Difference between revisions of "Part:BBa K4150005"
Jacky CHIN (Talk | contribs) |
Jacky CHIN (Talk | contribs) |
||
| (2 intermediate revisions by the same user not shown) | |||
| Line 10: | Line 10: | ||
<br> | <br> | ||
| − | <span style="color:#00000000">This</span>The vector was amplified from Part:BBa_K3431048 by PCR with a high-fidelity DNA polymerases (KOD-plus) as a backbone with a T7 promoter and a T7 terminator, followed by digested with SpeI and HindIII. The insert of g10.RBS-His-GFP was synthesized by Integrated DNA Technologies, Inc., followed by digested with XbaI and HindIII. E. coli DH5alpha competent cells were transformed with the resulting ligation product and checked by colony PCR and restriction enzymes, as well as confirmed by DNA sequencing. | + | <span style="color:#00000000">This</span>The vector was amplified from [[Part:BBa_K3431048]] by PCR with a high-fidelity DNA polymerases (KOD-plus) as a backbone with a T7 promoter and a T7 terminator, followed by digested with SpeI and HindIII. The insert of g10.RBS-His-GFP was synthesized by Integrated DNA Technologies, Inc., followed by digested with XbaI and HindIII. E. coli DH5alpha competent cells were transformed with the resulting ligation product and checked by colony PCR and restriction enzymes, as well as confirmed by DNA sequencing. |
<br> | <br> | ||
| Line 25: | Line 25: | ||
| − | <span style="color:#00000000">This</span>T7 bacteriophage genome was edited with the BioBrick of T7P-g10.RBS-His-GFP-T7T in the end of T7 gene 10 using the T7Select® 415-1 Cloning Kit (Merck Millipore) according to the manufacturer’s instruction. To select the correct T7 phage genome inserted with the BioBrick, the plaque assay on LB agar plate covered by the lawn of E. coli DH5alpha was performed. Several phages from the plaques were subjected to phage genomic DNA extraction and PCR with corresponding primers (Fig. | + | <span style="color:#00000000">This</span>T7 bacteriophage genome was edited with the BioBrick of T7P-g10.RBS-His-GFP-T7T in the end of T7 gene 10 using the T7Select® 415-1 Cloning Kit (Merck Millipore) according to the manufacturer’s instruction. To select the correct T7 phage genome inserted with the BioBrick, the plaque assay on LB agar plate covered by the lawn of E. coli DH5alpha was performed. Several phages from the plaques were subjected to phage genomic DNA extraction and PCR with corresponding primers (Fig. 2). |
<br> | <br> | ||
| Line 37: | Line 37: | ||
===Functional Assay=== | ===Functional Assay=== | ||
| − | <span style="color:#00000000">This</span>The mouse intestinal commensal E. coli isolates were grown to an OD around 0.6 in the M9 minimal medium supplemented with casamino acids (0.2%), D-glucose (0.3%), vitamin B1 (1 mg/ml), MgSO4 (0.2 mM), CaCl2 (0.1 mM)<ref>Vinay M, Franche N, Grégori G, Fantino JR, Pouillot F, Ansaldi M. Phage-Based Fluorescent Biosensor Prototypes to Specifically Detect Enteric Bacteria Such as E. coli and Salmonella enterica Typhimurium. PLoS One. 2015 Jul 17;10(7):e0131466. doi: 10.1371/journal.pone.0131466. PMID: 26186207; PMCID: PMC4506075.</ref>, followed by infected with the engineered phages at a MOI of 5 for 2 hr at 37°C. The data were read for OD600 and GFP at an ex/em = 483/513 nm every 10 min. Compared to wild-type T7 phage infection, the engineered phage::T7P-g10.RBS-His-GFP-T7T can infect and kill the E. coli in a similar way (Fig. | + | <span style="color:#00000000">This</span>The mouse intestinal commensal E. coli isolates were grown to an OD around 0.6 in the M9 minimal medium supplemented with casamino acids (0.2%), D-glucose (0.3%), vitamin B1 (1 mg/ml), MgSO4 (0.2 mM), CaCl2 (0.1 mM)<ref>Vinay M, Franche N, Grégori G, Fantino JR, Pouillot F, Ansaldi M. Phage-Based Fluorescent Biosensor Prototypes to Specifically Detect Enteric Bacteria Such as E. coli and Salmonella enterica Typhimurium. PLoS One. 2015 Jul 17;10(7):e0131466. doi: 10.1371/journal.pone.0131466. PMID: 26186207; PMCID: PMC4506075.</ref>, followed by infected with the engineered phages at a MOI of 5 for 2 hr at 37°C. The data were read for OD600 and GFP at an ex/em = 483/513 nm every 10 min. Compared to wild-type T7 phage infection, the engineered phage::T7P-g10.RBS-His-GFP-T7T can infect and kill the E. coli in a similar way (Fig. 3). In addition, the engineered phages are able to transfer the GFP reporter cassette into E. coli and make GFP production at the beginning of 40 min (Fig. 4), which is consistent with the time starting to lyse the bacteria (Fig. 3). |
<br> | <br> | ||
| Line 45: | Line 45: | ||
<br><br><br><br><br><br><br><br><br><br><br> | <br><br><br><br><br><br><br><br><br><br><br> | ||
| − | <b>Figure | + | <b>Figure 3 |</b> The infection of the wild-type and the engineered T7 phages on a mouse intestinal commensal E. coli. The E. coli were growing to an OD around 0.6 in the wells of a 96-well plate (Falcon® 96-well Black/Clear Flat Bottom) at 37°C with 200 μl of the supplemented M9 media, followed by the phage (as indicated) infection at a MOI of 5. The values of OD600 were read every 10 min till 2 hr by a microplate reader (Synergy H1 Hybrid Multi-Mode Reader - BioTek Instruments). The growth of E. coli without phages was set as a control. |
<br> | <br> | ||
| Line 53: | Line 53: | ||
<br><br><br><br><br><br><br><br><br><br><br> | <br><br><br><br><br><br><br><br><br><br><br> | ||
| − | <b>Figure | + | <b>Figure 4 |</b> The production of GFP by the engineered T7 phage::T7P-g10.RBS-His-GFP-T7T. The E. coli were growing to an OD around 0.6 in the wells of a 96-well plate (Falcon® 96-well Black/Clear Flat Bottom) at 37°C with 200μl of the supplemented M9 media, followed by the phage (as indicated) infection at a MOI of 5. The values of fluorescence intensity were read at an ex/em = 483/513 nm every 10 min till 2 hr by a microplate reader (Synergy H1 Hybrid Multi-Mode Reader - BioTek Instruments). The background levels were measured with the controls of E. coli without phages or infected with wild-type phages. |
<br><br> | <br><br> | ||
| + | |||
| + | ===<b>Reference</b>=== | ||
| + | <references /> | ||
| + | |||
<!-- Add more about the biology of this part here | <!-- Add more about the biology of this part here | ||
Latest revision as of 07:12, 20 September 2022
T7P-g10.RBS-His-GFP-T7T
Plasmid Construction
ThisTo verify the function of the engineered phage, we created a GFP reporter in the context of DNA elements composed of T7 promoter, g10.RBS and T7 terminator.
ThisThe vector was amplified from Part:BBa_K3431048 by PCR with a high-fidelity DNA polymerases (KOD-plus) as a backbone with a T7 promoter and a T7 terminator, followed by digested with SpeI and HindIII. The insert of g10.RBS-His-GFP was synthesized by Integrated DNA Technologies, Inc., followed by digested with XbaI and HindIII. E. coli DH5alpha competent cells were transformed with the resulting ligation product and checked by colony PCR and restriction enzymes, as well as confirmed by DNA sequencing.
Figure 1 | T7P-g10.RBS-His-GFP-T7T (BBa_K4150005) construction. DNAs were run electrophoresis on 1% agarose gel with 1kb marker. (A) Vector (2154 bp) and Insert (g10.RBS-His-GFP, 841 bp) were digested by indicated restriction enzymes before ligation. (B) 4 colonies were subject to colony PCR with VF2 forward primer and GFP reverse primers (PCR product size: 995 bp). (C) The plasmid DNAs were extracted and digested by EcoRI and PstI (Product size: 2029 and 954 bps).
Phage Engineering
ThisT7 bacteriophage genome was edited with the BioBrick of T7P-g10.RBS-His-GFP-T7T in the end of T7 gene 10 using the T7Select® 415-1 Cloning Kit (Merck Millipore) according to the manufacturer’s instruction. To select the correct T7 phage genome inserted with the BioBrick, the plaque assay on LB agar plate covered by the lawn of E. coli DH5alpha was performed. Several phages from the plaques were subjected to phage genomic DNA extraction and PCR with corresponding primers (Fig. 2).
Figure 2 | T7 phage engineering with T7P-g10.RBS-His-GFP-T7T. (A) The plaque assay was conducted on the LB agar plate, where E. coli DH5alpha cells were infected with the engineered phages. (B) The gDNAs of phages from 8 plaques were extracted and subjected to PCR with T7SelectUP primer and T7SelectDOWN primer provided in the kit (Product size: 1024 bp). DNAs were run electrophoresis on 1% agarose gel with 1kb marker.
Functional Assay
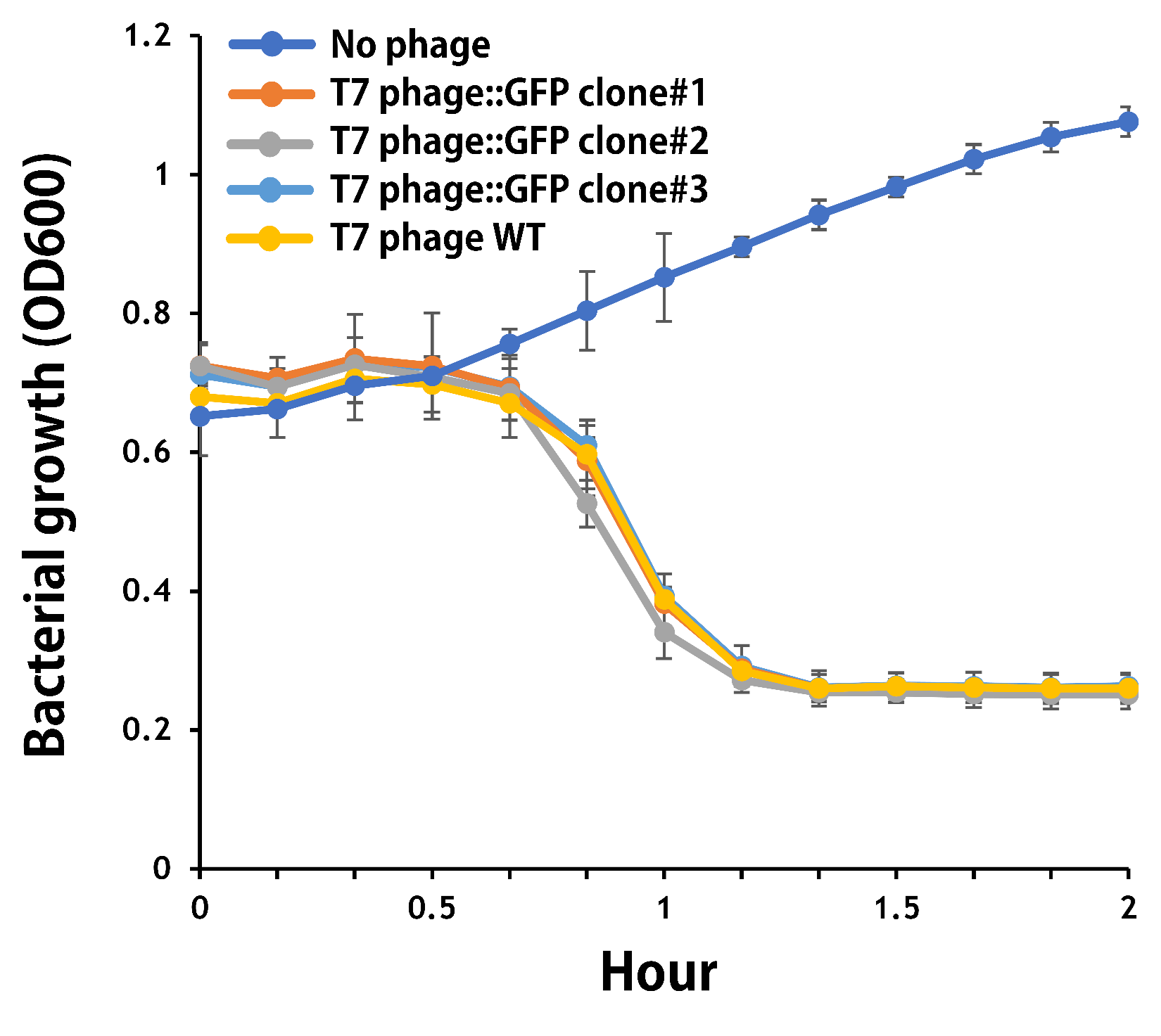
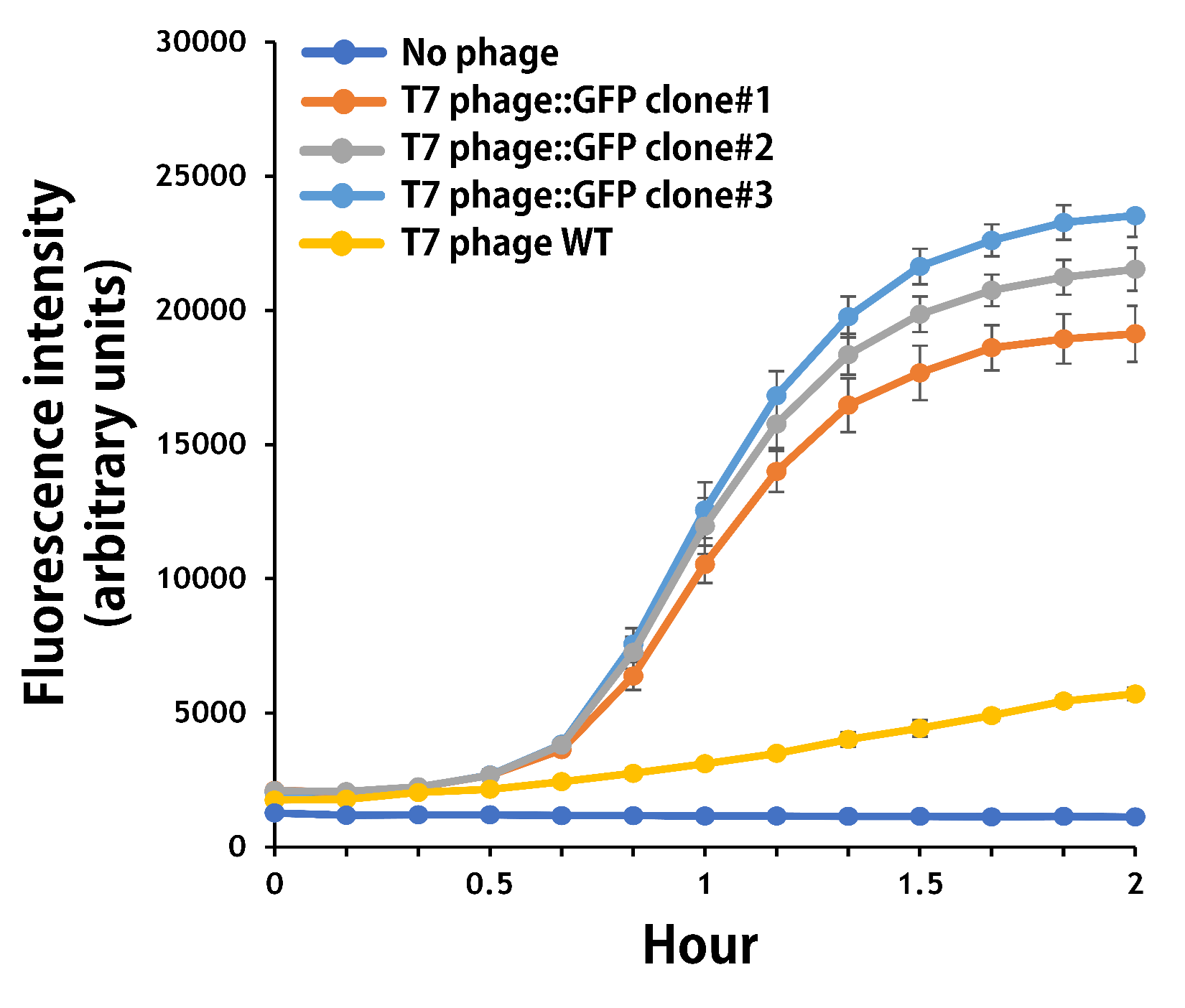
ThisThe mouse intestinal commensal E. coli isolates were grown to an OD around 0.6 in the M9 minimal medium supplemented with casamino acids (0.2%), D-glucose (0.3%), vitamin B1 (1 mg/ml), MgSO4 (0.2 mM), CaCl2 (0.1 mM)[1], followed by infected with the engineered phages at a MOI of 5 for 2 hr at 37°C. The data were read for OD600 and GFP at an ex/em = 483/513 nm every 10 min. Compared to wild-type T7 phage infection, the engineered phage::T7P-g10.RBS-His-GFP-T7T can infect and kill the E. coli in a similar way (Fig. 3). In addition, the engineered phages are able to transfer the GFP reporter cassette into E. coli and make GFP production at the beginning of 40 min (Fig. 4), which is consistent with the time starting to lyse the bacteria (Fig. 3).
Figure 3 | The infection of the wild-type and the engineered T7 phages on a mouse intestinal commensal E. coli. The E. coli were growing to an OD around 0.6 in the wells of a 96-well plate (Falcon® 96-well Black/Clear Flat Bottom) at 37°C with 200 μl of the supplemented M9 media, followed by the phage (as indicated) infection at a MOI of 5. The values of OD600 were read every 10 min till 2 hr by a microplate reader (Synergy H1 Hybrid Multi-Mode Reader - BioTek Instruments). The growth of E. coli without phages was set as a control.
Figure 4 | The production of GFP by the engineered T7 phage::T7P-g10.RBS-His-GFP-T7T. The E. coli were growing to an OD around 0.6 in the wells of a 96-well plate (Falcon® 96-well Black/Clear Flat Bottom) at 37°C with 200μl of the supplemented M9 media, followed by the phage (as indicated) infection at a MOI of 5. The values of fluorescence intensity were read at an ex/em = 483/513 nm every 10 min till 2 hr by a microplate reader (Synergy H1 Hybrid Multi-Mode Reader - BioTek Instruments). The background levels were measured with the controls of E. coli without phages or infected with wild-type phages.
Reference
- ↑ Vinay M, Franche N, Grégori G, Fantino JR, Pouillot F, Ansaldi M. Phage-Based Fluorescent Biosensor Prototypes to Specifically Detect Enteric Bacteria Such as E. coli and Salmonella enterica Typhimurium. PLoS One. 2015 Jul 17;10(7):e0131466. doi: 10.1371/journal.pone.0131466. PMID: 26186207; PMCID: PMC4506075.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 871
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 34
Illegal BsaI.rc site found at 787