Difference between revisions of "Part:BBa K525121"
(→Expression in E. coli) |
(→Expression in E. coli) |
||
| Line 43: | Line 43: | ||
The CspB|mRFP fusion protein was overexpressed in E. coli KRX after induction of T7 polymerase by supplementation of 0,1 % L-rhamnose using the [http://2011.igem.org/Team:Bielefeld-Germany/Protocols/Downstream-processing#Expression_of_S-layer_genes_in_E._coli autinduction protocol] from promega. | The CspB|mRFP fusion protein was overexpressed in E. coli KRX after induction of T7 polymerase by supplementation of 0,1 % L-rhamnose using the [http://2011.igem.org/Team:Bielefeld-Germany/Protocols/Downstream-processing#Expression_of_S-layer_genes_in_E._coli autinduction protocol] from promega. | ||
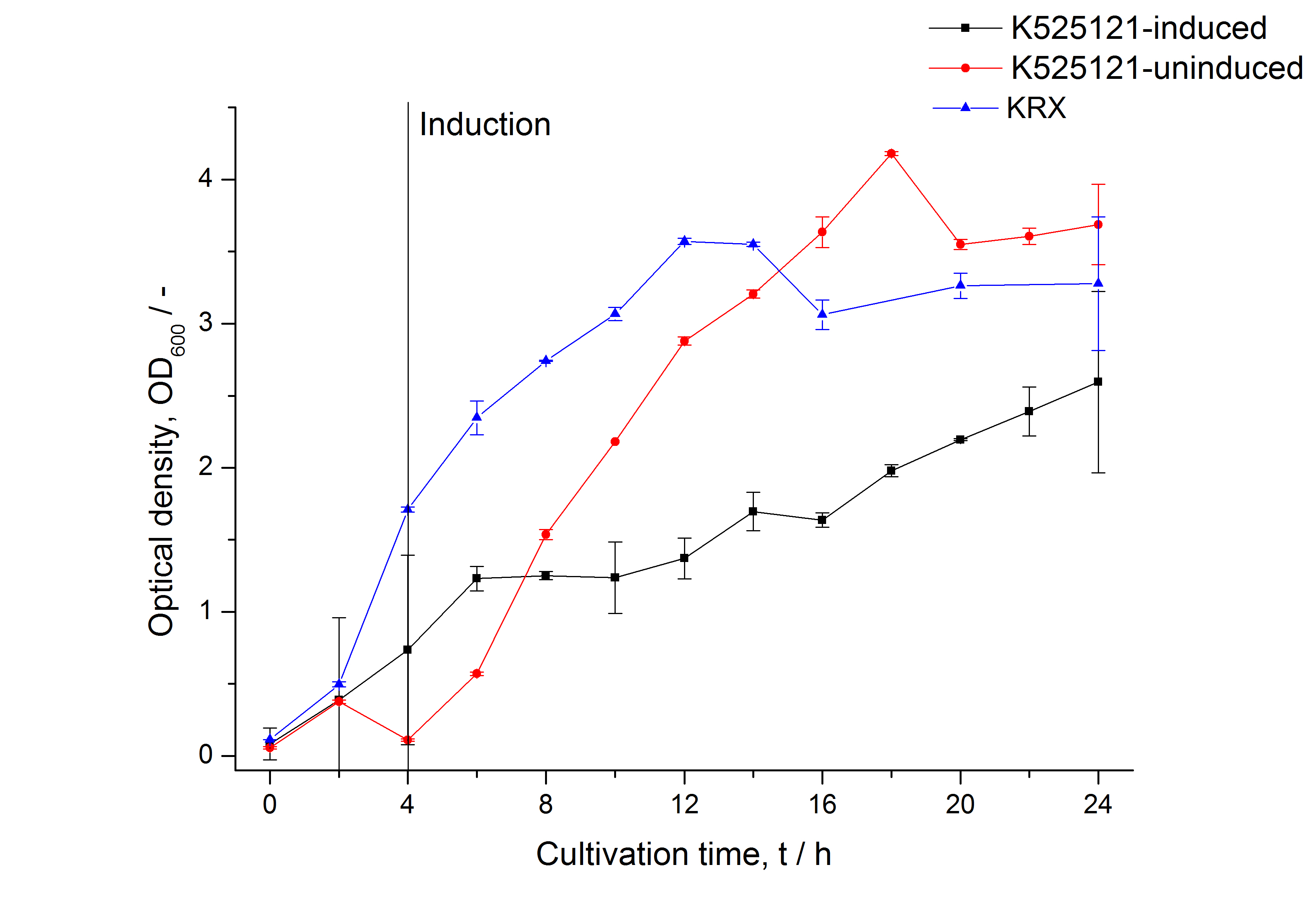
| − | [[Image:Bielefeld_2011_BF1_Growthcurve.png|600px|center|thumb| '''Figure 1: Growthcurve of ''E. coli'' KRX expressing the fusion protein of CspB and mRFP with and without induction. A curve depicting KRX wildtype is shown for comparsion.''']] | + | [[Image:Bielefeld_2011_BF1_Growthcurve.png|600px|center|thumb| '''Figure 1: Growthcurve of ''E. coli'' KRX expressing the fusion protein of CspB and mRFP with and without induction. A curve depicting KRX wildtype is shown for comparsion. Cultivations were carried out in autoinduction medium at 37 ˚C.''']] |
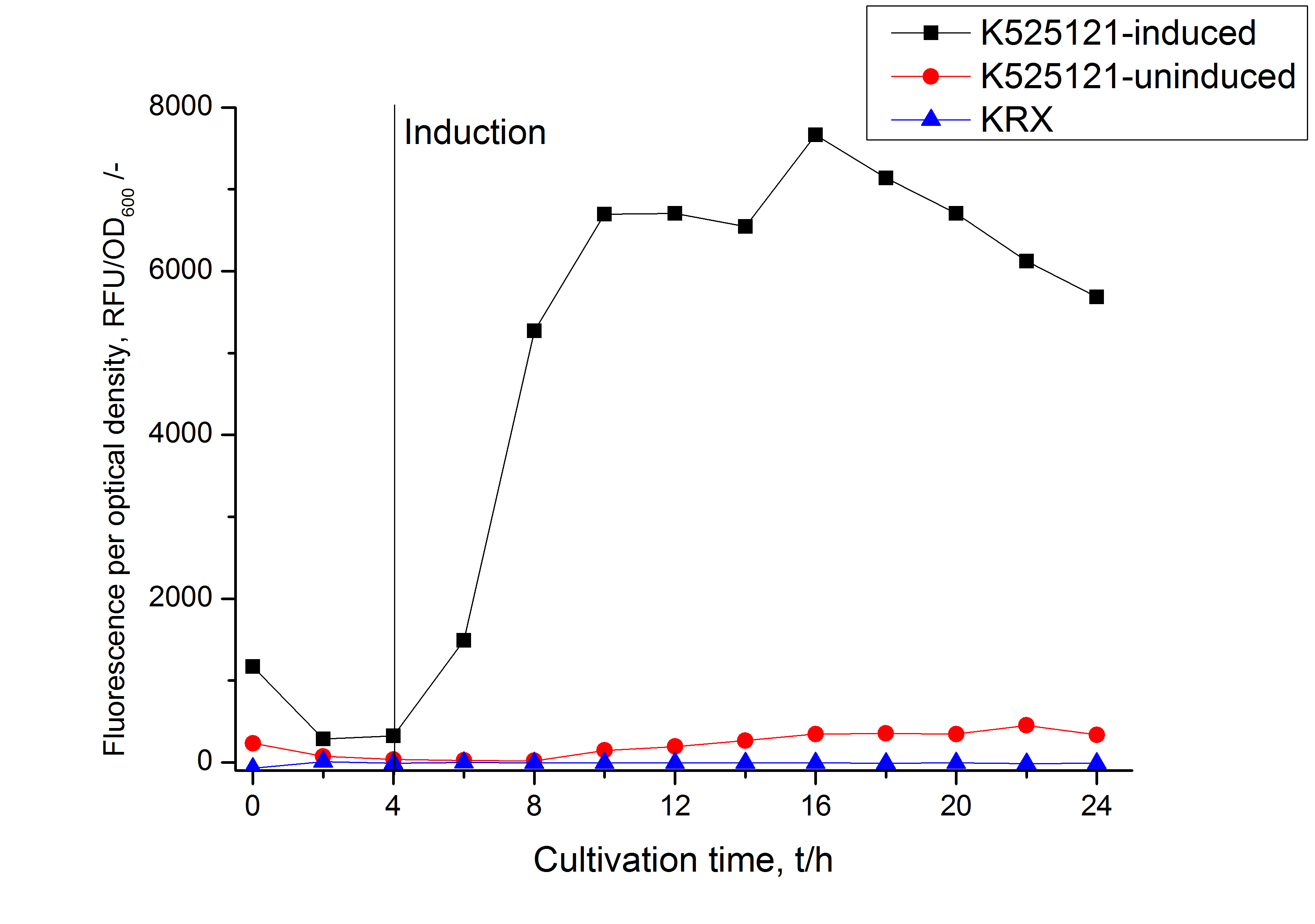
| − | [[Image:Bielefeld_2011_BF1_RFU_OD.png|600px|center|thumb| '''Figure 2: RFU to OD<sub>600</sub> ratio of ''E. coli'' KRX expressing the fusion protein of CspB and mRFP with and without induction. A curve depicting KRX wildtype is shown for comparsion.''']] | + | [[Image:Bielefeld_2011_BF1_RFU_OD.png|600px|center|thumb| '''Figure 2: RFU to OD<sub>600</sub> ratio of ''E. coli'' KRX expressing the fusion protein of CspB and mRFP with and without induction. A curve depicting KRX wildtype is shown for comparsion. Cultivations were carried out in autoinduction medium at 37 ˚C.''']] |
===Identification and localisation=== | ===Identification and localisation=== | ||
Revision as of 17:53, 21 September 2011
S-layer cspB from Corynebacterium glutamicum with TAT-Sequence and lipid anchor, PT7 and RBS
Usage and Biology
S-layer proteins can be used as scaffold for nanobiotechnological applications and devices by e.g. fusing the S-layer's self-assembly domain to other functional protein domains. It is possible to coat surfaces and liposomes with S-layers. A big advantage of S-layers: after expressing in E. coli and purification, the nanobiotechnological system is cell-free. This enhances the biological security of a device.
This fluorescent S-layer fusion protein is used to characterize purification methods and the S-layer's ability to self-assemble on surfaces.
Important parameters
| Experiment | Characteristic | Result |
|---|---|---|
| Expression (E. coli) | Localisation | cell membrane |
| Compatibility | E. coli KRX | |
| Inductor for expression | L-rhamnose for induction of T7 polymerase | |
| Characteristics | Molecular weight | |
| Theoretical pI |
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 1421
Illegal XhoI site found at 248
Illegal XhoI site found at 866 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 1400
Illegal SapI site found at 647
Illegal SapI site found at 859
Illegal SapI site found at 1407
Expression in E. coli
For characterization the CspB gen was fused with a monomeric RFP (BBa_E1010) using Gibson assembly.
The CspB|mRFP fusion protein was overexpressed in E. coli KRX after induction of T7 polymerase by supplementation of 0,1 % L-rhamnose using the [http://2011.igem.org/Team:Bielefeld-Germany/Protocols/Downstream-processing#Expression_of_S-layer_genes_in_E._coli autinduction protocol] from promega.
Identification and localisation
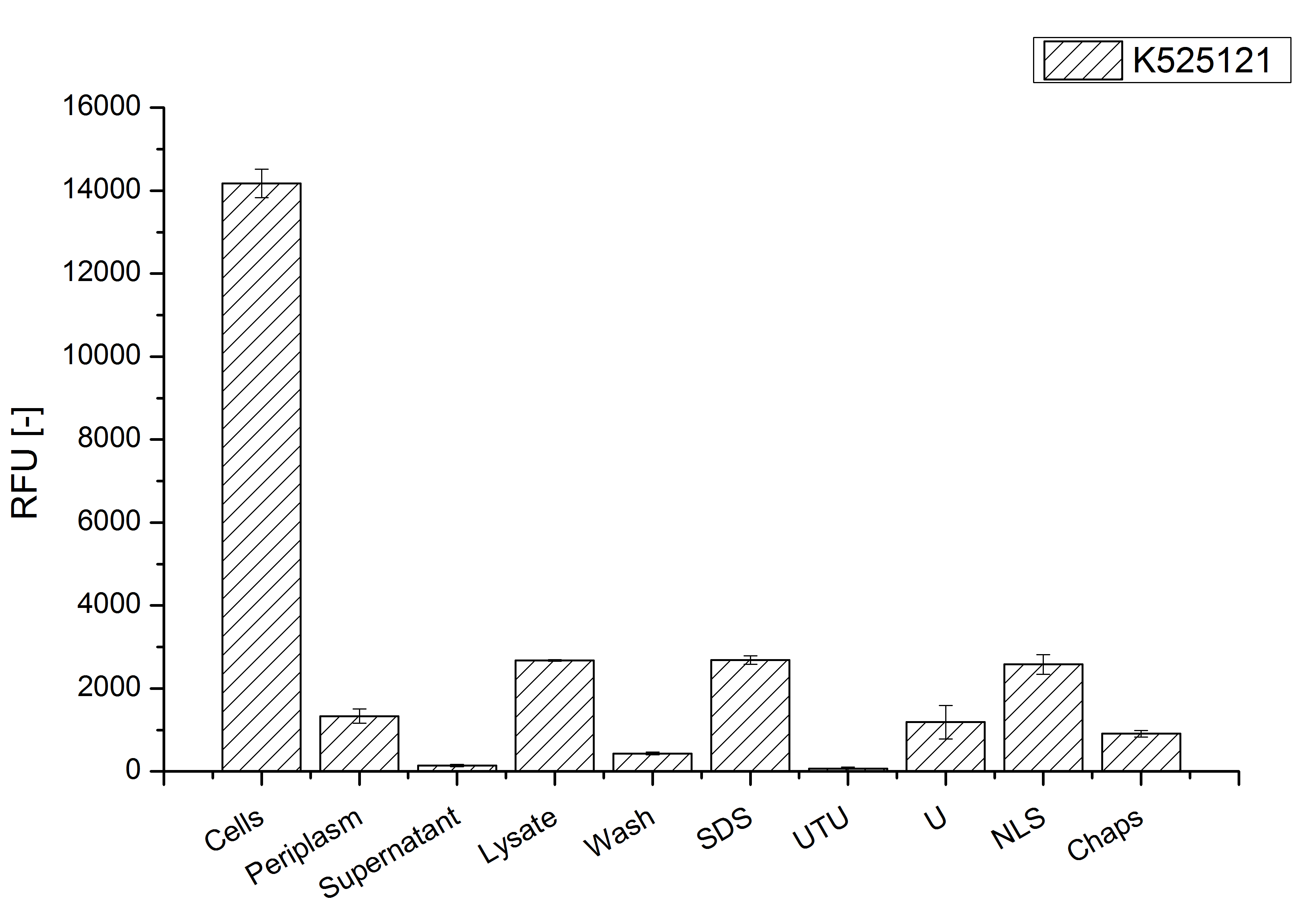
After a cultivation time of 18 h the CspB|mRFP fusion protein has to be localized in E. coli KRX. Therefor a part of the produced biomass was mechanically disrupted and the resulting lysate was wahed with ddH2O. From the other part the periplasm was detached by using a osmotic shock. The existance of fluorescene in the periplasma fraction, showed in fig. 3, indicates that Brevibacterium flavum TAT-signal sequence is at least in part functional in E. coli KRX.
The S-layer fusion protein could not be found in the polyacrylamide gel after a SDS-PAGE of the lysate and the cell depris were still red. This indicated that the fusion protein intigrates into the cell membrane with its lipid anchor. For testing this assumption the washed lysate was treted with ionic, nonionic and zwitterionic detergents to release the CspB|mRFP out of the membranes.
The proportionally high RFU in the detergent fractions and the MALDI-TOF analysis of the relevant size range in the polyacrylamid gel approved the insertion into the cell membrane (fig. 3). An insertion of these S-layer proteins might stabilize the membrane structure and increase the stability of cells against mechanical and chemical treatment. A stabilization of E. coli expressing S-lyer proteins was discribed by Lederer et al., (2010).