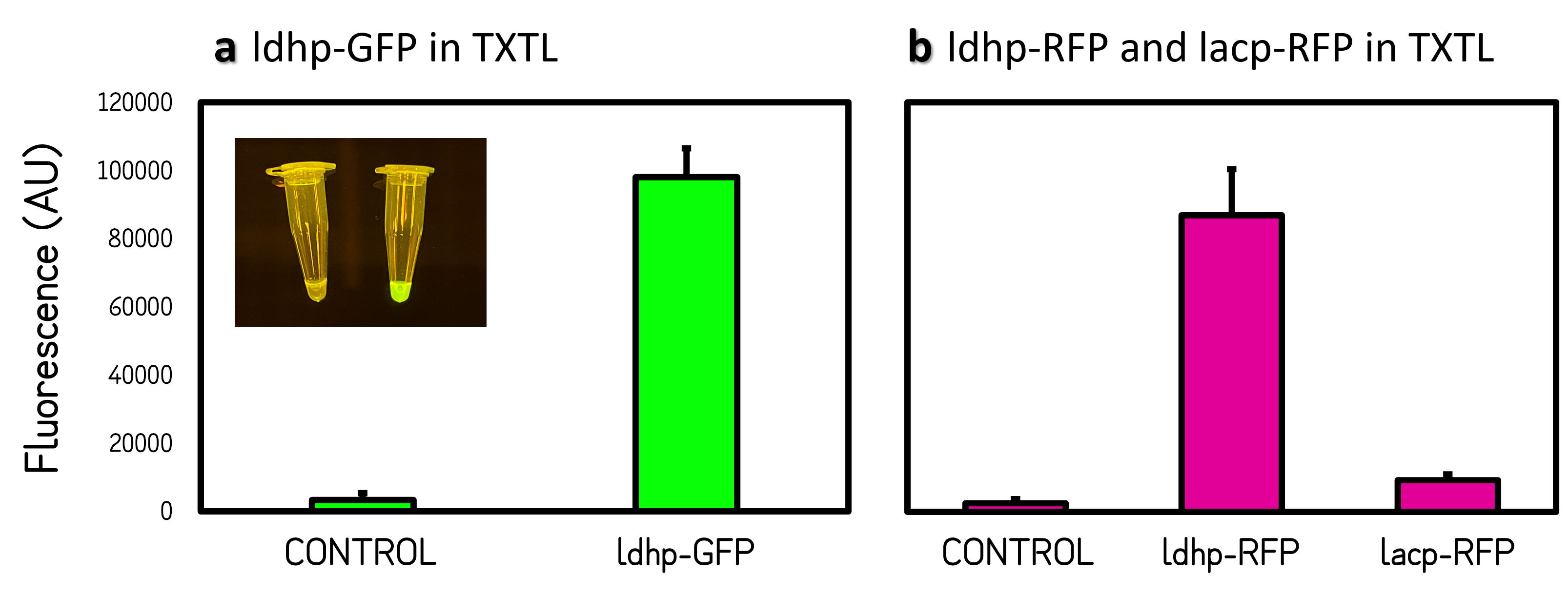Part:BBa_K3728006
ldhp-RFP-Tr/pTol2
ThisTol2 transposon system is highly used in zebrafish transgenesis. The transposase protein (TPase) is from the Medaka fish (Oryzias latipes) aka Japanese rice fish, which catalyzes the transposition of the Tol2 elements through cut-and-paste mechanism. The minimal transposable Tol2 sequence (mTol2) contains 200-bp left arm and 150-bp right arm[1]. Up to 11kb DNA insert between Tol2 sequence can be integrated into the genome of nearly all vertebrates including zebrafish, frog, chicken, mouse, and human [2].
ThisA further application in synthetic biology was demonstrated by Jun Ni, et. al.[3], in which the recombinant TPase protein is fully functional in HeLa cell line and Zebrafish germline cells. In addition, the TPase can be expressed under T7 promoter in E. coli BL21 and purified with N-terminal 6xHis tag. The transposase is active in vitro and mediated the integration of DNA fragments between plasmids with Tol2 elements.
ThisIn our study, we constructed BioBrick Parts of TPase (Part:BBa_K3728000) and the BioBrick compatible Tol2 vectors (Part:BBa_K3728003) with reporter (KanR:Part:BBa_K3728004; GFP:Part:BBa_K3728005; RFP:Part:BBa_K3728006; amilCP:Part:BBa_K3728007) and Phi29 DNA polymerase genes (Part:BBa_K3728008). We prepared the In vitro transcription-translation (TXTL) system [4][5]and expressed the functional reporter proteins. The recombinant TPase and Phil29 DNA polymerase with His tag were expressed in E. coli BL21. The purified proteins were functional in the plasmid integration assay and rolling circle amplification(RCA) application, respectively.
Note: The map was generated and sponsored by SnapGene.
CHARACTERIZATION - TXTL & REPORTER ASSAY
Cell-free TXTL system
ThisIn vitro transcription and translation (TXTL) is a convenient cell-free system that has increasingly been developed to apply in synthetic biology[6][7]. In addition to achieve biosafety level, TXTL becomes powerful in prototype characterization of genetic parts, devices and circuits. Moreover, TXTL is particularly useful to express and purify proteins which are toxic, insoluble or unstable in cell-based system. Furthermore and amazingly, Dr. Vincent Noireau’s lab has demonstrated cell-free TXTL application in infectious bacteriophage production, in which T7 phage (40kbp, 77 genes, dsDNA) and T4 phage (170kbp, 289 genes, dsDNA) genome replication, synthesis, assembly can be performed in vitro just in a single test tube[8][5]
Promoter Activity
ThisTo test the TPase activity and Tol2 transposon system, we inserted a kanamycin resistance gene (KanR) cassette (Part:BBa_K3376004) and the reporters of ldhp-GFP-Tr(Part:BBa_K3376005), ldhp-RFP-Tr(Part:BBa_K3376006) and ldhp-amilCP-Tr(Part:BBa_K3376007) between the transposable elements on the pTol2 vector. ldhp is a constitutive and broad-host-range promoter, which was originally cloned and driving the lactate dehydrogenase gene in S. mutans. We have characterized the ldhp activities in S. mutans and E. coli in our project of iGEM 2020, as well as Salmonella and TXTL in this project of iGEM 2021.(Part:BBa_K3376000)
ThisThe GFP and RFP fluorescence intensities driven by ldhp on pTol2 vectors were measured at high level in TXTL reaction (Fig. 2). The strong GFP fluorescence can even be visualized by naked eyes under a Blue LED Illuminator. Compared the activities of ldhp to lac promoter (lacp), lacp is inhibited in TXTL because the extracts of E. coli Rosetta 2 (DE3) contains LacI repressor, which can be relieved by IPTG induction or using E. coli DH5α as extracts.
Figure 2 | Promoter activities on pTol2 vector in TXTL. GFP fluorescence was measured at Ex/Em = 500/530 nm using a microplate reader of BioTek Synergy H1. RFP was at Ex/Em = 586/611 nm. KanR/pTol2 in TXTX was set as a background control. AU means arbitrary unit. (a) ldhp-GFP-Tr/pTol2 activity in TXTL. The inset photo was captured under a blue LED light. (b) ldhp-RFP-Tr/pTol2 and J04450/pTol2 (i.e., lacp-RFP-Tr) in TXTL.
Note: The map was generated and sponsored by SnapGene.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 725
Illegal AgeI site found at 837 - 1000COMPATIBLE WITH RFC[1000]
- ↑ Urasaki A, Morvan G, Kawakami K. Functional dissection of the Tol2 transposable element identified the minimal cis-sequence and a highly repetitive sequence in the subterminal region essential for transposition. Genetics. 2006 Oct;174(2):639-49. doi: 10.1534/genetics.106.060244.
- ↑ Kawakami K. Tol2: a versatile gene transfer vector in vertebrates. Genome Biol. 2007;8 Suppl 1(Suppl 1):S7. doi: 10.1186/gb-2007-8-s1-s7
- ↑ Ni J, Wangensteen KJ, Nelsen D, Balciunas D, Skuster KJ, Urban MD, Ekker SC. Active recombinant Tol2 transposase for gene transfer and gene discovery applications. Mob DNA. 2016 Mar 31;7:6. doi: 10.1186/s13100-016-0062-z.
- ↑ Garenne D, Noireaux V. Cell-free transcription-translation: engineering biology from the nanometer to the millimeter scale. Curr Opin Biotechnol. 2019 Aug;58:19-27. doi: 10.1016/j.copbio.2018.10.007.
- ↑ 5.0 5.1 Rustad M, Eastlund A, Marshall R, Jardine P, Noireaux V. Synthesis of Infectious Bacteriophages in an E. coli-based Cell-free Expression System. J Vis Exp. 2017 Aug 17;(126):56144. doi: 10.3791/56144.
- ↑ Marshall R, Noireaux V. Synthetic Biology with an All E. coli TXTL System: Quantitative Characterization of Regulatory Elements and Gene Circuits. Methods Mol Biol. 2018;1772:61-93. doi: 10.1007/978-1-4939-7795-6_4.
- ↑ Tinafar A, Jaenes K, Pardee K. Synthetic Biology Goes Cell-Free. BMC Biol. 2019 Aug 8;17(1):64. doi: 10.1186/s12915-019-0685-x.
- ↑ Shin J, Jardine P, Noireaux V. Genome replication, synthesis, and assembly of the bacteriophage T7 in a single cell-free reaction. ACS Synth Biol. 2012 Sep 21;1(9):408-13. doi: 10.1021/sb300049p.
| None |