Difference between revisions of "Part:BBa K2656106"
Alejovigno (Talk | contribs) |
Alejovigno (Talk | contribs) |
||
| (2 intermediate revisions by the same user not shown) | |||
| Line 13: | Line 13: | ||
<ul> | <ul> | ||
<li></html>[https://parts.igem.org/Part:BBa_K2656005 BBa_K2656005]: the [https://parts.igem.org/Part:J23102 J23102] constitutive promoter in its Golden Braid compatible version <html></li> | <li></html>[https://parts.igem.org/Part:BBa_K2656005 BBa_K2656005]: the [https://parts.igem.org/Part:J23102 J23102] constitutive promoter in its Golden Braid compatible version <html></li> | ||
| − | <li></html>[https://parts.igem.org/Part:BBa_K2656009 BBa_K2656009]: the [https://parts.igem.org/Part: | + | <li></html>[https://parts.igem.org/Part:BBa_K2656009 BBa_K2656009]: the [https://parts.igem.org/Part:BBa_B0030 B0030] strong ribosome biding site in its Golden Braid compatible version <html></li> |
<li></html> [https://parts.igem.org/Part:BBa_K2656022 BBa_K2656022]: the [https://parts.igem.org/Part:BBa_K2656022 BBa_K2656022] GFPmut3b sequence in its Golden Braid standardized version <html></li> | <li></html> [https://parts.igem.org/Part:BBa_K2656022 BBa_K2656022]: the [https://parts.igem.org/Part:BBa_K2656022 BBa_K2656022] GFPmut3b sequence in its Golden Braid standardized version <html></li> | ||
<li></html> Terminator [https://parts.igem.org/Part:BBa_K2656026 BBa_K2656026]: the [https://parts.igem.org/Part:BBa_B0015 B0015] transcriptional terminator in its Golden Braid compatible version<html></li> | <li></html> Terminator [https://parts.igem.org/Part:BBa_K2656026 BBa_K2656026]: the [https://parts.igem.org/Part:BBa_B0015 B0015] transcriptional terminator in its Golden Braid compatible version<html></li> | ||
| Line 25: | Line 25: | ||
{|class='wikitable' | {|class='wikitable' | ||
| − | |colspan=4|Table 1. Optimized parameters for the | + | |colspan=4|Table 1. Optimized parameters for the BBa_K2656106. |
|- | |- | ||
|'''Parameter''' | |'''Parameter''' | ||
| Line 31: | Line 31: | ||
|- | |- | ||
|Constitutive transcription rate CR | |Constitutive transcription rate CR | ||
| − | |CR = | + | |CR = 539.5 min-1 |
|- | |- | ||
| Dilution rate μ | | Dilution rate μ | ||
| − | |μ = 0. | + | |μ = 0.0072 min-1 |
|- | |- | ||
|} | |} | ||
| − | |||
Latest revision as of 01:50, 18 October 2018
GFP transcriptional unit 2
Transcriptional unit assembled with a one-pot [http://2018.igem.org/Team:Valencia_UPV/Design Level 1] Golden Gate reaction using BsaI type IIS endonuclease (Figure 1).
It is a composite part of our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection#com BioArt DNA toolkit palette: Printone ], as it can be used to express a green color with medium intensity.
This transcriptional unit is composed of the following standardized parts from our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection Part Collection]:
- BBa_K2656005: the J23102 constitutive promoter in its Golden Braid compatible version
- BBa_K2656009: the B0030 strong ribosome biding site in its Golden Braid compatible version
- BBa_K2656022: the BBa_K2656022 GFPmut3b sequence in its Golden Braid standardized version
- Terminator BBa_K2656026: the B0015 transcriptional terminator in its Golden Braid compatible version
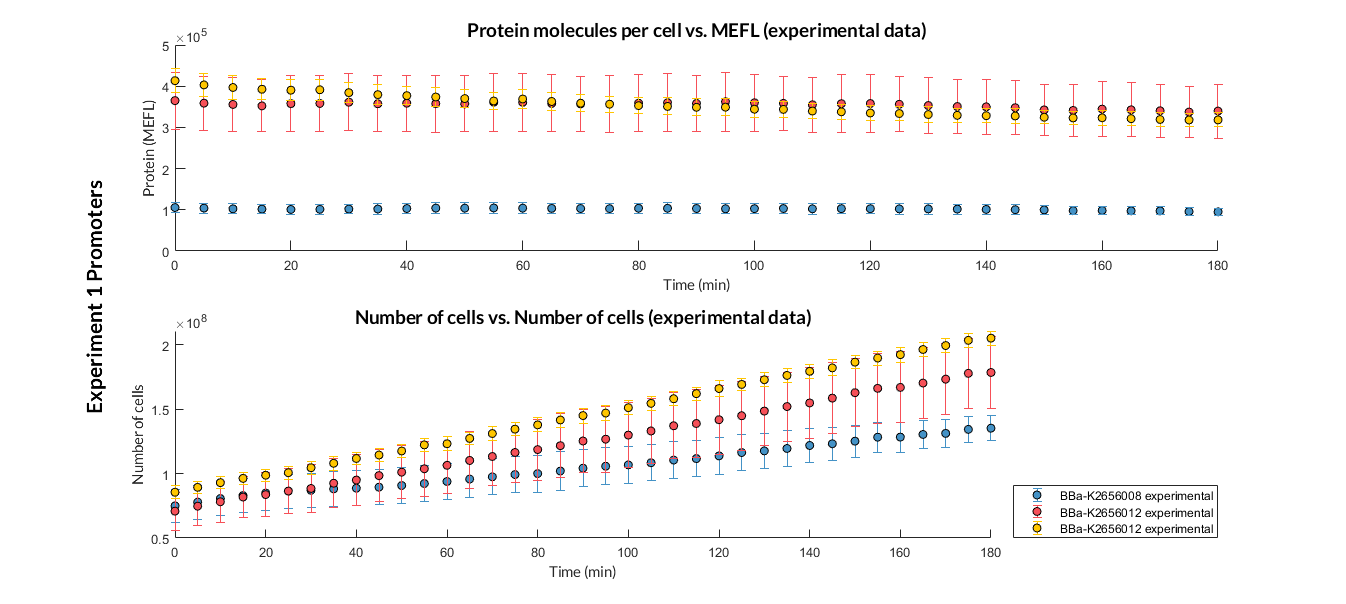
This transcriptional unit has been used to characterize the constitutive BBa_K2656005 promoter. Following this [http://2018.igem.org/Team:Valencia_UPV/Modeling#http://2018.igem.org/Team:Valencia_UPV/Experiments#exp_protocol experimental protocol], we have obtained the parameters to valide our [http://2018.igem.org/Team:Valencia_UPV/Modeling#models constitutive model] and so the results of the next graph have been obtained after an [http://2018.igem.org/Team:Valencia_UPV/Modeling#optimization optimization] and decision-making process, in which the optimal parameters have been selected and the constituent model for each objective has been simulated and compared with the experimental data.
| Table 1. Optimized parameters for the BBa_K2656106. | |||
| Parameter | Value | ||
| Constitutive transcription rate CR | CR = 539.5 min-1 | ||
| Dilution rate μ | μ = 0.0072 min-1 | ||
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 11
Illegal NheI site found at 34 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]