca S, Del Fabbro E (2009) Characterization experiment on BBa_J23100, BBa_J23101, BBa_J23118, https://parts.igem.org/Part:BBa_J23101:Experience.
User Reviews
UNIQ3d87fdc987ded1c9-partinfo-00000005-QINU UNIQ3d87fdc987ded1c9-partinfo-00000006-QINU