Difference between revisions of "Part:BBa K4247025"
| (2 intermediate revisions by one other user not shown) | |||
| Line 1: | Line 1: | ||
===mCherry-SnoopCatcher=== | ===mCherry-SnoopCatcher=== | ||
| − | |||
| − | |||
| + | This part codes for the red fluorescent protein mCherry (<partinfo>BBa_J06504</partinfo>) fused to SnoopCatcher (<partinfo>BBa_K4247009</partinfo>) via a linker. The SnoopCatcher enables the formation of a spontaneous isopeptide bond with any other protein containing the SnoopTag (<partinfo>BBa_K4247008</partinfo>). | ||
==Usage & Biology== | ==Usage & Biology== | ||
| Line 13: | Line 12: | ||
[[File:Snoop 2.png]] | [[File:Snoop 2.png]] | ||
| + | |||
| + | == Characterization == | ||
| + | |||
| + | ==== Addition of SnoopCatcher to mCherry ==== | ||
| + | |||
| + | '''Aim - '''To attach the SnoopCatcher sequence (Part BBa_K4247009) to the original mCherry part (Part BBa_J06504). | ||
| + | |||
| + | '''Results - '''We designed 6 primers with Uracil to perform a USER cloning reaction. Two primers to amplify the mCherry fragment while swapping the 6-His tag’s location from the C-terminus to the N-terminus, two for amplifying the backbone of the expression vector (pBAD) and two for amplifying the SnoopCatcher fragment from a part we already produced (BBa_K4247021 - Mfp151_SnoopCatcher). We verified the success of the cloning via PCR and sequencing. | ||
| + | |||
| + | From the final USER cloned plasmid coding for the mCherry_SnoopCtacher, we amplified mCherry, SnoopCatcher and the whole mCherry_SnoopCatcher fragment via PCR to confirm that our USER cloning worked. We also used mCherry and SnoopCatcher fragments as controls. | ||
| + | |||
| + | From a plasmid expressing just the mCherry, we used two sets of primers (seq primers and USER primers) to amplify the mCherry fragment as a control for mCherry. However, the seq primers also amplified some base pairs that were upstream and downstream and hence, one of the control fragments - mCherry Control (seq primers) - seems slightly higher than the other mCherry control fragment. | ||
| + | |||
| + | [[File:mccfig2.jpg|500px]] | ||
| + | |||
| + | '''Conclusion - '''Hence, it is clear that our USER cloning was successful in adding the SnoopCatcher to the mCherry fragment as seen in the PCR. | ||
| + | |||
| + | ==== Protein expression of mCherry_SnoopCatcher ==== | ||
| + | |||
| + | '''Aim - '''To express and purify mCherry_SnoopCatcher. | ||
| + | |||
| + | '''Results - '''We expressed the mCherry_SnoopCatcher and mCherry in E.coli BL21(DE3) and induced protein expression using 0.5% arabinose overnight. The next day, we obtained a bright red E.coli culture for both proteins. Then, we performed SDS-PAGE and a Western blot to check for protein expression. It was clear that both proteins were expressed since we found bands of appropriate size only in the induced lanes. | ||
| + | |||
| + | [[File:mccfig3a.jpg|500px]] | ||
| + | |||
| + | [[File:mccfig3b.jpg|500px]] | ||
| + | |||
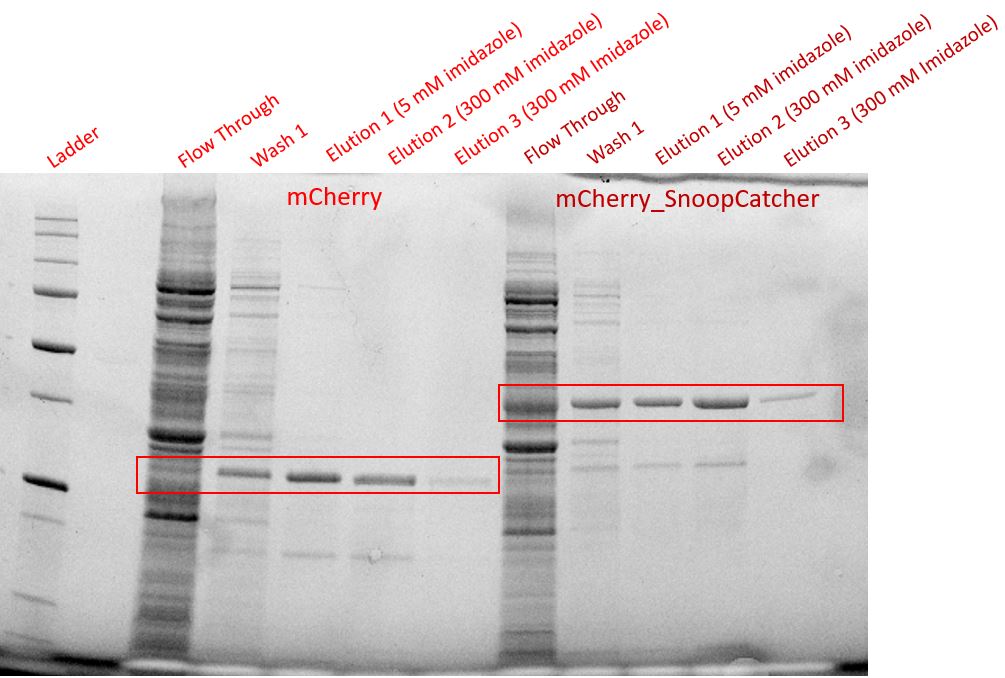
| + | We then performed IMAC using Ni-NTA resin since the protein was expressed with a 6x His-tag and successfully purified mCherry_SnoopCatcher as well as mCherry. Then, we performed SDS-PAGE and a Western blot on the purification fractions of both proteins which confirmed that both mCherry_SnoopCatcher (40.89 KDa) and mCherry (26.7 KDa) were expressed since we found bands at the expected molecular weights. Further, we noticed that mCherry has a leaky expression (see mCherry control in the western blot) and that mCherry_Catcher does not stain as well as mCherry. The purification was not as efficient since an important portion of the protein was lost in the washes, probably due to saturation of the beads. However, we only needed small amounts of proteins for our assays. | ||
| + | |||
| + | [[File:mccfig4.jpg|500px]] | ||
| + | |||
| + | '''Conclusion - '''Thus, mCherry and mCherry_SnoopCatcher were successfully expressed and purified. | ||
| + | |||
| + | ==== Effect of the SnoopCtacher on mCherry’s fluorescence ==== | ||
| + | |||
| + | '''Aim - '''To ensure the addition of the SnoopCatcher would not impair the main characteristic of mCherry, its fluorescence. | ||
| + | |||
| + | '''Results - '''We noticed that mCherry_SnoopCatcher also emits fluorescence in the red wavelengths. It was clear from the lysates that mCherry_SnoopCatcher fluoresces just as much, if not more than the original mCherry, considering that the lysate of mCherry comes from a 0.5L culture while the lysate of mCherry-SnoopCatcher comes from a 150mL culture. This leads us to believe that the addition of SnoopCatcher might improve the protein expression of mCherry or might even improve the fluorescence property of mCherry. However, these results need to be validated since we could not quantify and compare the protein yields or the fluorescence intensity due to time constraints. | ||
| + | |||
| + | [[File:mccfig5a.jpg|500px]] | ||
| + | |||
| + | [[File:mccfig5b.jpg|500px]] | ||
| + | |||
| + | '''Conclusion - '''Hence, the addition of SnoopCatcher does not affect the properties of mCherry. | ||
| + | |||
| + | ==== Validation of SnoopCatcher’s function ==== | ||
| + | |||
| + | '''Aim - '''To demonstrate the spontaneous isopeptide bond formation between our minispidroin protein displaying the SnoopTag and mCherry_SnoopCatcher. | ||
| + | |||
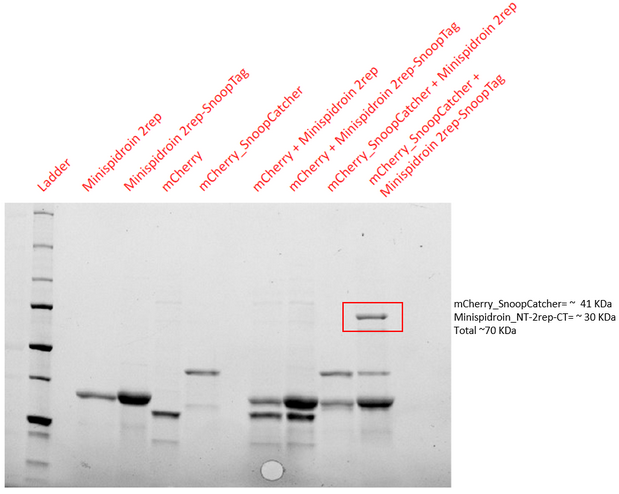
| + | '''Results - '''The proteins were allowed to interact with each other by mixing equal amounts of both proteins. The reaction was carried out at 25°C for 40 minutes and the pH of the reaction was maintained at 8 using pH adjusted PBS. After 40 minutes, the reaction was stopped by adding SDS sample buffer. Then, we performed an SDS-PAGE and a Western blot to visualize the results. We would expect to see a band whose molecular weight is the sum of the molecular weights of both proteins to indicate that both proteins are bound to each via an isopeptide bond between the SnoopTag and the SnoopCatcher. We see such a band only in the reaction where mCherry_SnoopCatcher and Minispidroin 2rep with a SnoopTag were allowed to interact. The fact that we don't see this band in any of the controls confirms that the proteins were able to bind to each other solely due to the SnoopTag-Catcher system. | ||
| + | |||
| + | [[File:mccfig6.jpg|500px]] | ||
| + | |||
| + | '''Conclusion - '''The SnoopCatcher enabled the mCherry_SnoopCatcher to bind to Minispidroin 2rep with a SnoopTag and hence, the function of the SnoopTag-Catcher system was validated. | ||
Latest revision as of 18:58, 11 October 2022
Contents
mCherry-SnoopCatcher
This part codes for the red fluorescent protein mCherry (BBa_J06504) fused to SnoopCatcher (BBa_K4247009) via a linker. The SnoopCatcher enables the formation of a spontaneous isopeptide bond with any other protein containing the SnoopTag (BBa_K4247008).
Usage & Biology
Since the first successful cloning of the green fluorescent protein (GFP) of Aequorea victoria in 1992 (Prasher et al. 1992) fluorescent proteins became a widely used tool in many fields of research. Now an array of these proteins exists, with improved variants and a wide colour spectrum. mCherry is a 26.7 kDa protein, and has a short maturation time (15 min) with an excitation max at 587 nm and emission max at 610 nm (www.fpbase.org). In 2006, its crystal structure was published (Shu and Remington 2006).
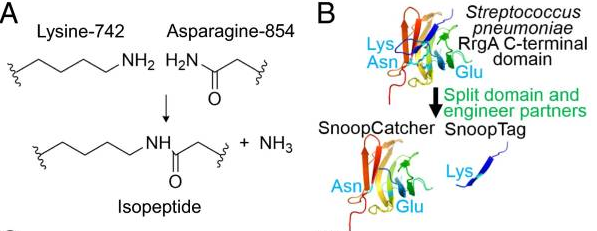
A covalent peptide/protein pair was developed to enable spontaneous isopeptide bond formation between peptide tags. This system was developed based on Gram-positive surface protein, the pilus adhesin RrgA of S. pneumoniae. The D4 domain of this protein is stabilised by an isopeptide forming between a lysine (K742) and an asparagine (N854). This domain was split into a scaffold protein called SnoopCatcher and a 12-residue peptide termed SnoopTag, which can spontaneously form a covalent isopeptide bond upon mixing. The initiation, extension, and release steps use mild conditions, independent of redox state, and therefore should be applicable to a wide range of proteins according to the original paper.
Here, we provide mCherry with an additional feature, the SnoopCatcher. After the fusion of SnoopCatcher, the mCherry protein has a MW of 40.9 kDa. The Tag-Catcher system is a protein conjugation tool that enables the formation of an irreversible isopeptide bond between these two components.
Characterization
Addition of SnoopCatcher to mCherry
Aim - To attach the SnoopCatcher sequence (Part BBa_K4247009) to the original mCherry part (Part BBa_J06504).
Results - We designed 6 primers with Uracil to perform a USER cloning reaction. Two primers to amplify the mCherry fragment while swapping the 6-His tag’s location from the C-terminus to the N-terminus, two for amplifying the backbone of the expression vector (pBAD) and two for amplifying the SnoopCatcher fragment from a part we already produced (BBa_K4247021 - Mfp151_SnoopCatcher). We verified the success of the cloning via PCR and sequencing.
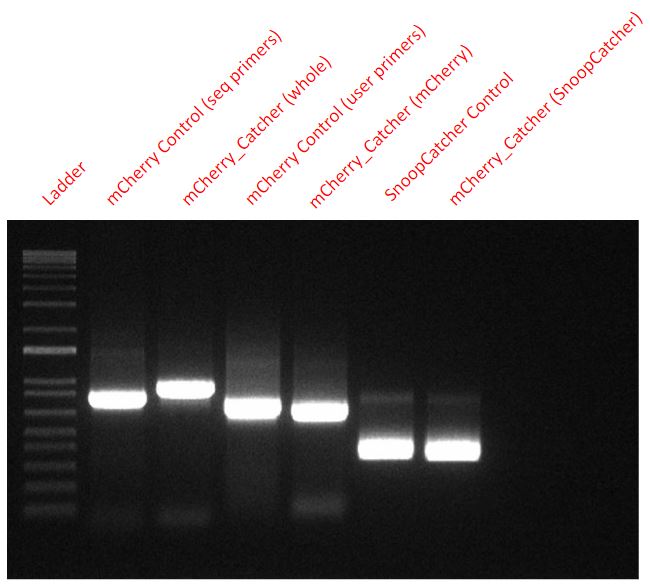
From the final USER cloned plasmid coding for the mCherry_SnoopCtacher, we amplified mCherry, SnoopCatcher and the whole mCherry_SnoopCatcher fragment via PCR to confirm that our USER cloning worked. We also used mCherry and SnoopCatcher fragments as controls.
From a plasmid expressing just the mCherry, we used two sets of primers (seq primers and USER primers) to amplify the mCherry fragment as a control for mCherry. However, the seq primers also amplified some base pairs that were upstream and downstream and hence, one of the control fragments - mCherry Control (seq primers) - seems slightly higher than the other mCherry control fragment.
Conclusion - Hence, it is clear that our USER cloning was successful in adding the SnoopCatcher to the mCherry fragment as seen in the PCR.
Protein expression of mCherry_SnoopCatcher
Aim - To express and purify mCherry_SnoopCatcher.
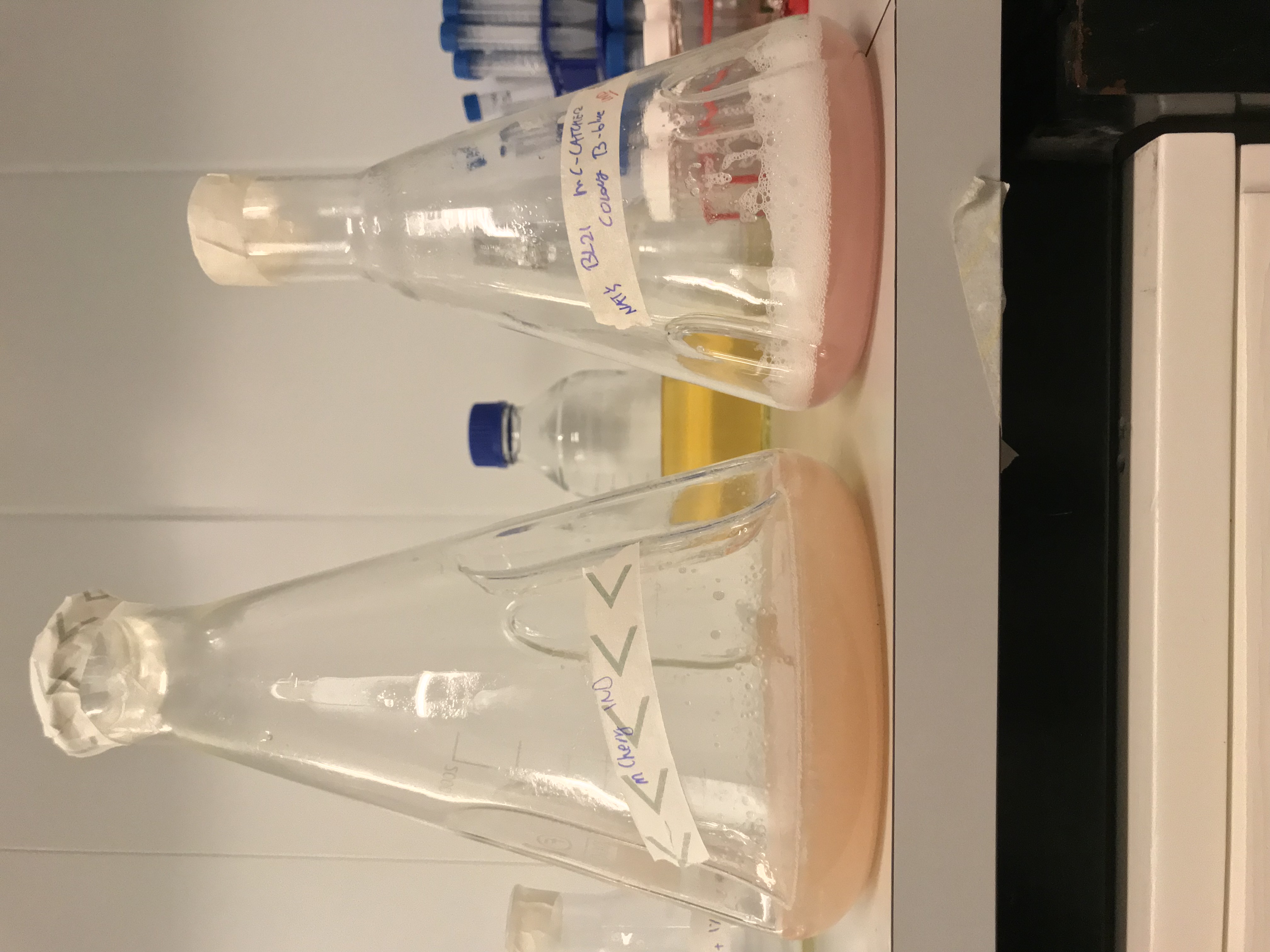
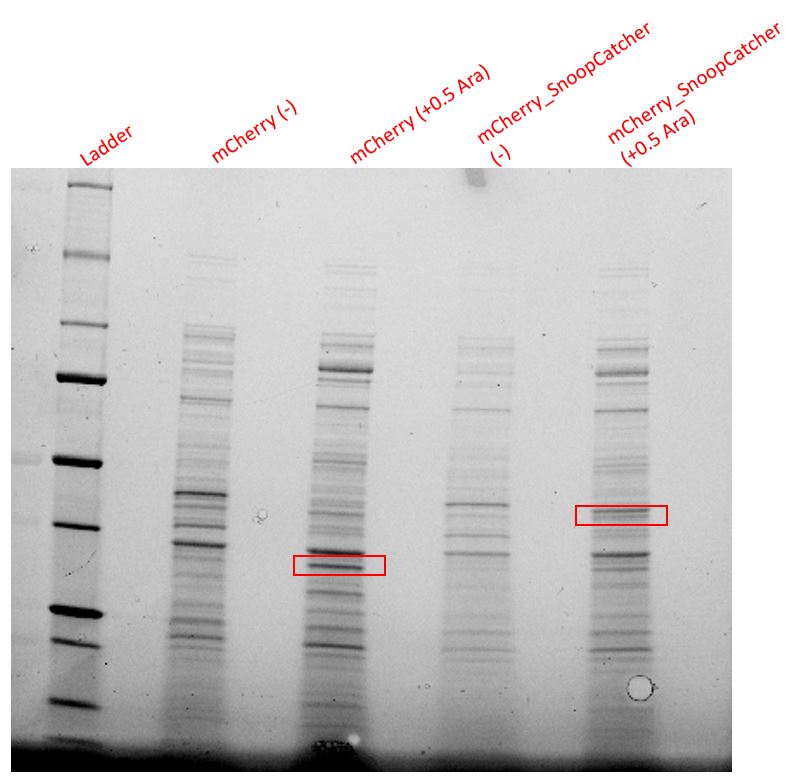
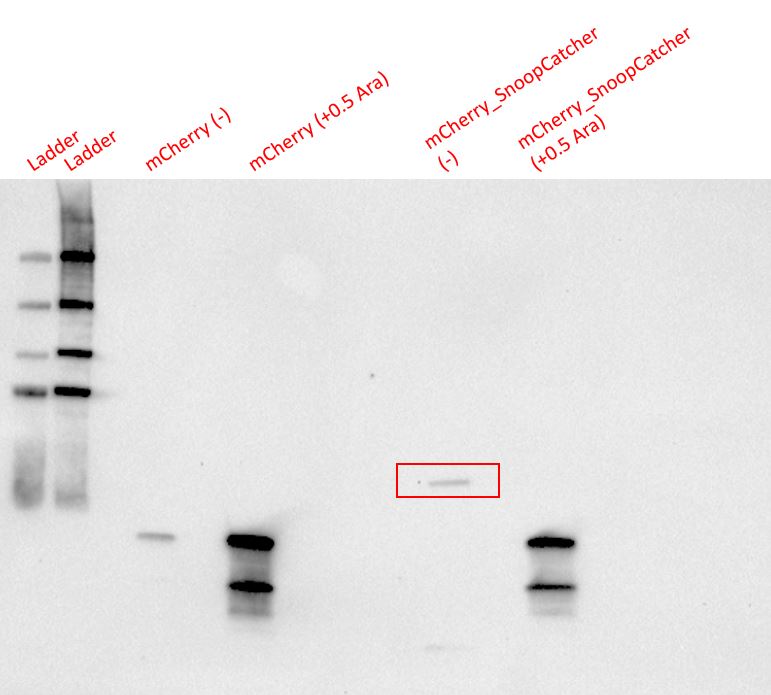
Results - We expressed the mCherry_SnoopCatcher and mCherry in E.coli BL21(DE3) and induced protein expression using 0.5% arabinose overnight. The next day, we obtained a bright red E.coli culture for both proteins. Then, we performed SDS-PAGE and a Western blot to check for protein expression. It was clear that both proteins were expressed since we found bands of appropriate size only in the induced lanes.
We then performed IMAC using Ni-NTA resin since the protein was expressed with a 6x His-tag and successfully purified mCherry_SnoopCatcher as well as mCherry. Then, we performed SDS-PAGE and a Western blot on the purification fractions of both proteins which confirmed that both mCherry_SnoopCatcher (40.89 KDa) and mCherry (26.7 KDa) were expressed since we found bands at the expected molecular weights. Further, we noticed that mCherry has a leaky expression (see mCherry control in the western blot) and that mCherry_Catcher does not stain as well as mCherry. The purification was not as efficient since an important portion of the protein was lost in the washes, probably due to saturation of the beads. However, we only needed small amounts of proteins for our assays.
Conclusion - Thus, mCherry and mCherry_SnoopCatcher were successfully expressed and purified.
Effect of the SnoopCtacher on mCherry’s fluorescence
Aim - To ensure the addition of the SnoopCatcher would not impair the main characteristic of mCherry, its fluorescence.
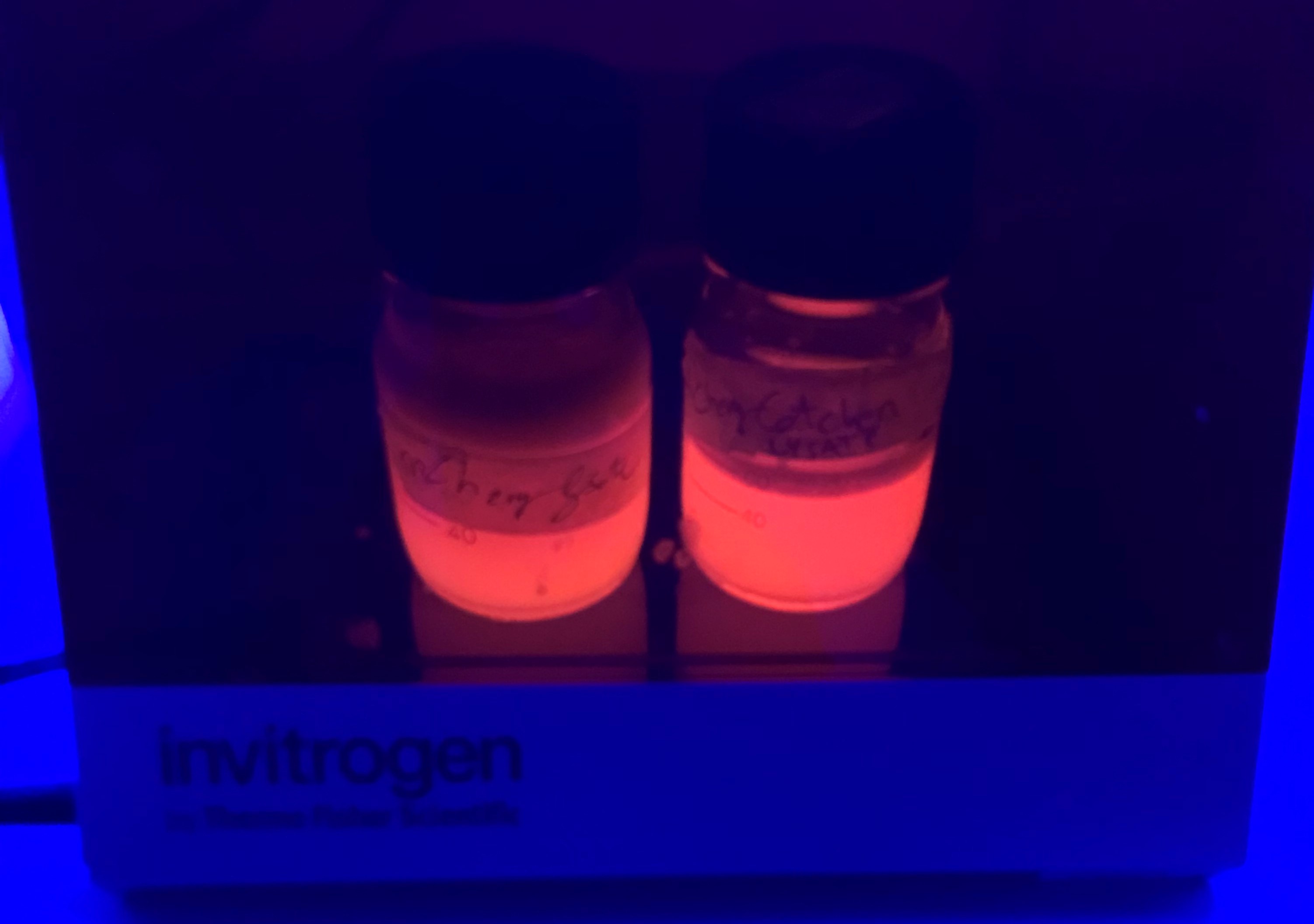
Results - We noticed that mCherry_SnoopCatcher also emits fluorescence in the red wavelengths. It was clear from the lysates that mCherry_SnoopCatcher fluoresces just as much, if not more than the original mCherry, considering that the lysate of mCherry comes from a 0.5L culture while the lysate of mCherry-SnoopCatcher comes from a 150mL culture. This leads us to believe that the addition of SnoopCatcher might improve the protein expression of mCherry or might even improve the fluorescence property of mCherry. However, these results need to be validated since we could not quantify and compare the protein yields or the fluorescence intensity due to time constraints.
Conclusion - Hence, the addition of SnoopCatcher does not affect the properties of mCherry.
Validation of SnoopCatcher’s function
Aim - To demonstrate the spontaneous isopeptide bond formation between our minispidroin protein displaying the SnoopTag and mCherry_SnoopCatcher.
Results - The proteins were allowed to interact with each other by mixing equal amounts of both proteins. The reaction was carried out at 25°C for 40 minutes and the pH of the reaction was maintained at 8 using pH adjusted PBS. After 40 minutes, the reaction was stopped by adding SDS sample buffer. Then, we performed an SDS-PAGE and a Western blot to visualize the results. We would expect to see a band whose molecular weight is the sum of the molecular weights of both proteins to indicate that both proteins are bound to each via an isopeptide bond between the SnoopTag and the SnoopCatcher. We see such a band only in the reaction where mCherry_SnoopCatcher and Minispidroin 2rep with a SnoopTag were allowed to interact. The fact that we don't see this band in any of the controls confirms that the proteins were able to bind to each other solely due to the SnoopTag-Catcher system.
Conclusion - The SnoopCatcher enabled the mCherry_SnoopCatcher to bind to Minispidroin 2rep with a SnoopTag and hence, the function of the SnoopTag-Catcher system was validated.