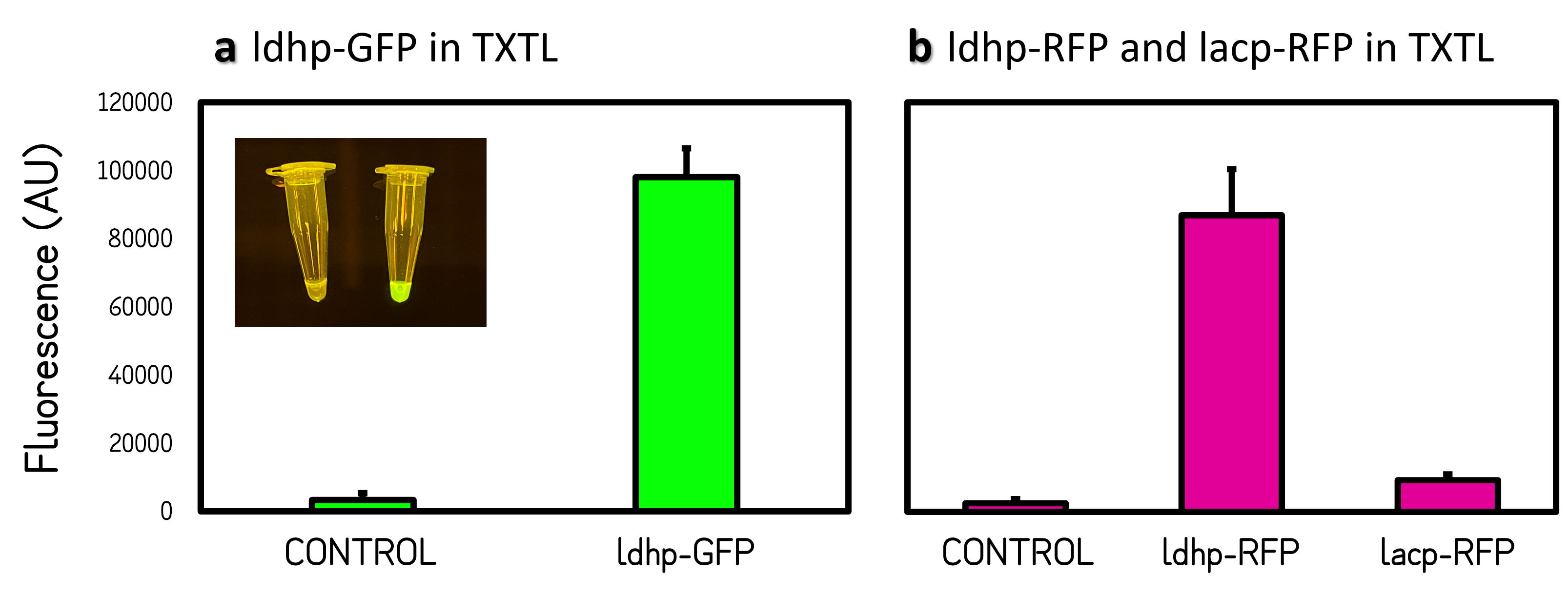Difference between revisions of "Part:BBa K3728002"
| (88 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
| − | |||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K3728002 short</partinfo> | <partinfo>BBa_K3728002 short</partinfo> | ||
| + | [[File:T--Mingdao--BBa K3728002-1.png|600px|right]] | ||
| + | <span style="color:#00000000">This</span>Tol2 transposon system is highly used in zebrafish transgenesis. The transposase protein (TPase) is from the Medaka fish (Oryzias latipes) aka Japanese rice fish, which catalyzes the transposition of the Tol2 elements through cut-and-paste mechanism. The minimal transposable Tol2 sequence (mTol2) contains 200-bp left arm and 150-bp right arm<ref name="z">Urasaki A, Morvan G, Kawakami K. Functional dissection of the Tol2 transposable element identified the minimal cis-sequence and a highly repetitive sequence in the subterminal region essential for transposition. Genetics. 2006 Oct;174(2):639-49. doi: 10.1534/genetics.106.060244.</ref>. Up to 11kb DNA insert between Tol2 sequence can be integrated into the genome of nearly all vertebrates including zebrafish, frog, chicken, mouse, and human <ref>Kawakami K. Tol2: a versatile gene transfer vector in vertebrates. Genome Biol. 2007;8 Suppl 1(Suppl 1):S7. doi: 10.1186/gb-2007-8-s1-s7</ref>. | ||
| − | <span style="color:# | + | <span style="color:#00000000">This</span>A further application in synthetic biology was demonstrated by Jun Ni, et. al.<ref>Ni J, Wangensteen KJ, Nelsen D, Balciunas D, Skuster KJ, Urban MD, Ekker SC. Active recombinant Tol2 transposase for gene transfer and gene discovery applications. Mob DNA. 2016 Mar 31;7:6. doi: 10.1186/s13100-016-0062-z.</ref>, in which the recombinant TPase protein is fully functional in HeLa cell line and Zebrafish germline cells. In addition, the TPase can be expressed under T7 promoter in E. coli BL21 and purified with N-terminal 6xHis tag. The transposase is active in vitro and mediated the integration of DNA fragments between plasmids with Tol2 elements. |
| − | [[ | + | <span style="color:#00000000">This</span>In our study, we constructed BioBrick Parts of TPase ([[Part:BBa_K3728000]]) and the BioBrick compatible Tol2 vectors ([[Part:BBa_K3728003]]) with reporter (KanR:[[Part:BBa_K3728004]]; GFP:[[Part:BBa_K3728005]]; RFP:[[Part:BBa_K3728006]]; amilCP:[[Part:BBa_K3728007]]) and Phi29 DNA polymerase genes ([[Part:BBa_K3728008]]). We prepared the In vitro transcription-translation (TXTL) system <ref>Garenne D, Noireaux V. Cell-free transcription-translation: engineering biology from the nanometer to the millimeter scale. Curr Opin Biotechnol. 2019 Aug;58:19-27. doi: 10.1016/j.copbio.2018.10.007.</ref><ref name="x">Rustad M, Eastlund A, Marshall R, Jardine P, Noireaux V. Synthesis of Infectious Bacteriophages in an E. coli-based Cell-free Expression System. J Vis Exp. 2017 Aug 17;(126):56144. doi: 10.3791/56144.</ref>and expressed the functional reporter proteins. The recombinant TPase and Phil29 DNA polymerase with His tag were expressed in E. coli BL21. The purified proteins were functional in the plasmid integration assay and rolling circle amplification(RCA) application, respectively. |
| − | |||
| + | === CONSTRUCTION – BIOBRICK COMPATIBLE VECTOR === | ||
| + | [[File:T--Mingdao--BBa K3728003-1.png|800px|center]] | ||
| + | <br> | ||
| + | <span style="color:#00000000">This</span>Minimal Tol2 transposable element (mTol2) has been characterized that is composed of 200-bp left arm and 150-bp right arm<ref name="z">Urasaki A, Morvan G, Kawakami K. Functional dissection of the Tol2 transposable element identified the minimal cis-sequence and a highly repetitive sequence in the subterminal region essential for transposition. Genetics. 2006 Oct;174(2):639-49. doi: 10.1534/genetics.106.060244.</ref>. The 19-bp to 11-kbp DNA inserts between the arms can be excised and transposed efficiently by Tol2 transposase (TPase). Therefore, we’d like to make a BioBrick compatible vector based on Tol2 mobile element (pTol2), which can be further assembled through a BioBrick standard EcoRI-XbaI-SpeI-PstI rule. | ||
| + | <br><br> | ||
| + | <span style="color:#00000000">This</span>We obtained the backbone of pBSII-SK-mTol2-MCS from Addgene (Plasmid #51817), which was given by Elly Tanaka<ref>Khattak S, Murawala P, Andreas H, Kappert V, Schuez M, Sandoval-Guzmán T, Crawford K, Tanaka EM. Optimized axolotl (Ambystoma mexicanum) husbandry, breeding, metamorphosis, transgenesis and tamoxifen-mediated recombination. Nat Protoc. 2014 Mar;9(3):529-40. doi: 0.1038/nprot.2014.040.</ref>. We deleted restriction enzyme sites in MCS and generated novel BioBrick Prefix (EcoRI-NotI-XbaI) and BioBrick Suffix (SpeI-NotI-PstI) elements in the both ends by PCR. The resulting DNA plasmid backbone called pTol2 ([[Part:BBa_K3376002]]) was further assembled with the [[Part:BBa_J04450]] (i.e., the iGEM official standard insert on pSB1C3). The resulting J04450/pTol2 ([[Part:BBa_K3376003]]) was checked by PCR (Fig. 1a) and restriction enzymes (Fig. 1b) and also confirmed by sequencing. | ||
| − | + | [[File:T--Mingdao--008photo2 1019.png|400px|left]] | |
| + | <br><br><br><br><br><br>Figure 1 | pTol2 and J04450/pTol2 constructs check. DNAs were run electrophoresis on 1% agarose gel with 1kb marker. (a) PCR producs of pTol2 (lane 1, 3429 bp) and BBa_J04450 (lane 2, 1112 bp). (b) 4 clones of J04450/pTol2 were checked by restriction enzymes (~ 3432 bp and ~1110 bp). Lanes 1-4 by EcoRI and SpeI. Lanes 5-8 by XbaI and PstI. | ||
| + | <br> | ||
| + | |||
| + | === CHARACTERIZATION - TXTL & REPORTER ASSAY === | ||
| + | ====Cell-free TXTL system==== | ||
| + | <span style="color:#00000000">This</span>In vitro transcription and translation (TXTL) is a convenient cell-free system that has increasingly been developed to apply in synthetic biology<ref>Marshall R, Noireaux V. Synthetic Biology with an All E. coli TXTL System: Quantitative Characterization of Regulatory Elements and Gene Circuits. Methods Mol Biol. 2018;1772:61-93. doi: 10.1007/978-1-4939-7795-6_4.</ref><ref>Tinafar A, Jaenes K, Pardee K. Synthetic Biology Goes Cell-Free. BMC Biol. 2019 Aug 8;17(1):64. doi: 10.1186/s12915-019-0685-x.</ref>. In addition to achieve biosafety level, TXTL becomes powerful in prototype characterization of genetic parts, devices and circuits. Moreover, TXTL is particularly useful to express and purify proteins which are toxic, insoluble or unstable in cell-based system. Furthermore and amazingly, Dr. Vincent Noireau’s lab has demonstrated cell-free TXTL application in infectious bacteriophage production, in which T7 phage (40kbp, 77 genes, dsDNA) and T4 phage (170kbp, 289 genes, dsDNA) genome replication, synthesis, assembly can be performed in vitro just in a single test tube<ref name="y">Shin J, Jardine P, Noireaux V. Genome replication, synthesis, and assembly of the bacteriophage T7 in a single cell-free reaction. ACS Synth Biol. 2012 Sep 21;1(9):408-13. doi: 10.1021/sb300049p.</ref><ref name="x">Rustad M, Eastlund A, Jardine P, Noireaux V. Cell-free TXTL synthesis of infectious bacteriophage T4 in a single test tube reaction. Synth Biol (Oxf). 2018 Jan 22;3(1):ysy002. doi: 10.1093/synbio/ysy002.</ref> | ||
| + | <br><br> | ||
| + | ==== Promoter Activity ==== | ||
| + | <span style="color:#00000000">This</span>To test the TPase activity and Tol2 transposon system, we inserted a kanamycin resistance gene (KanR) cassette ([[Part:BBa_K3376004]]) and the reporters of ldhp-GFP-Tr([[Part:BBa_K3376005]]), ldhp-RFP-Tr([[Part:BBa_K3376006]]) and ldhp-amilCP-Tr([[Part:BBa_K3376007]]) between the transposable elements on the pTol2 vector. ldhp is a constitutive and broad-host-range promoter, which was originally cloned and driving the lactate dehydrogenase gene in S. mutans. We have characterized the ldhp activities in S. mutans and E. coli in our project of iGEM 2020, as well as Salmonella and TXTL in this project of iGEM 2021.([[Part:BBa_K3376000]]) | ||
| + | <br><br> | ||
| + | <span style="color:#00000000">This</span>The GFP and RFP fluorescence intensities driven by ldhp on pTol2 vectors were measured at high level in TXTL reaction (Fig. 2). The strong GFP fluorescence can even be visualized by naked eyes under a Blue LED Illuminator. Compared the activities of ldhp to lac promoter (lacp), lacp is inhibited in TXTL because the extracts of E. coli Rosetta 2 (DE3) contains LacI repressor, which can be relieved by IPTG induction or using E. coli DH5α as extracts. | ||
| + | |||
| + | [[File:T--Mingdao--008photo4 1019.png|600px|left]] | ||
| + | <br><br>Figure 2 | Promoter activities on pTol2 vector in TXTL. GFP fluorescence was measured at Ex/Em = 500/530 nm using a microplate reader of BioTek Synergy H1. RFP was at Ex/Em = 586/611 nm. KanR/pTol2 in TXTX was set as a background control. AU means arbitrary unit. (a) ldhp-GFP-Tr/pTol2 activity in TXTL. The inset photo was captured under a blue LED light. (b) ldhp-RFP-Tr/pTol2 and J04450/pTol2 (i.e., lacp-RFP-Tr) in TXTL. | ||
| − | |||
=== CHARACTERIZATION – PLASMID INTEGRATION === | === CHARACTERIZATION – PLASMID INTEGRATION === | ||
| + | ==== In vitro Integration Assay ==== | ||
| + | [[File:T--Mingdao--001photo4 1017.png|400px|right]] | ||
| + | <span style="color:#00000000">This</span>In vitro integration assay was used by Jun Ni, et al. to characterize the activity of purified recombinant Tol2 transposase (TPase, [[Part:BBa_K3728000]]) and the transposition of Tol2 mobile element<ref name="a">Ni J, Wangensteen KJ, Nelsen D, Balciunas D, Skuster KJ, Urban MD, Ekker SC. Active recombinant Tol2 transposase for gene transfer and gene discovery applications. Mob DNA. 2016 Mar 31;7:6. doi: 10.1186/s13100-016-0062-z</ref>. We prepared the purified TPase from TXTL (Fig. 2) and performed PCR to generate KanR, ldhp-GFP-Tr and ldhp-amilCP-Tr (expressing blue color) DNA fragments flanked by 200-bp right and 150-bp left arms of pTol2 ([[Part:BBa_K3728002]]). The mixtures of TPase, Tol2 mobile inserts and a target plasmid of pSB1C3 were incubated at 30°C for 2 hours. The resulting DNAs were cleaned up and subjected to transform E. coli DH5α competent cells. The colonies displaying kanamycin resistance, green fluorescence or blue color were counted as successful jumping to plasmids by active purified TPase. And the integration rate was calculated by comparing with chloramphenicol resistance or red colonies from pSB1C3 backbone carrying [[Part:BBa_J04450]] part (i.e., RFP coding device). | ||
| + | <span style="color:#00000000">This</span>GFP/Tol2-integrated plasmid can transform E. coli to exhibit weak to strong green fluorescence in Fig. 3. Two plasmids of GFP-positive bacteria were extracted and checked by restriction enzymes. They are larger than pSB1C3 when single cut on the backbone by ApaLI (Fig. 4b). The schematic map of Fig. 4a showed the possible position of integration by a BamHI-cut on the insert and a ApaLI-cut on the backbone (Fig. 4c). The rate of successful integration was calculated by the ratio of numbers of KanR, GFP and BLUE colonies to CmR or RED colonies, respectively (Fig. 4d). The ratio was between 0.2% to 0.9%, of which data are consistent with the observation by Jun Ni, et al<ref name="a">Ni J, Wangensteen KJ, Nelsen D, Balciunas D, Skuster KJ, Urban MD, Ekker SC. Active recombinant Tol2 transposase for gene transfer and gene discovery applications. Mob DNA. 2016 Mar 31;7:6. doi: 10.1186/s13100-016-0062-z</ref>. In sum, we can modify plasmid DNAs in vitro with an insert between Tol2 mobile elements (DONOR) and purified TPase enzymes (HELPER) from TXTL reaction. | ||
| − | + | [[File:T--Mingdao--001photo5 1017.png|400px|center]] | |
| + | ::Figure 3 | E. coli colonies on Cm agar plates were transformed by the mixture of GFP/Tol2 and pSB1C3 with TPase or without TPase as a control. | ||
| + | [[File:T--Mingdao--001photo6 1017.png|600px|center]] | ||
| + | ::Figure 4 | Possible integration map and ratio. (a) Schematic maps showed the predicted integration sites. (b, c) pSB1C3::GFP/Tol2 Clone #1 (lane 1) and #2 (lane 2) or pSB1C3 as a control (lane 3) were cut by ApaLI on the backbone (b) or cut by ApaLI with a BamHI cut on the insert (c). DNA was analyzed by electrophoresis on 1% agarose gel with a 1kb marker. (d) The successful integration ratios are calculated by the numbers of colonies of pSB1C3::KanR/Tol2 on Kan agar plate divided by those of pSBC13 (CmR) on Cm agar plates or by the numbers of pSB1C3::GFP or pSB1C3::BLUE divided by colony numbers of pSB1C3 (RED) on Cm agar plates such as shown in Figure 3. | ||
| + | <br> | ||
| + | === APPLICATION – SALMONELLA REPORTER PHAGE CREATED BY PHAGE GENOME ENGINEERING THROUGH TOL2 TRANSPOSON SYSTEM=== | ||
| + | ==== Construction of Tol2 Transposable Element - ldhp-Phi29 DNA pol-Tr/pTol2 ==== | ||
| + | <span style="color:#00000000">This</span>To engineer RCA reporter Salmonella phage, we requested [[Part:BBa_K3352001]] (Φ29 DNA Polymerase with His-Tag and GS linker Sequence) from iGEM team TAS_Taipei in 2020. The part was assembled with ldhp promoter [[Part:BBa_K3376000]] and a double terminator [[Part:BBa_B0015]] onto the Tol2 mobile element vector (pTol2, [[Part:BBa_K3728002]]). The resulting composite part named ldhp-Phi29 DNA pol-Tr/pTol2 ([[Part:BBa_K3728008]]) was checked by colony PCR, restriction enzymes and sequencing (Fig. 2). | ||
| + | [[File:T--Mingdao--008photo3 1019.png|400px|left]] | ||
| + | <br><br><br><br><br><br><br>Figure 2 | ldhp-Phi29 DNA pol-Tr/pTol2 construct check. DNAs were run electrophoresis on 1% agarose gel with 1kb marker. (a) 4 colonies were subject to PCR with ldhp forward primer and a reverse primer in the end of Phi29 gene (PCR product size: 1965 bp). (b) The DNAs were extracted and digested by EcoRI and PstI (3573 and 1973 bps). | ||
| + | |||
| + | |||
| + | ==== Protein Expression & Functional Assay - Ф29 DNA Polymerase ==== | ||
| + | <span style="color:#00000000">This</span>To characterize the function of recombinant phi29 DNA polymerase, TXTL cell-free system can solve the problem caused by the difficulty in bacterial transformation or without suitable plasmid vectors. We took the DNA of ldhp-Phi29 DNA pol-Tr/pTol2([[Part:BBa K3728008]])mixed into TXTL reaction with Salmonella extracts. The recombinant His-tagged phi29 DNA polymerase protein was purified through Nickel column and analyzed by SDS-PAGE and Coomassie Blue Staining (Fig. 4). The isolated proteins in Elution #4 and #5 were at the size of around 70 kDa as predicted (His-phi29 DNA polymerase: 590 amino acids, 68 kDa) and collected for the following studies. | ||
| + | |||
| + | [[File:T--Mingdao--008photo5 1019.png|500px|left]] | ||
| + | <br><br><br><br><br><br>Figure 4 | His-phi29 DNA polymerase was expressed in TXTL using Salmonella extracts and purified by Nickel column. 5 μg of protein lysates were analyzed by SDS-PAGE and Coomassie Blue Staining using 4–12% gradient gel (NuPAGE™, Thermo Fisher Scientific Inc.) Lane: (1) PageRuler™ Prestained Protein Ladder, (2) Salmonella cell extracts (no DNA control), (3) total lysates in TXTL, (4) flow-through, (5) wash-through, (6) Elution #3, (7) Elution #4, (8) Elution #5, (9) Elution #6, (10) Elution #7, (11) Elution #8. | ||
| + | |||
| + | <br><br> | ||
| + | <span style="color:#00000000">This</span>To test functionality of phi29 DNA polymerase, we performed RCA by mixing a circular ssDNA and primer in the buffer with our purified phi29 DNA polymerase (Φ29) from TXTL using Salmonella extracts or a commercial recombinant Φ29 from New England Biolabs Inc. (NEB) as a positive control. The Φ29 enzymes were diluted with various factors in the assay. After incubation at 30°C for 1 hour, the RCA products were stained with EvaGreen DNA dye and subjected to a microplate reader to measure signals at Ex/Em=500/530 nm. And the fold changes in fluorescence intensity were calculated by dividing the values from Φ29-untreated groups (as controls). As shown in Fig. 11, a 12-fold change was achieved with our Φ29, indicating the functionality and the activity of ldhp-Phi29 DNA pol-Tr/pTol2 that are comparable to the commercial NEB Φ29 enzymes. Moreover, the green fluorescence can readily be seen with a blue led light, even using a 4 times diluted Φ29, proving the super high processivity of phi29 DNA polymerase in DNA amplification. | ||
| + | |||
| + | [[File:T--Mingdao--008photo6 1019.png|400px|left]] | ||
| + | <br><br><br><br><br><br>Figure 5 | RCA assay using TXTL-expressed (MINGDAO) or commercial (NEB) phi29 DNA polymerase (Φ29). The commercial Φ29 was purchased from NEB with a defined activity by units. The control was set without Φ29 treatment. The concentration and dilutions of enzymes were, respectively, 4, 2, 1 g/l for MINGDAO Φ29 and 1, 0.5, 0.25 units for NEB Φ29. The EvaGreen DNA binding signals were read at Ex/Em=500/530 nm in BioTek Synergy H1 Microplate Reader. | ||
| + | <br><br> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ==== Salmonella Phage Engineering ==== | ||
| + | [[File:T--Mingdao--001photo1 1019.png|450px|right]] | ||
| + | <br> | ||
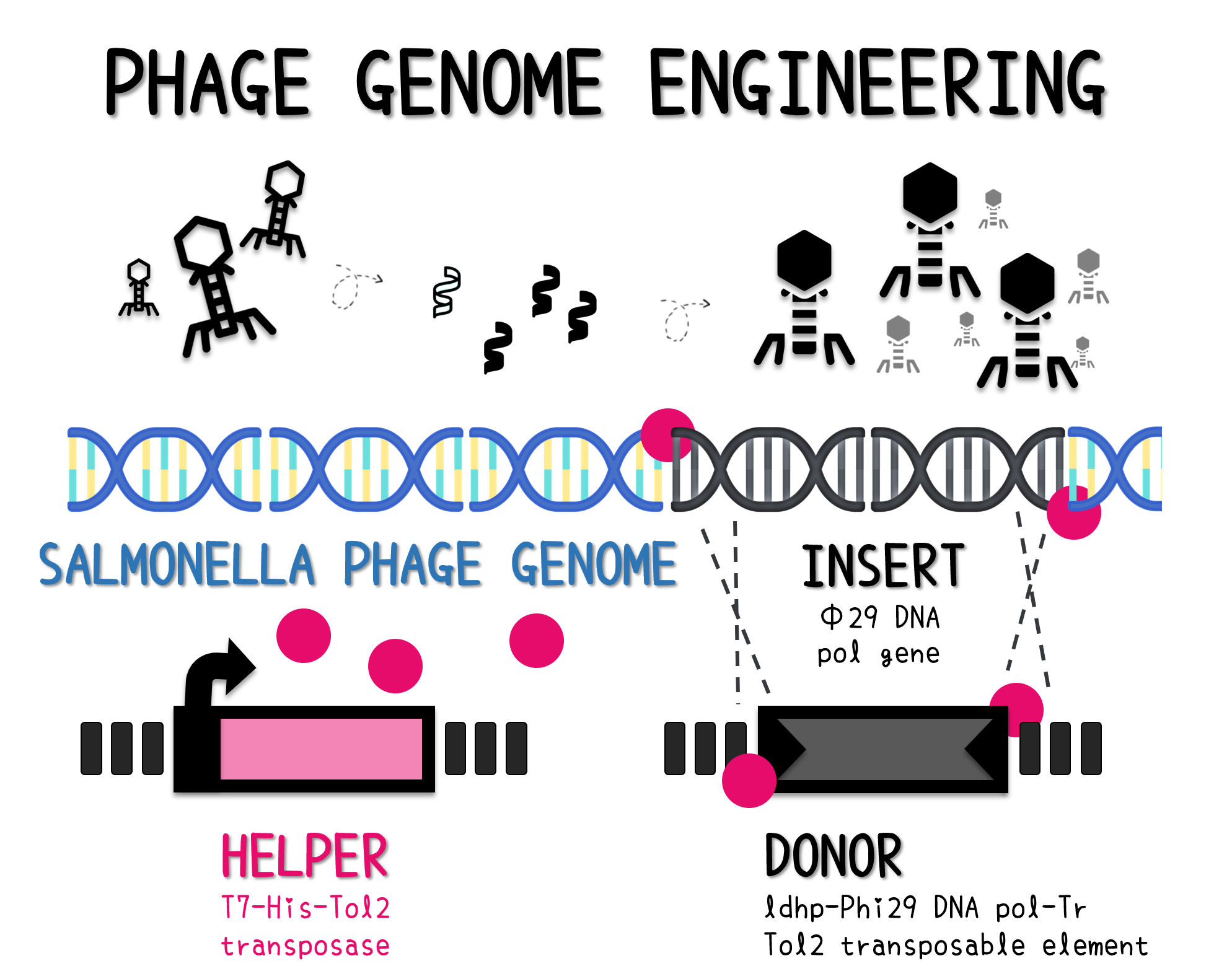
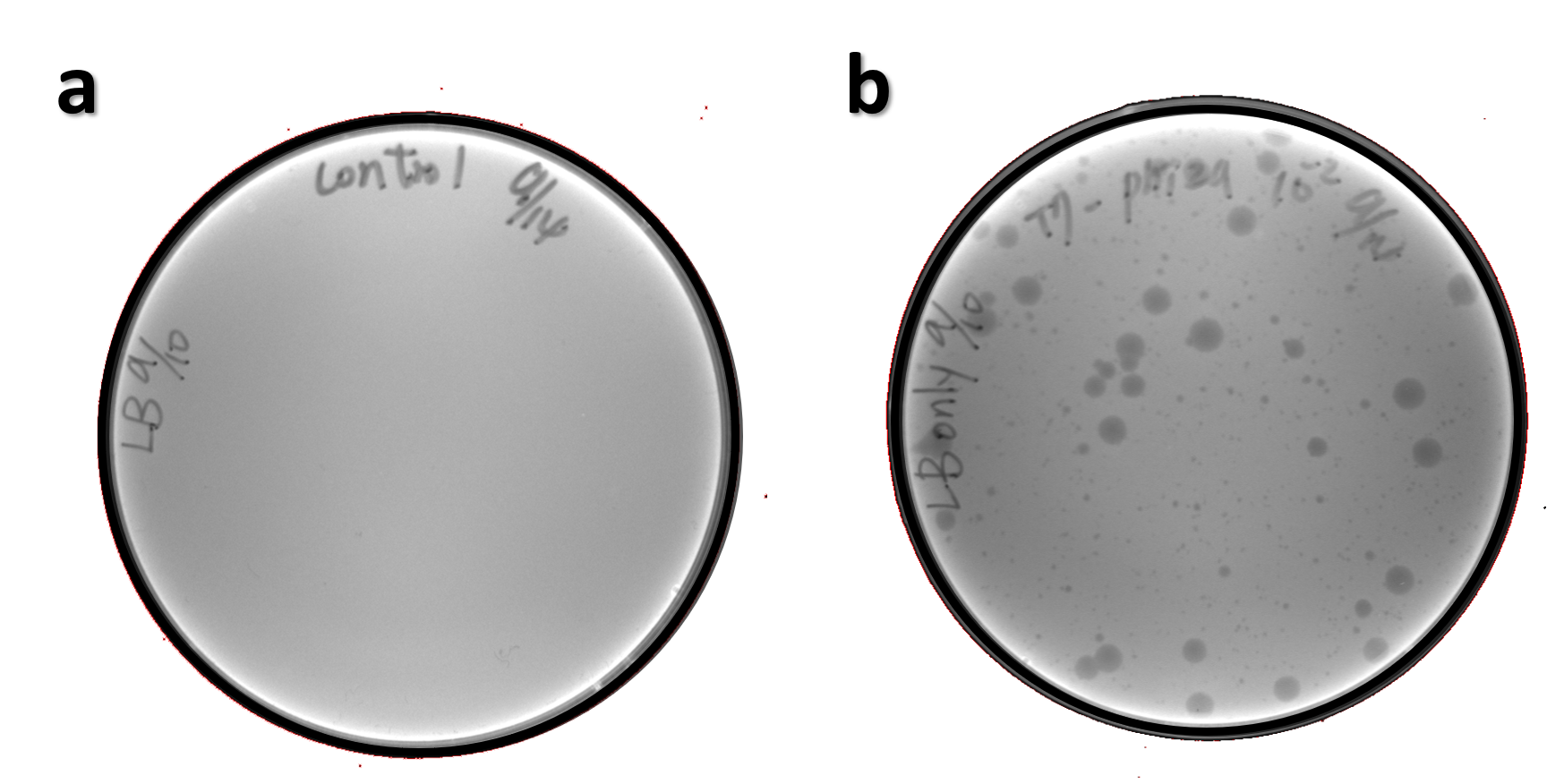
| + | <span style="color:#00000000">This</span>The genome of Salmonella phage #ST1 was extracted and engineered to carry phi29 DNA polymerase gene through the Tol2 transposon system. T7-His-Tol2 transposase was expressed in TXTL using IPTG-induced E. coli Rosetta 2(DE3) extracts, followed by purification with Nickel column. The DNA fragment of Tol2 transposable element carrying ldhp-Phi29 DNA polymerase-Tr (Ф29/Tol2) was amplified by PCR. The extracted Salmonella phage genome and the Ф29/Tol2 DNA fragment were incubated with Tol2 transposase at 30°C for 2 hours. The recombinant phage genome was subjected to TXTL based on the work of Jonghyeon Shin<ref name="y">Shin J, Jardine P, Noireaux V. Genome replication, synthesis, and assembly of the bacteriophage T7 in a single cell-free reaction. ACS Synth Biol. 2012 Sep 21;1(9):408-13. doi: 10.1021/sb300049p.</ref>, where T7 phage genome can be replicated, synthesized, and assembled in a single cell-free reaction. The infectious phage of Salmonella phage #ST1::Ф29 made by TXTL using Salmonella extracts was tested on plaque assay with a culture of Salmonella on the LB agar plate. As shown in Fig. 5b, visible different sizes of plaques were formed on the agar plate, demonstrating infectious phages were produced in our TXTL system. As a control, the phage gDNA without TXTL reaction displayed no plaques (Fig. 5a), indicating the live phages from TXTL are not from the contaminated DNA in the process of genomic DNA extraction. | ||
| + | |||
| + | [[File:T--Mingdao--001photo2 1019.png|400px|left]] | ||
| + | <br><br><br><br><br><br><br>Figure 5 | The recombinant phage synthesis in TXTL. Plaques were formed on a lawn of Salmonella culture on the LB agar plate from TXTL reaction (b) compared to no plaques without TXTL reaction (a). | ||
| + | |||
| + | <br><br><br> | ||
| + | <span style="color:#00000000">This</span>Dozens of plaques were screened by PCR with phi29 DNA polymerase gene-specific primers. A representative result on DNA gel electrophoresis was shown in Fig. 6, in which the successful Ф29/Tol2 insertion (Salmonella phage::Ф29 DNA polymerase, or #ST1::Ф29 for short) can be amplified by PCR with either Tol2 transposable element-specific primers or phi29 DNA polymerase-specific primers, compared to no PCR products from wild-type Salmonella phage #ST1, showing the success of our Salmonella phage engineering with phi29 DNA polymerase gene. | ||
| + | |||
| + | [[File:T--Mingdao--001photo3 1019.png|220px|left]] | ||
| + | <br><br><br><br><br>Figure 6 | Salmonella genome were checked by PCR with Tol2 transposable element-specific primers (lanes 1, 2) or with phi29 DNA polymerase gene-specific primer set 1 (lanes 3, 4) or set 2 (lane 5, 6). The odd numbers refer to Salmonella phage::Ф29 DNA polymerase (#ST1::Ф29), and the even numbers refer to wild-type Salmonella phage #ST1. The gel electrophoresis was performed on a 1% agarose gel with a 1kb DNA ladder. | ||
| + | |||
| + | <br><br> | ||
| + | ==== Salmonella Detection with the Engineered Φ29 Reporter Phage ==== | ||
| + | ===== Comparison to commercial phi29 DNA polymerase ===== | ||
| + | <span style="color:#00000000">This</span>An overnight culture of Salmonella Typhimurium LT2 (~109 cells/ml) were infected by the engineered reporter phage #ST1::Ф29 at MOI=0.1 to produce phi29 DNA polymerase. The lysates were collected after 2 hr or 4 hr of treatment, and then subjected to RCA test. The lysate of phage-infected Salmonella at 2 hr can induce strong RCA reaction (24-fold change) comparable to 2.5U of a commercial phi29 DNA polymerase (NEB) in Fig. 7. Surprisingly, the lysate at 4 hr can not trigger any signal in RCA, suggesting a quick decay of phi29 DNA polymerase in the phage-infected bacterial lysates. | ||
| + | |||
| + | [[File:T--Mingdao--001photo4 1019.png|400px|left]] | ||
| + | <br><br><br><br><br><br>Figure 7 | RCA assay with NEB phi29 DNA polymerase or the Salmonella (10<sup>9</sup> cells/ml) lysates infected by #ST1::Ф29 at MOI=1 which were collected at 2 hr or 4 hr post infection. The fold changes were calculated by the fluorescence intensity of EvaGreen DNA binding of RCA materials without phi29 DNA polymerase as a background control. RCA was performed at 30°C for 1 hr. | ||
| + | <br><br><br><br><br> | ||
| + | |||
| + | ===== Detection Time ===== | ||
| + | <span style="color:#00000000">This</span>We are wondering whether the time of bacterial lysis by phage infection affects the stability of phi29 DNA polymerase protein. An isolated phage may be featured by a latent time (phage generation time in a bacterial cell) and a burst size (numbers of phage produced per bacterial cell). And the latent time and burst size are in a relationship in terms of bacterial density<ref>Abedon ST. Selection for bacteriophage latent period length by bacterial density: A theoretical examination. Microb Ecol. 1989 Sep;18(2):79-88. doi: 10.1007/BF02030117.</ref> and MOI of phage infection<ref>Atel, IR, and K. K. Rao. Bacteriophage burst size as a function of multiplicity of infection. Current Science 1984 53(4): 198–200.</ref>. | ||
| + | <br> | ||
| + | <span style="color:#00000000">This</span>Therefore, we performed the experiment to figure out the latent time and burst size of our Salmonella phage #ST1::Ф29 and the time course of RCA signals during phage infection in Salmonella. 10<sup>5</sup> cells/ml of Salmonella were infected by phage #ST1::Ф29 at MOI=1. The lysates collected by an interval of 5 min until 60 min and subjected to plaque assays and RCA test. As Fig. 8 demonstrated, the phages were released at around 40 min (latent time) to a plateau level with a burst size of average 98.4±14. Interestingly, the RCA signal increased dramatically at 35 min, achieved a high level around 40-45 min, and dropped significantly thereafter, that are consistent with our speculation of the correlation between phage lysis time (latent time) and phi29 DNA polymerase protein functionality. | ||
| + | |||
| + | [[File:T--Mingdao--001photo5 1019.png|480px|left]] | ||
| + | <br><br><br><br><br><br>Figure 8 | Salmonella phage #ST1::Ф29 latent time (min) and burst size (PFU per infected cell, the left Y axis) at MOI=1, and the relationship to RCA assay (fold change, the right Y axis). Phage-infected Salmonella lysates were harvested for 1 hour at an interval of 5 min. The lysates were subjected to plaque assays and RCA reaction followed by stained with EvaGreen dye. The burst sizes were counted by numbers of plaques. The RCA signal were read at Ex/Em=500/530 nm and divided by the background level without phi29 DNA polymerase. | ||
| + | <br><br><br><br> | ||
| + | ===== Detection limit with our hardware ===== | ||
| + | [[File:T--Mingdao--001photo6 1019.png|450px|right]] | ||
| + | |||
| + | <span style="color:#00000000">This</span>Imagine an application of real Salmonella diagnosis in a food or drink. A contaminated sample collected in a large volume may find 1-100 CFU/ml of bacteria<ref>Gwimbi P, George M, Ramphalile M. Bacterial contamination of drinking water sources in rural villages of Mohale Basin, Lesotho: exposures through neighbourhood sanitation and hygiene practices. Environ Health Prev Med. 2019 May 15;24(1):33. doi: 10.1186/s12199-019-0790-z.</ref>. Large volume and bacterial density are key parameters to affect the result of Salmonella detection with our product. | ||
| + | <br> | ||
| + | <span style="color:#00000000">This</span>To overcome the large volume of a sample, we were inspired by Nickel column purification, in which His-tagged phi29 DNA polymerase were bound. Therefore, we think this method may enrich the His-phi29 DNA polymerase from the sample of large volume. Go to our page of ([https://2021.igem.org/Team:Mingdao/Hardware HARDWARE]) for such a design. And the schematic diagram was shown here. | ||
| + | <br> | ||
| + | <span style="color:#00000000">This</span>To examine the feasibility, we made a serial dilution of Salmonella from 10<sup>7</sup> to 10<sup>3</sup> cells in a beaker of 500ml water and prepared the water without bacteria as a control. The water were mixed with Salmonella phage #ST1::Ф29 at the concentration of 10<sup>5</sup> PFU/500ml at room temperature for 25 min. The 3D-printed Luer locker embedded a mini Ni-column was assembled onto a syringe, followed by repeatedly drawing up and pushing back the water in order to pass through the Ni-column. Then, the RCA materials were drawn onto the Ni-column. If His-phi29 DNA polymerase is present, the RCA reaction may be turned on. After 30 min incubation for RCA reaction, the mixtures were push back into a well of a 96-well black plate containing EvaGreen Dye in a total volume of 50 μl. The fluorescence signals were measured at Ex/Em=500/530 nm. Significant RCA signals began to appear in 2x10<sup>2</sup> bacterial cells/ml (Fig. 9). 20 cells/ml can be detected with a slight enhanced signal that is able to be distinguished from the background. However, we can’t measure the cell density under 10 cells/ml of a liquid to be examined. | ||
| + | |||
| + | [[File:T--Mingdao--001photo7 1019.png|400px|left]] | ||
| + | <br><br><br><br><br><br>Figure 9 | Salmonella test at various concentrations between 2-2x10<sup>4</sup> cells/ml in 500ml water with engineered Salmonella phage carrying His-phi29 DNA polymerase gene at the concentration of 200 PFU/ml. RCA was performed on the embedded Ni-column in a 3D-printerd Luer lock adapter. The amplified DNAs were stained with EvaGreen Dye and measured at Ex/Em=500/530 nm in a microplate reader (BioTek Synergy H1). | ||
| + | |||
| + | <br><br><br><br><br> | ||
| + | |||
| + | ===== Comparison with current methods ===== | ||
| + | <span style="color:#00000000">This</span>We demonstrated a proof of concept that it is possible to detect Salmonella by an isolated genetically engineered Salmonella-specific phage carrying phi29 DNA polymerase gene. The His-tagged phi29 DNA polymerase can be enriched from a large volume of a sample onto the homemade mini Ni-column in a Luer-lock adaptor format, where isothermal RCA may be triggered in the presence of the extraordinarily processive phi29 DNA polymerase at room temperature in a time as short as within 30 min. This</span>Bacteriophages possess features that can produce large amounts of phage progeny (the burst size, usually dozens to hundreds PFU per infected bacterium) to release by killing bacterial host in a short time (the latent time, usually 20 min to 1 hr). We mathematically modeled the latent time (min) as a function of cell density (cells/ml) and applied it to simulate a possible real condition using reporter phage to detect Salmonella in a poisoning case. Go to our [https://2021.igem.org/Team:Mingdao/Model MODEL] page for a detail. We predicted the optimal bacterial lysis time (latent time) for RCA is between 16.545 and 24.427 min for 1 to 100 cells/ml of bacteria. Finally, we developed a hardware of 3D-printed Luer-lock adapter containing mini Ni-column. We used it to measure the Salmonella in 500 ml of water. We can detect 200 bacterial cells/ml within 1 hour. About 20 cells/ml of bacteria may be the most probable limit of detection in our device that generated a slight but distinguishable signal compared to the background level of no bacteria control. Compared to traditional Salmonella tests and published engineered reporter phage, we think our product is better than traditional method in term of test time, and our product is comparable and competitive in terms of test limit and time to the current designer phages carrying reporter genes<ref>Smartt AE, Ripp S. Bacteriophage reporter technology for sensing and detecting microbial targets. Anal Bioanal Chem. 2011 May;400(4):991-1007. doi: 10.1007/s00216-010-4561-3</ref><ref>Vinay M, Franche N, Grégori G, Fantino JR, Pouillot F, Ansaldi M. Phage-Based Fluorescent Biosensor Prototypes to Specifically Detect Enteric Bacteria Such as E. coli and Salmonella enterica Typhimurium. PLoS One. 2015 Jul 17;10(7):e0131466. doi: 10.1371/journal.pone.0131466.</ref><ref>Kim S, Kim M, Ryu S. Development of an engineered bioluminescent reporter phage for the sensitive detection of viable Salmonella typhimurium. Anal Chem. 2014 Jun 17;86(12):5858-64. doi: 10.1021/ac500645c.</ref> (Table 1). | ||
| + | [[File:T--Mingdao--001photo8 1019.png|700px|center]] | ||
| + | <br> | ||
| + | |||
| + | ===Reference=== | ||
| + | <references /> | ||
| + | |||
| + | <strong>Note: The map was generated and sponsored by SnapGene.</strong> | ||
| Line 41: | Line 147: | ||
<partinfo>BBa_K3728002 parameters</partinfo> | <partinfo>BBa_K3728002 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
| − | |||
| − | |||
Latest revision as of 08:05, 19 October 2021
pTol2: a Tol2 vector - the Tol2 transposable element
ThisTol2 transposon system is highly used in zebrafish transgenesis. The transposase protein (TPase) is from the Medaka fish (Oryzias latipes) aka Japanese rice fish, which catalyzes the transposition of the Tol2 elements through cut-and-paste mechanism. The minimal transposable Tol2 sequence (mTol2) contains 200-bp left arm and 150-bp right arm[1]. Up to 11kb DNA insert between Tol2 sequence can be integrated into the genome of nearly all vertebrates including zebrafish, frog, chicken, mouse, and human [2].
ThisA further application in synthetic biology was demonstrated by Jun Ni, et. al.[3], in which the recombinant TPase protein is fully functional in HeLa cell line and Zebrafish germline cells. In addition, the TPase can be expressed under T7 promoter in E. coli BL21 and purified with N-terminal 6xHis tag. The transposase is active in vitro and mediated the integration of DNA fragments between plasmids with Tol2 elements.
ThisIn our study, we constructed BioBrick Parts of TPase (Part:BBa_K3728000) and the BioBrick compatible Tol2 vectors (Part:BBa_K3728003) with reporter (KanR:Part:BBa_K3728004; GFP:Part:BBa_K3728005; RFP:Part:BBa_K3728006; amilCP:Part:BBa_K3728007) and Phi29 DNA polymerase genes (Part:BBa_K3728008). We prepared the In vitro transcription-translation (TXTL) system [4][5]and expressed the functional reporter proteins. The recombinant TPase and Phil29 DNA polymerase with His tag were expressed in E. coli BL21. The purified proteins were functional in the plasmid integration assay and rolling circle amplification(RCA) application, respectively.
CONSTRUCTION – BIOBRICK COMPATIBLE VECTOR
ThisMinimal Tol2 transposable element (mTol2) has been characterized that is composed of 200-bp left arm and 150-bp right arm[1]. The 19-bp to 11-kbp DNA inserts between the arms can be excised and transposed efficiently by Tol2 transposase (TPase). Therefore, we’d like to make a BioBrick compatible vector based on Tol2 mobile element (pTol2), which can be further assembled through a BioBrick standard EcoRI-XbaI-SpeI-PstI rule.
ThisWe obtained the backbone of pBSII-SK-mTol2-MCS from Addgene (Plasmid #51817), which was given by Elly Tanaka[6]. We deleted restriction enzyme sites in MCS and generated novel BioBrick Prefix (EcoRI-NotI-XbaI) and BioBrick Suffix (SpeI-NotI-PstI) elements in the both ends by PCR. The resulting DNA plasmid backbone called pTol2 (Part:BBa_K3376002) was further assembled with the Part:BBa_J04450 (i.e., the iGEM official standard insert on pSB1C3). The resulting J04450/pTol2 (Part:BBa_K3376003) was checked by PCR (Fig. 1a) and restriction enzymes (Fig. 1b) and also confirmed by sequencing.
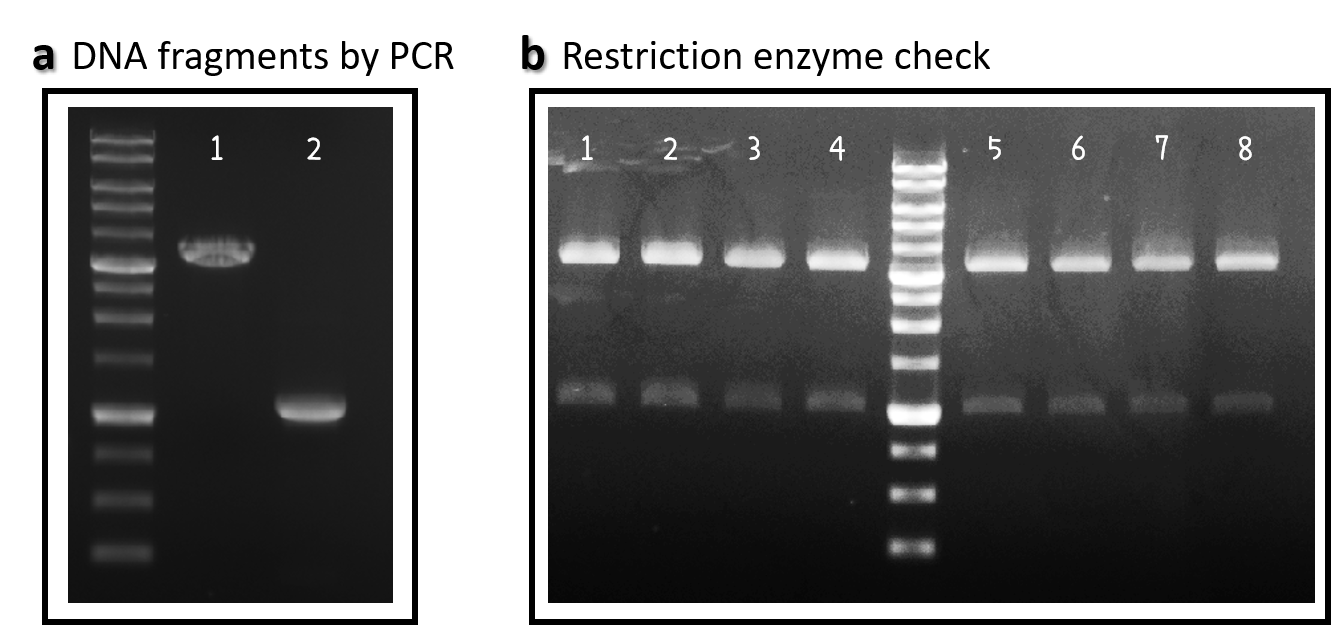
Figure 1 | pTol2 and J04450/pTol2 constructs check. DNAs were run electrophoresis on 1% agarose gel with 1kb marker. (a) PCR producs of pTol2 (lane 1, 3429 bp) and BBa_J04450 (lane 2, 1112 bp). (b) 4 clones of J04450/pTol2 were checked by restriction enzymes (~ 3432 bp and ~1110 bp). Lanes 1-4 by EcoRI and SpeI. Lanes 5-8 by XbaI and PstI.
CHARACTERIZATION - TXTL & REPORTER ASSAY
Cell-free TXTL system
ThisIn vitro transcription and translation (TXTL) is a convenient cell-free system that has increasingly been developed to apply in synthetic biology[7][8]. In addition to achieve biosafety level, TXTL becomes powerful in prototype characterization of genetic parts, devices and circuits. Moreover, TXTL is particularly useful to express and purify proteins which are toxic, insoluble or unstable in cell-based system. Furthermore and amazingly, Dr. Vincent Noireau’s lab has demonstrated cell-free TXTL application in infectious bacteriophage production, in which T7 phage (40kbp, 77 genes, dsDNA) and T4 phage (170kbp, 289 genes, dsDNA) genome replication, synthesis, assembly can be performed in vitro just in a single test tube[9][5]
Promoter Activity
ThisTo test the TPase activity and Tol2 transposon system, we inserted a kanamycin resistance gene (KanR) cassette (Part:BBa_K3376004) and the reporters of ldhp-GFP-Tr(Part:BBa_K3376005), ldhp-RFP-Tr(Part:BBa_K3376006) and ldhp-amilCP-Tr(Part:BBa_K3376007) between the transposable elements on the pTol2 vector. ldhp is a constitutive and broad-host-range promoter, which was originally cloned and driving the lactate dehydrogenase gene in S. mutans. We have characterized the ldhp activities in S. mutans and E. coli in our project of iGEM 2020, as well as Salmonella and TXTL in this project of iGEM 2021.(Part:BBa_K3376000)
ThisThe GFP and RFP fluorescence intensities driven by ldhp on pTol2 vectors were measured at high level in TXTL reaction (Fig. 2). The strong GFP fluorescence can even be visualized by naked eyes under a Blue LED Illuminator. Compared the activities of ldhp to lac promoter (lacp), lacp is inhibited in TXTL because the extracts of E. coli Rosetta 2 (DE3) contains LacI repressor, which can be relieved by IPTG induction or using E. coli DH5α as extracts.
Figure 2 | Promoter activities on pTol2 vector in TXTL. GFP fluorescence was measured at Ex/Em = 500/530 nm using a microplate reader of BioTek Synergy H1. RFP was at Ex/Em = 586/611 nm. KanR/pTol2 in TXTX was set as a background control. AU means arbitrary unit. (a) ldhp-GFP-Tr/pTol2 activity in TXTL. The inset photo was captured under a blue LED light. (b) ldhp-RFP-Tr/pTol2 and J04450/pTol2 (i.e., lacp-RFP-Tr) in TXTL.
CHARACTERIZATION – PLASMID INTEGRATION
In vitro Integration Assay
ThisIn vitro integration assay was used by Jun Ni, et al. to characterize the activity of purified recombinant Tol2 transposase (TPase, Part:BBa_K3728000) and the transposition of Tol2 mobile element[10]. We prepared the purified TPase from TXTL (Fig. 2) and performed PCR to generate KanR, ldhp-GFP-Tr and ldhp-amilCP-Tr (expressing blue color) DNA fragments flanked by 200-bp right and 150-bp left arms of pTol2 (Part:BBa_K3728002). The mixtures of TPase, Tol2 mobile inserts and a target plasmid of pSB1C3 were incubated at 30°C for 2 hours. The resulting DNAs were cleaned up and subjected to transform E. coli DH5α competent cells. The colonies displaying kanamycin resistance, green fluorescence or blue color were counted as successful jumping to plasmids by active purified TPase. And the integration rate was calculated by comparing with chloramphenicol resistance or red colonies from pSB1C3 backbone carrying Part:BBa_J04450 part (i.e., RFP coding device).
ThisGFP/Tol2-integrated plasmid can transform E. coli to exhibit weak to strong green fluorescence in Fig. 3. Two plasmids of GFP-positive bacteria were extracted and checked by restriction enzymes. They are larger than pSB1C3 when single cut on the backbone by ApaLI (Fig. 4b). The schematic map of Fig. 4a showed the possible position of integration by a BamHI-cut on the insert and a ApaLI-cut on the backbone (Fig. 4c). The rate of successful integration was calculated by the ratio of numbers of KanR, GFP and BLUE colonies to CmR or RED colonies, respectively (Fig. 4d). The ratio was between 0.2% to 0.9%, of which data are consistent with the observation by Jun Ni, et al[10]. In sum, we can modify plasmid DNAs in vitro with an insert between Tol2 mobile elements (DONOR) and purified TPase enzymes (HELPER) from TXTL reaction.
- Figure 3 | E. coli colonies on Cm agar plates were transformed by the mixture of GFP/Tol2 and pSB1C3 with TPase or without TPase as a control.
- Figure 4 | Possible integration map and ratio. (a) Schematic maps showed the predicted integration sites. (b, c) pSB1C3::GFP/Tol2 Clone #1 (lane 1) and #2 (lane 2) or pSB1C3 as a control (lane 3) were cut by ApaLI on the backbone (b) or cut by ApaLI with a BamHI cut on the insert (c). DNA was analyzed by electrophoresis on 1% agarose gel with a 1kb marker. (d) The successful integration ratios are calculated by the numbers of colonies of pSB1C3::KanR/Tol2 on Kan agar plate divided by those of pSBC13 (CmR) on Cm agar plates or by the numbers of pSB1C3::GFP or pSB1C3::BLUE divided by colony numbers of pSB1C3 (RED) on Cm agar plates such as shown in Figure 3.
APPLICATION – SALMONELLA REPORTER PHAGE CREATED BY PHAGE GENOME ENGINEERING THROUGH TOL2 TRANSPOSON SYSTEM
Construction of Tol2 Transposable Element - ldhp-Phi29 DNA pol-Tr/pTol2
ThisTo engineer RCA reporter Salmonella phage, we requested Part:BBa_K3352001 (Φ29 DNA Polymerase with His-Tag and GS linker Sequence) from iGEM team TAS_Taipei in 2020. The part was assembled with ldhp promoter Part:BBa_K3376000 and a double terminator Part:BBa_B0015 onto the Tol2 mobile element vector (pTol2, Part:BBa_K3728002). The resulting composite part named ldhp-Phi29 DNA pol-Tr/pTol2 (Part:BBa_K3728008) was checked by colony PCR, restriction enzymes and sequencing (Fig. 2).
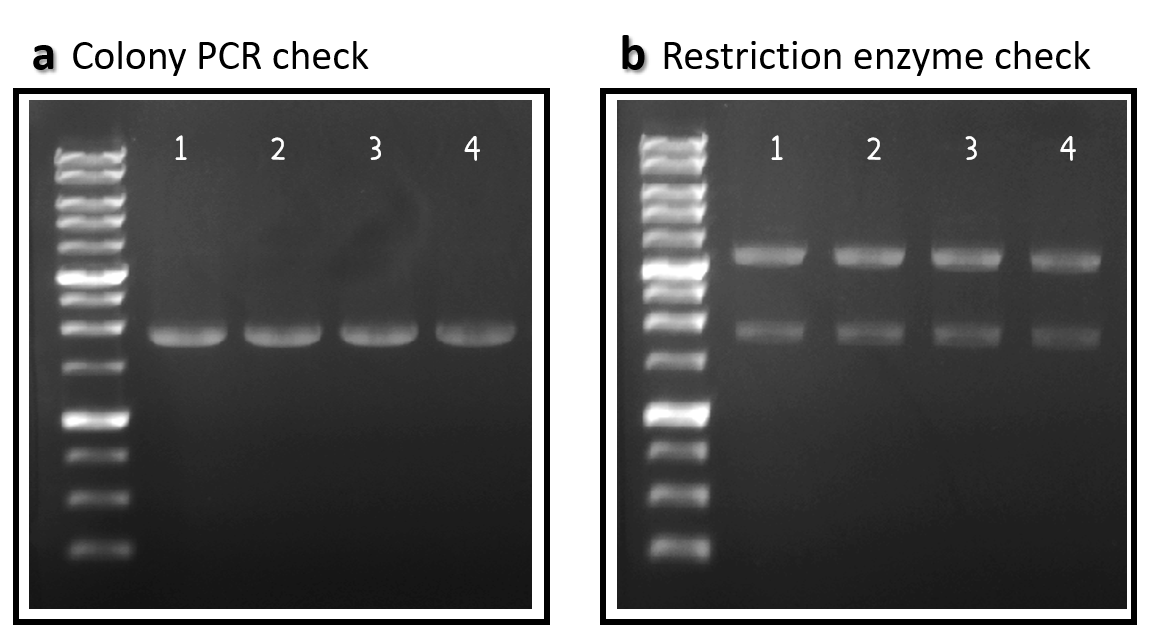
Figure 2 | ldhp-Phi29 DNA pol-Tr/pTol2 construct check. DNAs were run electrophoresis on 1% agarose gel with 1kb marker. (a) 4 colonies were subject to PCR with ldhp forward primer and a reverse primer in the end of Phi29 gene (PCR product size: 1965 bp). (b) The DNAs were extracted and digested by EcoRI and PstI (3573 and 1973 bps).
Protein Expression & Functional Assay - Ф29 DNA Polymerase
ThisTo characterize the function of recombinant phi29 DNA polymerase, TXTL cell-free system can solve the problem caused by the difficulty in bacterial transformation or without suitable plasmid vectors. We took the DNA of ldhp-Phi29 DNA pol-Tr/pTol2(Part:BBa K3728008)mixed into TXTL reaction with Salmonella extracts. The recombinant His-tagged phi29 DNA polymerase protein was purified through Nickel column and analyzed by SDS-PAGE and Coomassie Blue Staining (Fig. 4). The isolated proteins in Elution #4 and #5 were at the size of around 70 kDa as predicted (His-phi29 DNA polymerase: 590 amino acids, 68 kDa) and collected for the following studies.
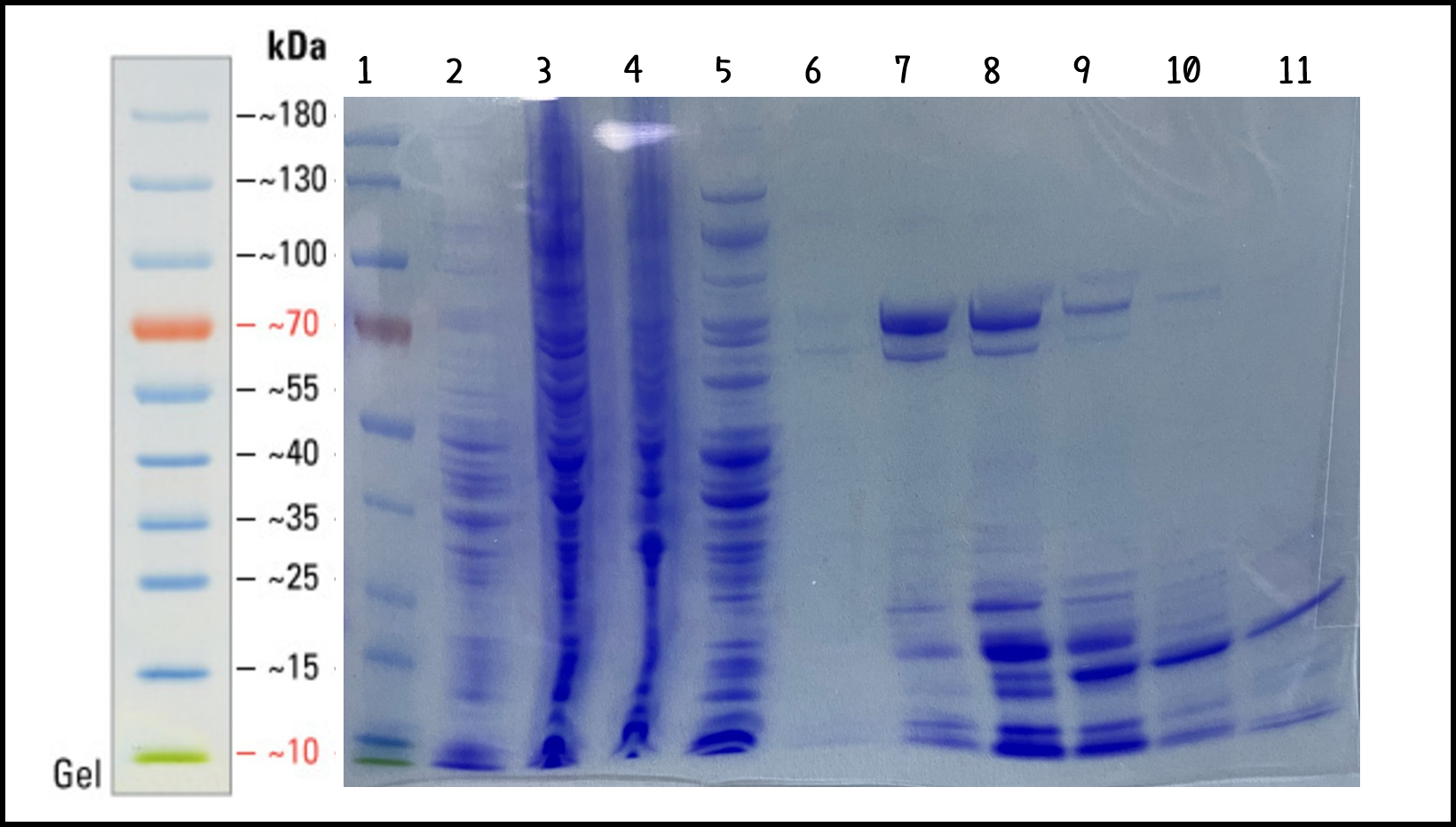
Figure 4 | His-phi29 DNA polymerase was expressed in TXTL using Salmonella extracts and purified by Nickel column. 5 μg of protein lysates were analyzed by SDS-PAGE and Coomassie Blue Staining using 4–12% gradient gel (NuPAGE™, Thermo Fisher Scientific Inc.) Lane: (1) PageRuler™ Prestained Protein Ladder, (2) Salmonella cell extracts (no DNA control), (3) total lysates in TXTL, (4) flow-through, (5) wash-through, (6) Elution #3, (7) Elution #4, (8) Elution #5, (9) Elution #6, (10) Elution #7, (11) Elution #8.
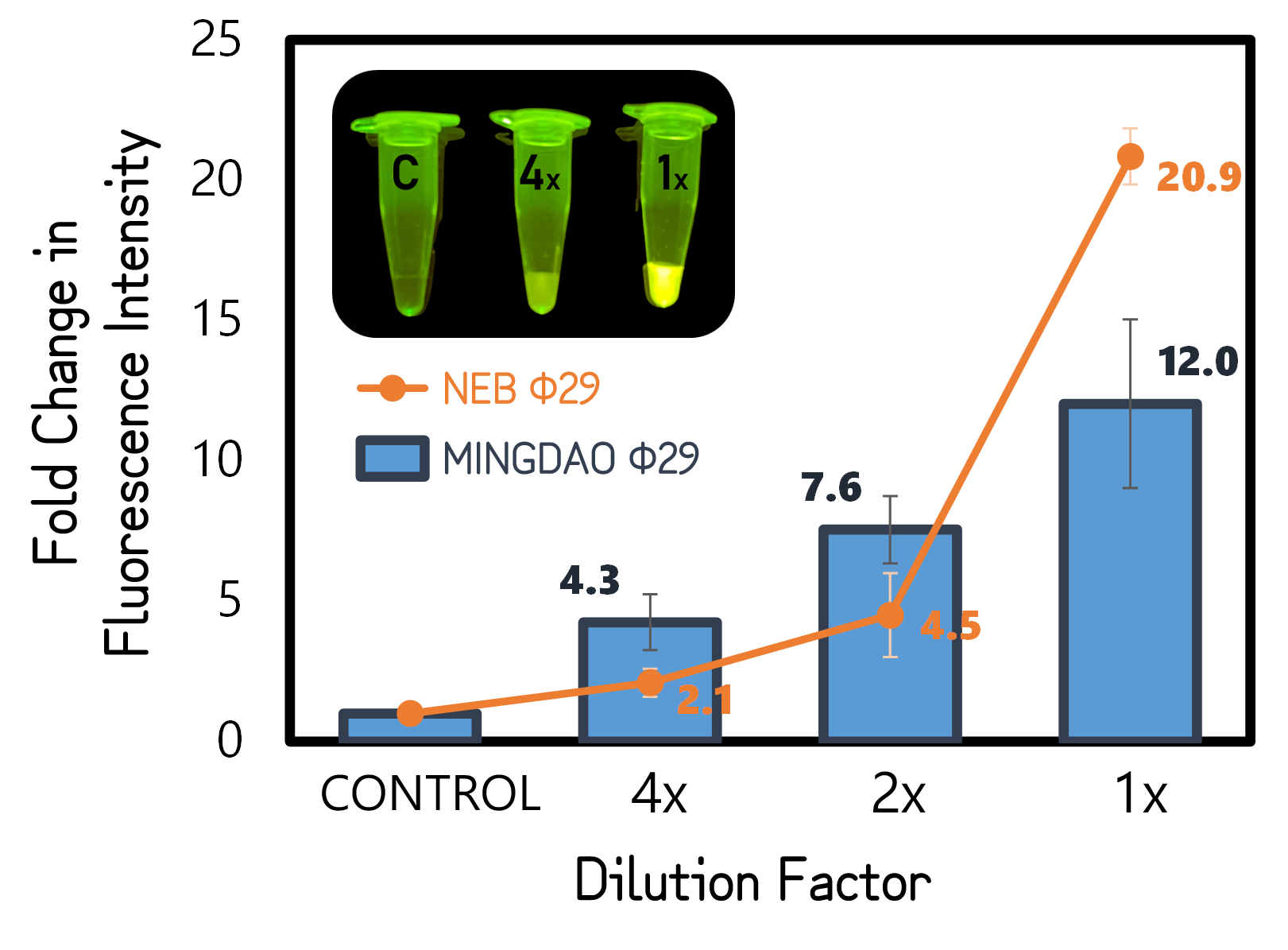
ThisTo test functionality of phi29 DNA polymerase, we performed RCA by mixing a circular ssDNA and primer in the buffer with our purified phi29 DNA polymerase (Φ29) from TXTL using Salmonella extracts or a commercial recombinant Φ29 from New England Biolabs Inc. (NEB) as a positive control. The Φ29 enzymes were diluted with various factors in the assay. After incubation at 30°C for 1 hour, the RCA products were stained with EvaGreen DNA dye and subjected to a microplate reader to measure signals at Ex/Em=500/530 nm. And the fold changes in fluorescence intensity were calculated by dividing the values from Φ29-untreated groups (as controls). As shown in Fig. 11, a 12-fold change was achieved with our Φ29, indicating the functionality and the activity of ldhp-Phi29 DNA pol-Tr/pTol2 that are comparable to the commercial NEB Φ29 enzymes. Moreover, the green fluorescence can readily be seen with a blue led light, even using a 4 times diluted Φ29, proving the super high processivity of phi29 DNA polymerase in DNA amplification.
Figure 5 | RCA assay using TXTL-expressed (MINGDAO) or commercial (NEB) phi29 DNA polymerase (Φ29). The commercial Φ29 was purchased from NEB with a defined activity by units. The control was set without Φ29 treatment. The concentration and dilutions of enzymes were, respectively, 4, 2, 1 g/l for MINGDAO Φ29 and 1, 0.5, 0.25 units for NEB Φ29. The EvaGreen DNA binding signals were read at Ex/Em=500/530 nm in BioTek Synergy H1 Microplate Reader.
Salmonella Phage Engineering
ThisThe genome of Salmonella phage #ST1 was extracted and engineered to carry phi29 DNA polymerase gene through the Tol2 transposon system. T7-His-Tol2 transposase was expressed in TXTL using IPTG-induced E. coli Rosetta 2(DE3) extracts, followed by purification with Nickel column. The DNA fragment of Tol2 transposable element carrying ldhp-Phi29 DNA polymerase-Tr (Ф29/Tol2) was amplified by PCR. The extracted Salmonella phage genome and the Ф29/Tol2 DNA fragment were incubated with Tol2 transposase at 30°C for 2 hours. The recombinant phage genome was subjected to TXTL based on the work of Jonghyeon Shin[9], where T7 phage genome can be replicated, synthesized, and assembled in a single cell-free reaction. The infectious phage of Salmonella phage #ST1::Ф29 made by TXTL using Salmonella extracts was tested on plaque assay with a culture of Salmonella on the LB agar plate. As shown in Fig. 5b, visible different sizes of plaques were formed on the agar plate, demonstrating infectious phages were produced in our TXTL system. As a control, the phage gDNA without TXTL reaction displayed no plaques (Fig. 5a), indicating the live phages from TXTL are not from the contaminated DNA in the process of genomic DNA extraction.
Figure 5 | The recombinant phage synthesis in TXTL. Plaques were formed on a lawn of Salmonella culture on the LB agar plate from TXTL reaction (b) compared to no plaques without TXTL reaction (a).
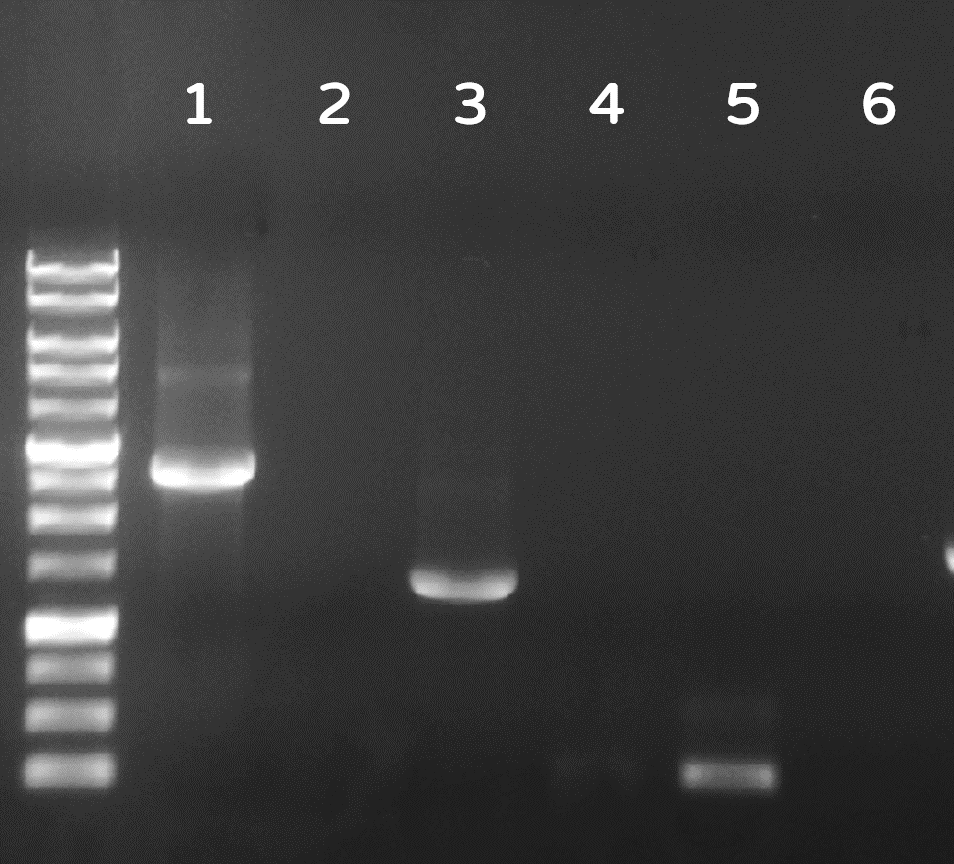
ThisDozens of plaques were screened by PCR with phi29 DNA polymerase gene-specific primers. A representative result on DNA gel electrophoresis was shown in Fig. 6, in which the successful Ф29/Tol2 insertion (Salmonella phage::Ф29 DNA polymerase, or #ST1::Ф29 for short) can be amplified by PCR with either Tol2 transposable element-specific primers or phi29 DNA polymerase-specific primers, compared to no PCR products from wild-type Salmonella phage #ST1, showing the success of our Salmonella phage engineering with phi29 DNA polymerase gene.
Figure 6 | Salmonella genome were checked by PCR with Tol2 transposable element-specific primers (lanes 1, 2) or with phi29 DNA polymerase gene-specific primer set 1 (lanes 3, 4) or set 2 (lane 5, 6). The odd numbers refer to Salmonella phage::Ф29 DNA polymerase (#ST1::Ф29), and the even numbers refer to wild-type Salmonella phage #ST1. The gel electrophoresis was performed on a 1% agarose gel with a 1kb DNA ladder.
Salmonella Detection with the Engineered Φ29 Reporter Phage
Comparison to commercial phi29 DNA polymerase
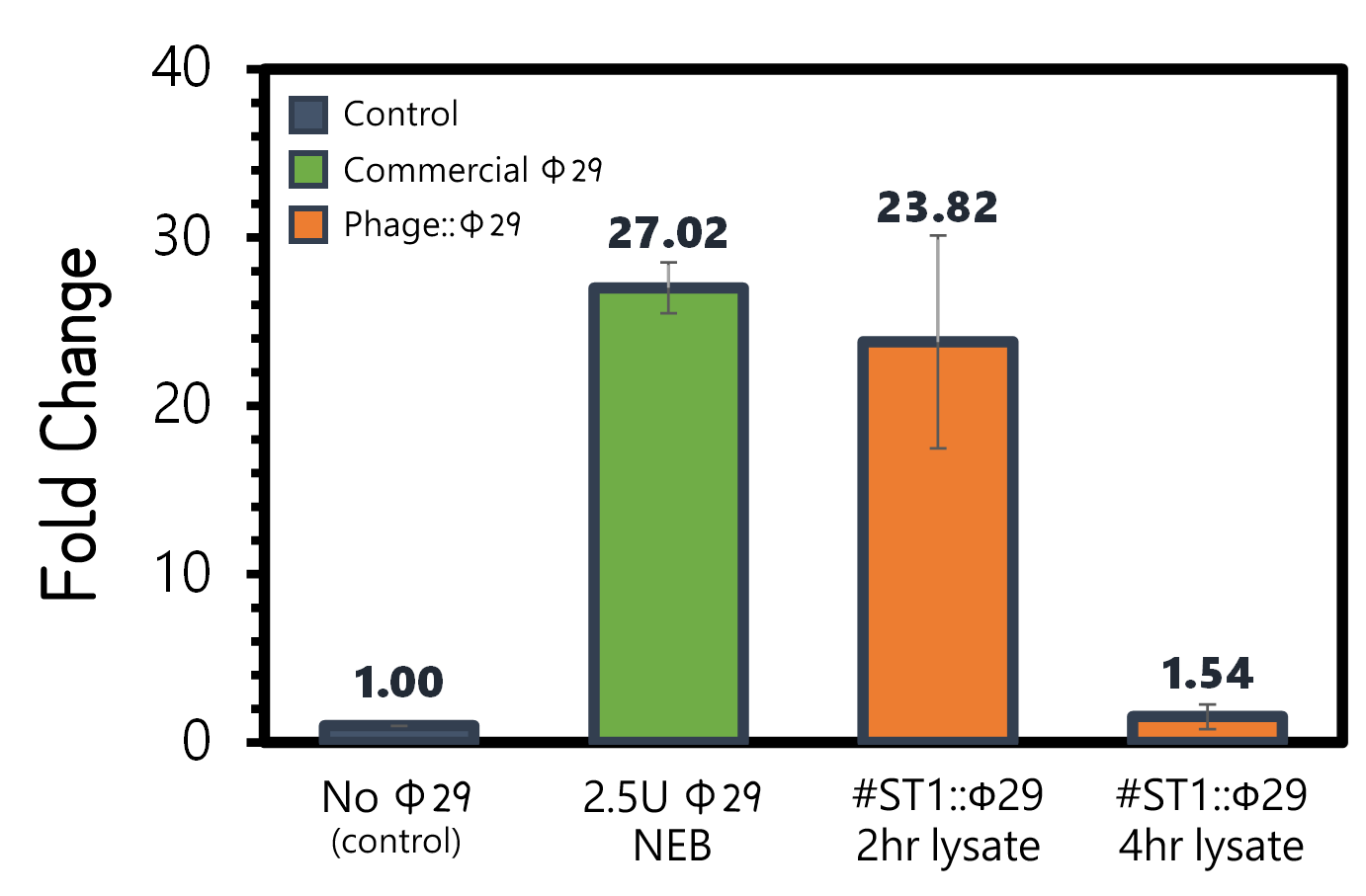
ThisAn overnight culture of Salmonella Typhimurium LT2 (~109 cells/ml) were infected by the engineered reporter phage #ST1::Ф29 at MOI=0.1 to produce phi29 DNA polymerase. The lysates were collected after 2 hr or 4 hr of treatment, and then subjected to RCA test. The lysate of phage-infected Salmonella at 2 hr can induce strong RCA reaction (24-fold change) comparable to 2.5U of a commercial phi29 DNA polymerase (NEB) in Fig. 7. Surprisingly, the lysate at 4 hr can not trigger any signal in RCA, suggesting a quick decay of phi29 DNA polymerase in the phage-infected bacterial lysates.
Figure 7 | RCA assay with NEB phi29 DNA polymerase or the Salmonella (109 cells/ml) lysates infected by #ST1::Ф29 at MOI=1 which were collected at 2 hr or 4 hr post infection. The fold changes were calculated by the fluorescence intensity of EvaGreen DNA binding of RCA materials without phi29 DNA polymerase as a background control. RCA was performed at 30°C for 1 hr.
Detection Time
ThisWe are wondering whether the time of bacterial lysis by phage infection affects the stability of phi29 DNA polymerase protein. An isolated phage may be featured by a latent time (phage generation time in a bacterial cell) and a burst size (numbers of phage produced per bacterial cell). And the latent time and burst size are in a relationship in terms of bacterial density[11] and MOI of phage infection[12].
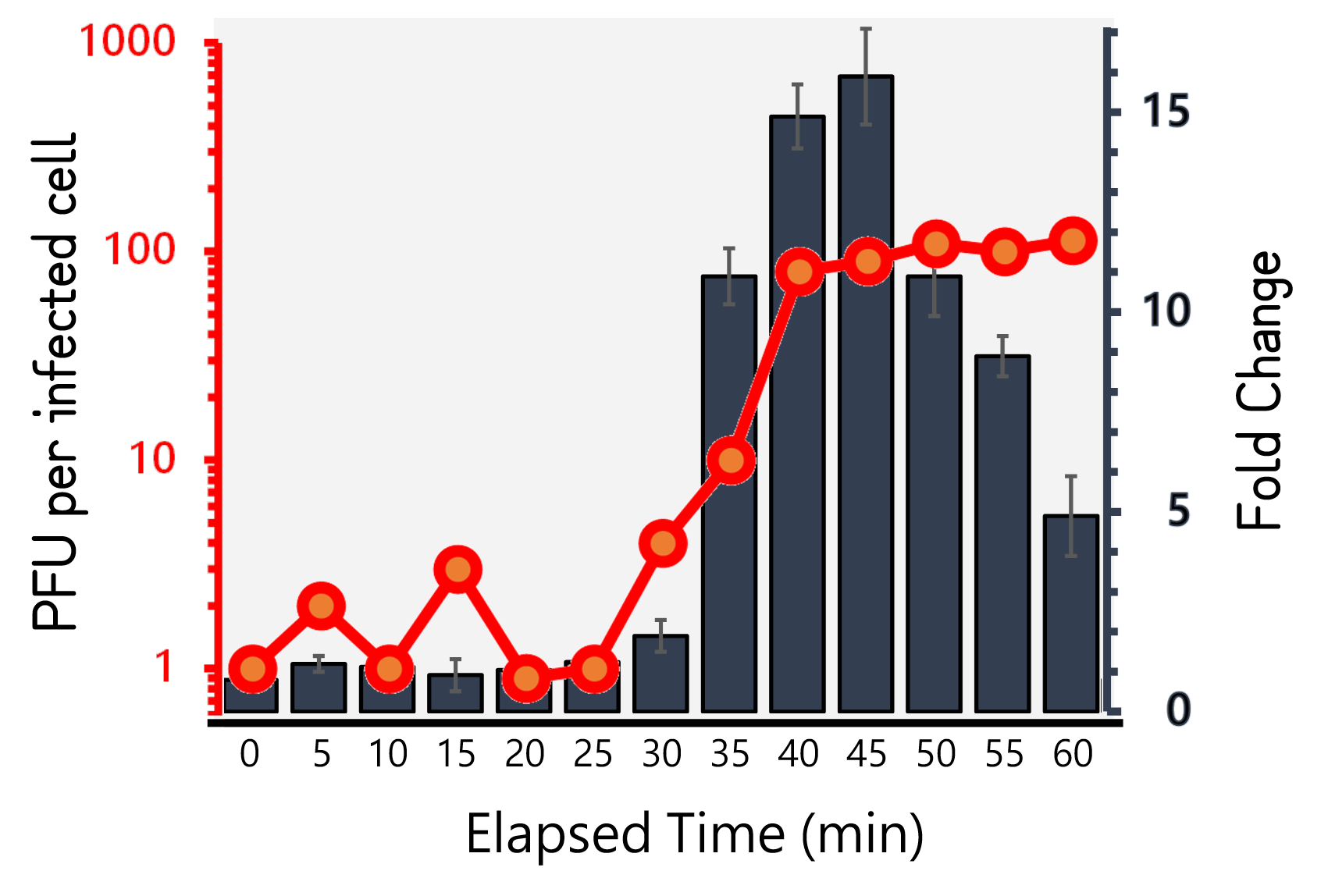
ThisTherefore, we performed the experiment to figure out the latent time and burst size of our Salmonella phage #ST1::Ф29 and the time course of RCA signals during phage infection in Salmonella. 105 cells/ml of Salmonella were infected by phage #ST1::Ф29 at MOI=1. The lysates collected by an interval of 5 min until 60 min and subjected to plaque assays and RCA test. As Fig. 8 demonstrated, the phages were released at around 40 min (latent time) to a plateau level with a burst size of average 98.4±14. Interestingly, the RCA signal increased dramatically at 35 min, achieved a high level around 40-45 min, and dropped significantly thereafter, that are consistent with our speculation of the correlation between phage lysis time (latent time) and phi29 DNA polymerase protein functionality.
Figure 8 | Salmonella phage #ST1::Ф29 latent time (min) and burst size (PFU per infected cell, the left Y axis) at MOI=1, and the relationship to RCA assay (fold change, the right Y axis). Phage-infected Salmonella lysates were harvested for 1 hour at an interval of 5 min. The lysates were subjected to plaque assays and RCA reaction followed by stained with EvaGreen dye. The burst sizes were counted by numbers of plaques. The RCA signal were read at Ex/Em=500/530 nm and divided by the background level without phi29 DNA polymerase.
Detection limit with our hardware
ThisImagine an application of real Salmonella diagnosis in a food or drink. A contaminated sample collected in a large volume may find 1-100 CFU/ml of bacteria[13]. Large volume and bacterial density are key parameters to affect the result of Salmonella detection with our product.
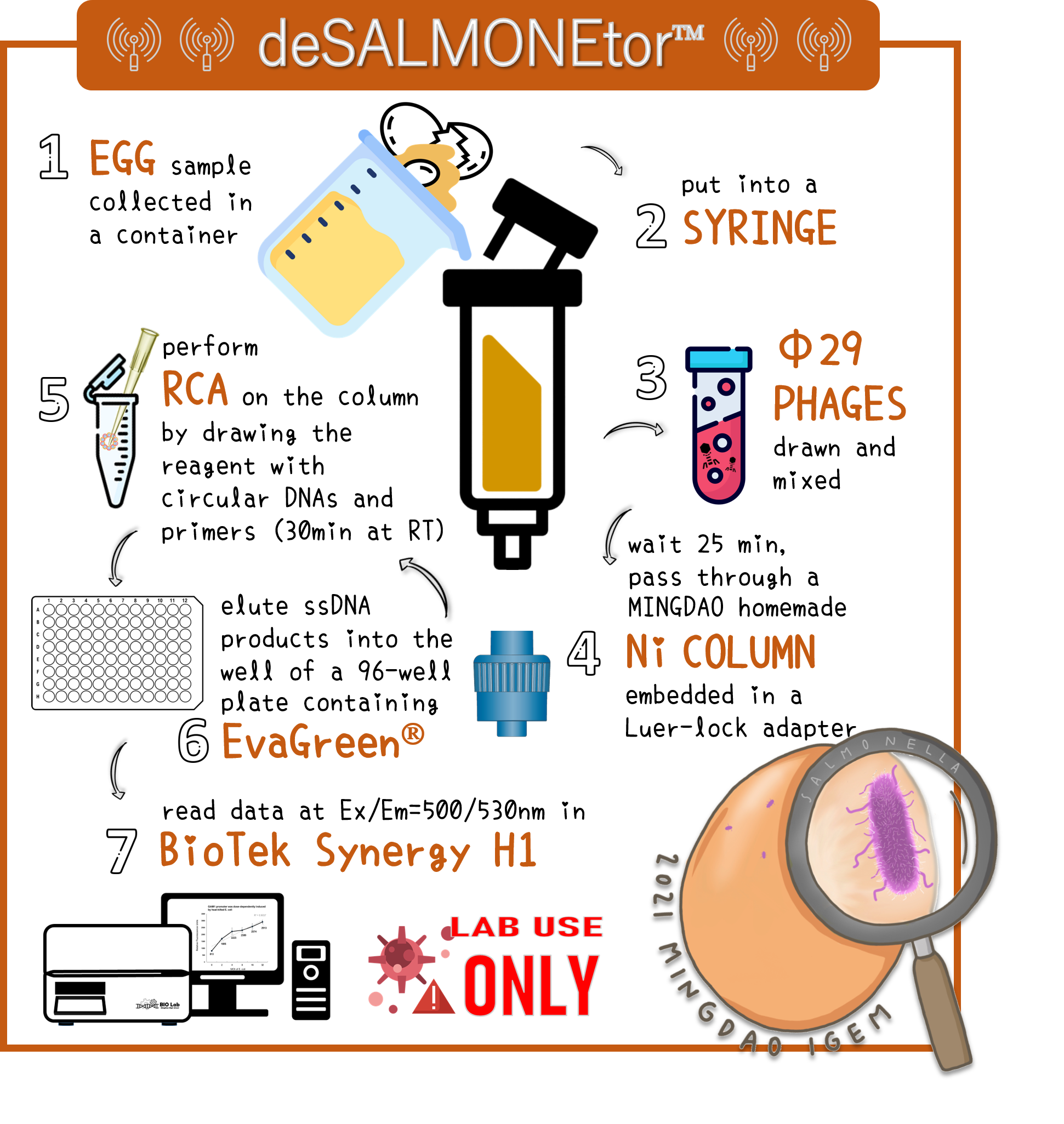
ThisTo overcome the large volume of a sample, we were inspired by Nickel column purification, in which His-tagged phi29 DNA polymerase were bound. Therefore, we think this method may enrich the His-phi29 DNA polymerase from the sample of large volume. Go to our page of (HARDWARE) for such a design. And the schematic diagram was shown here.
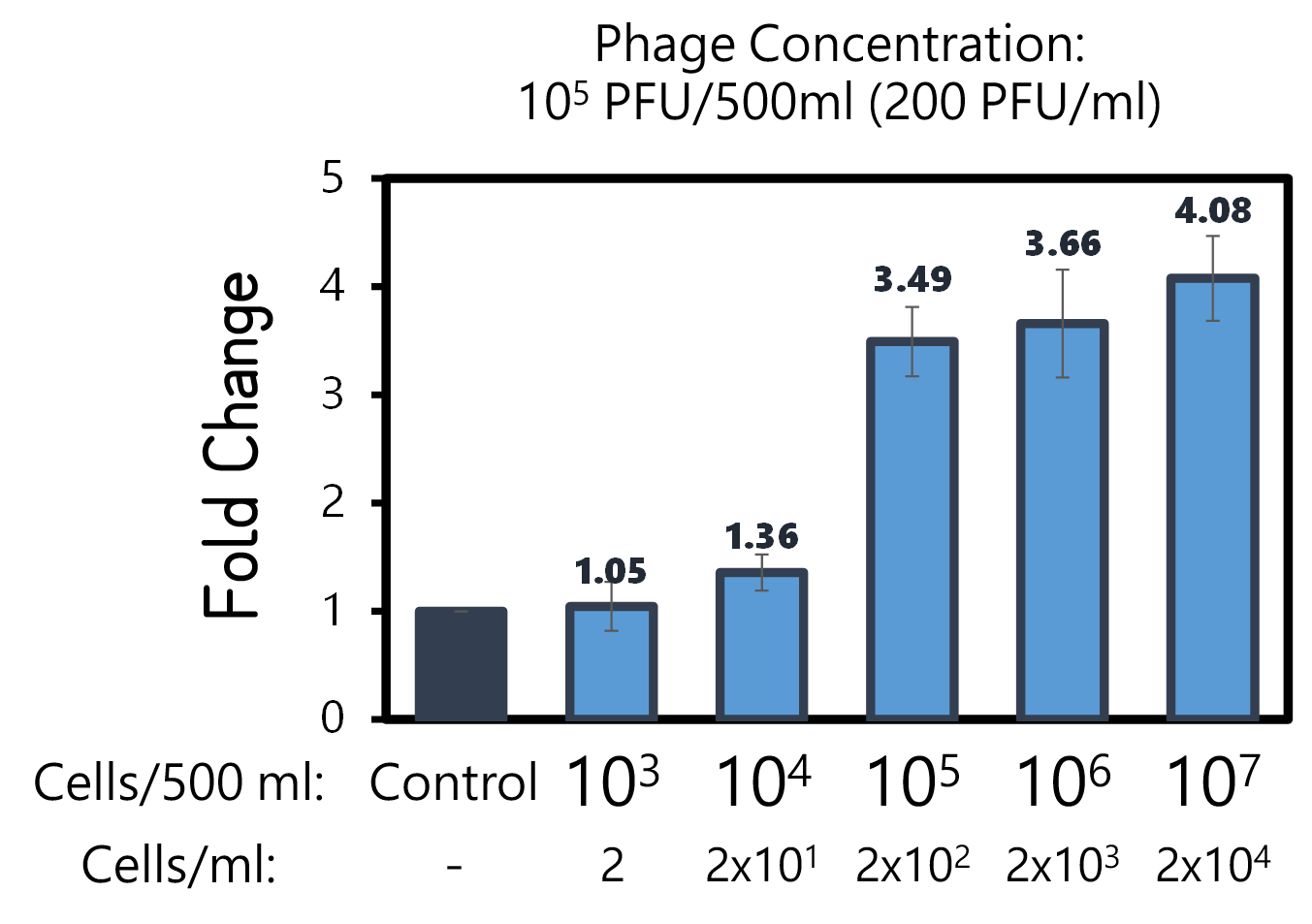
ThisTo examine the feasibility, we made a serial dilution of Salmonella from 107 to 103 cells in a beaker of 500ml water and prepared the water without bacteria as a control. The water were mixed with Salmonella phage #ST1::Ф29 at the concentration of 105 PFU/500ml at room temperature for 25 min. The 3D-printed Luer locker embedded a mini Ni-column was assembled onto a syringe, followed by repeatedly drawing up and pushing back the water in order to pass through the Ni-column. Then, the RCA materials were drawn onto the Ni-column. If His-phi29 DNA polymerase is present, the RCA reaction may be turned on. After 30 min incubation for RCA reaction, the mixtures were push back into a well of a 96-well black plate containing EvaGreen Dye in a total volume of 50 μl. The fluorescence signals were measured at Ex/Em=500/530 nm. Significant RCA signals began to appear in 2x102 bacterial cells/ml (Fig. 9). 20 cells/ml can be detected with a slight enhanced signal that is able to be distinguished from the background. However, we can’t measure the cell density under 10 cells/ml of a liquid to be examined.
Figure 9 | Salmonella test at various concentrations between 2-2x104 cells/ml in 500ml water with engineered Salmonella phage carrying His-phi29 DNA polymerase gene at the concentration of 200 PFU/ml. RCA was performed on the embedded Ni-column in a 3D-printerd Luer lock adapter. The amplified DNAs were stained with EvaGreen Dye and measured at Ex/Em=500/530 nm in a microplate reader (BioTek Synergy H1).
Comparison with current methods
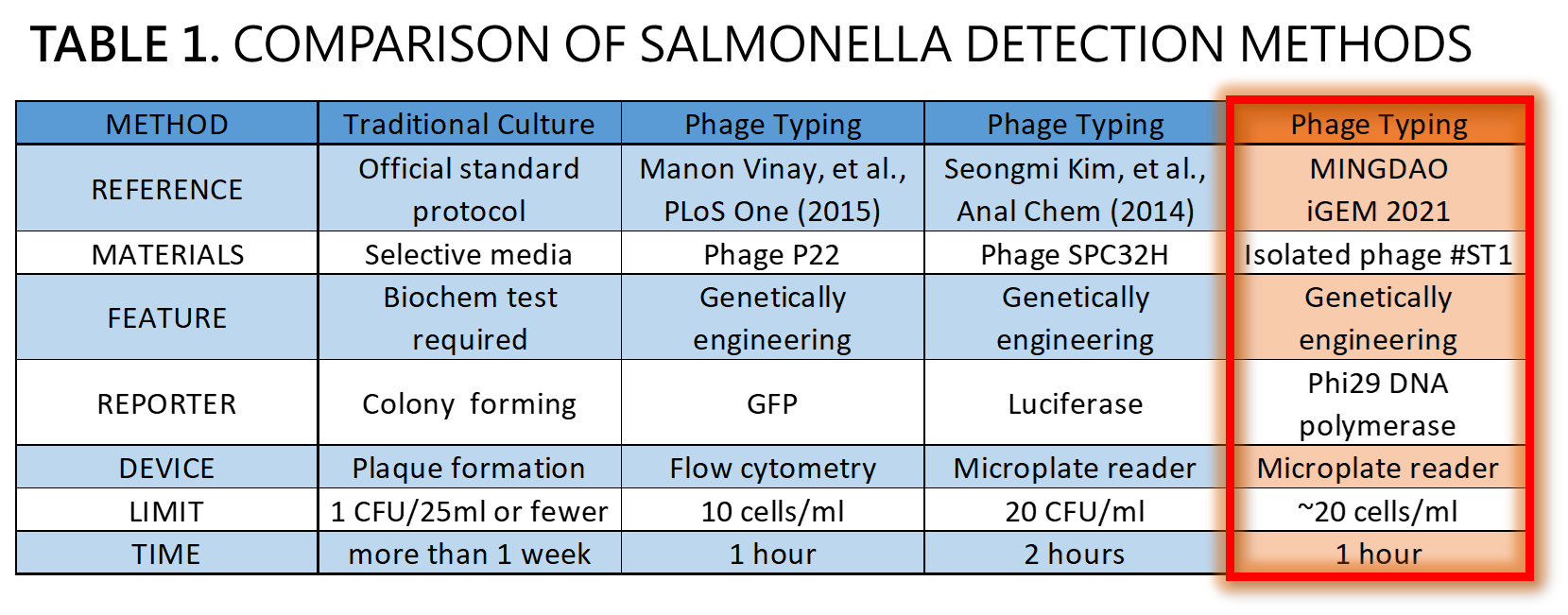
ThisWe demonstrated a proof of concept that it is possible to detect Salmonella by an isolated genetically engineered Salmonella-specific phage carrying phi29 DNA polymerase gene. The His-tagged phi29 DNA polymerase can be enriched from a large volume of a sample onto the homemade mini Ni-column in a Luer-lock adaptor format, where isothermal RCA may be triggered in the presence of the extraordinarily processive phi29 DNA polymerase at room temperature in a time as short as within 30 min. This</span>Bacteriophages possess features that can produce large amounts of phage progeny (the burst size, usually dozens to hundreds PFU per infected bacterium) to release by killing bacterial host in a short time (the latent time, usually 20 min to 1 hr). We mathematically modeled the latent time (min) as a function of cell density (cells/ml) and applied it to simulate a possible real condition using reporter phage to detect Salmonella in a poisoning case. Go to our MODEL page for a detail. We predicted the optimal bacterial lysis time (latent time) for RCA is between 16.545 and 24.427 min for 1 to 100 cells/ml of bacteria. Finally, we developed a hardware of 3D-printed Luer-lock adapter containing mini Ni-column. We used it to measure the Salmonella in 500 ml of water. We can detect 200 bacterial cells/ml within 1 hour. About 20 cells/ml of bacteria may be the most probable limit of detection in our device that generated a slight but distinguishable signal compared to the background level of no bacteria control. Compared to traditional Salmonella tests and published engineered reporter phage, we think our product is better than traditional method in term of test time, and our product is comparable and competitive in terms of test limit and time to the current designer phages carrying reporter genes[14][15][16] (Table 1).
Reference
- ↑ 1.0 1.1 Urasaki A, Morvan G, Kawakami K. Functional dissection of the Tol2 transposable element identified the minimal cis-sequence and a highly repetitive sequence in the subterminal region essential for transposition. Genetics. 2006 Oct;174(2):639-49. doi: 10.1534/genetics.106.060244.
- ↑ Kawakami K. Tol2: a versatile gene transfer vector in vertebrates. Genome Biol. 2007;8 Suppl 1(Suppl 1):S7. doi: 10.1186/gb-2007-8-s1-s7
- ↑ Ni J, Wangensteen KJ, Nelsen D, Balciunas D, Skuster KJ, Urban MD, Ekker SC. Active recombinant Tol2 transposase for gene transfer and gene discovery applications. Mob DNA. 2016 Mar 31;7:6. doi: 10.1186/s13100-016-0062-z.
- ↑ Garenne D, Noireaux V. Cell-free transcription-translation: engineering biology from the nanometer to the millimeter scale. Curr Opin Biotechnol. 2019 Aug;58:19-27. doi: 10.1016/j.copbio.2018.10.007.
- ↑ 5.0 5.1 Rustad M, Eastlund A, Marshall R, Jardine P, Noireaux V. Synthesis of Infectious Bacteriophages in an E. coli-based Cell-free Expression System. J Vis Exp. 2017 Aug 17;(126):56144. doi: 10.3791/56144.
- ↑ Khattak S, Murawala P, Andreas H, Kappert V, Schuez M, Sandoval-Guzmán T, Crawford K, Tanaka EM. Optimized axolotl (Ambystoma mexicanum) husbandry, breeding, metamorphosis, transgenesis and tamoxifen-mediated recombination. Nat Protoc. 2014 Mar;9(3):529-40. doi: 0.1038/nprot.2014.040.
- ↑ Marshall R, Noireaux V. Synthetic Biology with an All E. coli TXTL System: Quantitative Characterization of Regulatory Elements and Gene Circuits. Methods Mol Biol. 2018;1772:61-93. doi: 10.1007/978-1-4939-7795-6_4.
- ↑ Tinafar A, Jaenes K, Pardee K. Synthetic Biology Goes Cell-Free. BMC Biol. 2019 Aug 8;17(1):64. doi: 10.1186/s12915-019-0685-x.
- ↑ 9.0 9.1 Shin J, Jardine P, Noireaux V. Genome replication, synthesis, and assembly of the bacteriophage T7 in a single cell-free reaction. ACS Synth Biol. 2012 Sep 21;1(9):408-13. doi: 10.1021/sb300049p.
- ↑ 10.0 10.1 Ni J, Wangensteen KJ, Nelsen D, Balciunas D, Skuster KJ, Urban MD, Ekker SC. Active recombinant Tol2 transposase for gene transfer and gene discovery applications. Mob DNA. 2016 Mar 31;7:6. doi: 10.1186/s13100-016-0062-z
- ↑ Abedon ST. Selection for bacteriophage latent period length by bacterial density: A theoretical examination. Microb Ecol. 1989 Sep;18(2):79-88. doi: 10.1007/BF02030117.
- ↑ Atel, IR, and K. K. Rao. Bacteriophage burst size as a function of multiplicity of infection. Current Science 1984 53(4): 198–200.
- ↑ Gwimbi P, George M, Ramphalile M. Bacterial contamination of drinking water sources in rural villages of Mohale Basin, Lesotho: exposures through neighbourhood sanitation and hygiene practices. Environ Health Prev Med. 2019 May 15;24(1):33. doi: 10.1186/s12199-019-0790-z.
- ↑ Smartt AE, Ripp S. Bacteriophage reporter technology for sensing and detecting microbial targets. Anal Bioanal Chem. 2011 May;400(4):991-1007. doi: 10.1007/s00216-010-4561-3
- ↑ Vinay M, Franche N, Grégori G, Fantino JR, Pouillot F, Ansaldi M. Phage-Based Fluorescent Biosensor Prototypes to Specifically Detect Enteric Bacteria Such as E. coli and Salmonella enterica Typhimurium. PLoS One. 2015 Jul 17;10(7):e0131466. doi: 10.1371/journal.pone.0131466.
- ↑ Kim S, Kim M, Ryu S. Development of an engineered bioluminescent reporter phage for the sensitive detection of viable Salmonella typhimurium. Anal Chem. 2014 Jun 17;86(12):5858-64. doi: 10.1021/ac500645c.
Note: The map was generated and sponsored by SnapGene.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Plasmid lacks a prefix.
Plasmid lacks a suffix.
Illegal EcoRI site found at 3452
Illegal SpeI site found at 2
Illegal PstI site found at 16
Illegal NotI site found at 9
Illegal NotI site found at 29
Illegal NotI site found at 3458 - 21INCOMPATIBLE WITH RFC[21]Plasmid lacks a prefix.
Plasmid lacks a suffix.
Illegal EcoRI site found at 3452
Illegal BglII site found at 3340
Illegal XhoI site found at 3419 - 23INCOMPATIBLE WITH RFC[23]Illegal prefix found at 3452
Illegal suffix found at 2 - 25INCOMPATIBLE WITH RFC[25]Illegal prefix found at 3452
Plasmid lacks a suffix.
Illegal XbaI site found at 3467
Illegal SpeI site found at 2
Illegal PstI site found at 16
Illegal NgoMIV site found at 642 - 1000INCOMPATIBLE WITH RFC[1000]Plasmid lacks a prefix.
Plasmid lacks a suffix.
Illegal BsaI site found at 1818
Illegal SapI.rc site found at 2900