Difference between revisions of "Cell-free chassis/Commercial E. coli T7 S30"
(→Commercial ''E. coli'' T7 S30) |
|||
| (7 intermediate revisions by 3 users not shown) | |||
| Line 1: | Line 1: | ||
| − | + | [[Cell-free chassis|< Back to Cell-free chassis]] | |
| + | |||
{| border="1" width="100%" | {| border="1" width="100%" | ||
| Line 6: | Line 7: | ||
|- | |- | ||
|<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30 <font face=georgia color=#000066 size=4>Chassis description</font>]</center> || The Commercial ''E. coli'' T7 S30 extract was purchased from Promega. It was prepared by modifications of the method described by Zubay et al (1980). '''''E. coli'' strain B''' deficient in OmpT endoproteinase and lon protease activity was used. | |<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30 <font face=georgia color=#000066 size=4>Chassis description</font>]</center> || The Commercial ''E. coli'' T7 S30 extract was purchased from Promega. It was prepared by modifications of the method described by Zubay et al (1980). '''''E. coli'' strain B''' deficient in OmpT endoproteinase and lon protease activity was used. | ||
| − | The simple constitutive gene expression device [https://parts.igem.org/wiki/index.php/Part: | + | The simple constitutive gene expression device [https://parts.igem.org/wiki/index.php/Part:BBa_I719015 BBa_I719015] has been used to characterize this cell-free chassis |
|- | |- | ||
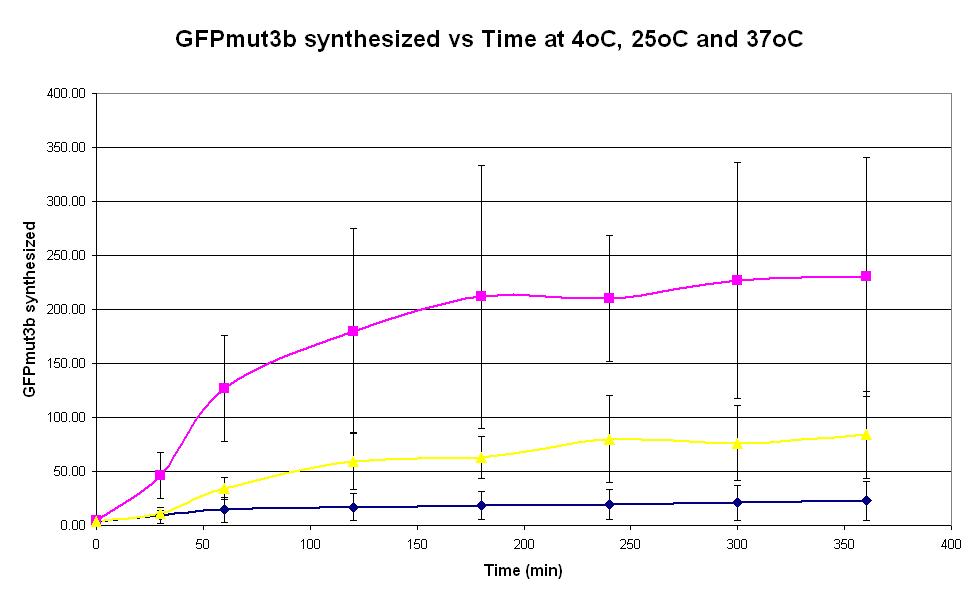
| − | |<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30/Optimum_Temp <font face=georgia color=#000066 size=4>Temperature dependence</font>]</center> || [[Image:T7_Optimum_Temp.JPG|thumb|200px|center|]] '''Graph 1. Graph of GFPmut3b synthesized (pmol) against time (minutes) at 4oC (dark blue), 25oC (pink) and 37oC.''' The simple constitutive gene expression device [https://parts.igem.org/wiki/index.php/Part: | + | |<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30/Optimum_Temp <font face=georgia color=#000066 size=4>Temperature dependence</font>]</center> || [[Image:T7_Optimum_Temp.JPG|thumb|200px|center|]] '''Graph 1. Graph of GFPmut3b synthesized (pmol) against time (minutes) at 4oC (dark blue), 25oC (pink) and 37oC.''' The simple constitutive gene expression device [https://parts.igem.org/wiki/index.php/Part:BBa_I719015 BBa_I719015] has been used. Each of the three coloured lines show average measurements based on three replicates of the experiment. The error bars represent the standard deviation of the measurements. |
|- | |- | ||
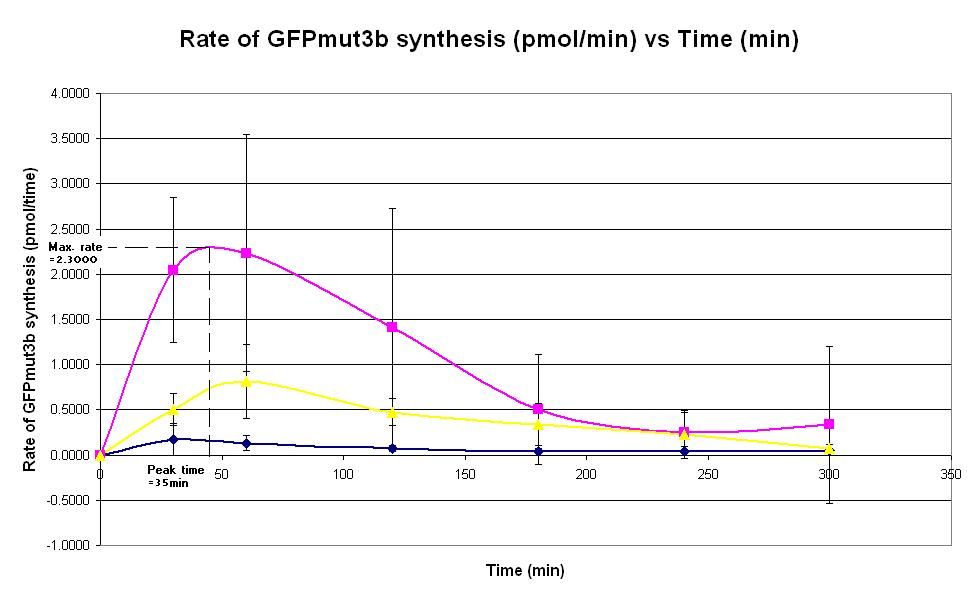
| − | |<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30/Peak_Time <font face=georgia color=#000066 size=4>Peak time</font>]</center> || [[Image:T7_Peak_Time.JPG|thumb|200px|center|]]'''Graph 2. Graph of rate of | + | |<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30/Peak_Time <font face=georgia color=#000066 size=4>Peak time</font>]</center> || [[Image:T7_Peak_Time.JPG|thumb|200px|center|]]'''Graph 2. Graph of rate of GFPmut3b synthesis (pmol per min) against time (minutes) at 4oC (dark blue), 25oC (pink) and 37oC (yellow).''' The simple constitutive gene expression device [https://parts.igem.org/wiki/index.php/Part:BBa_I719015 BBa_I719015] has been used. Each of the three coloured lines show average measurements based on three replicates of the experiment. The error bars represent the standard deviation of the measurements. |
|- | |- | ||
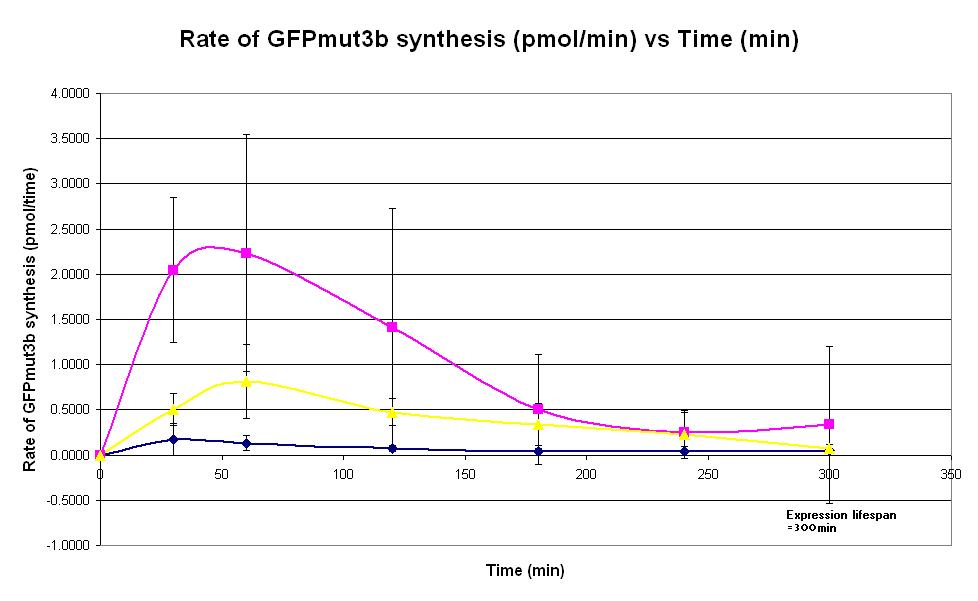
| − | |<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30/Expression_Lifespan <font face=georgia color=#000066 size=4>Expression lifespan</font>]</center> || [[Image: | + | |<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30/Expression_Lifespan <font face=georgia color=#000066 size=4>Expression lifespan</font>]</center> || [[Image:T7_Expression_Lifespan.JPG|thumb|200px|center|]] '''Graph 3. Graph of rate of GFP synthesis (mol per min) against time (minutes) at 4oC (blue), 25oC (red) and 37oC (green).''' The simple constitutive gene expression device [https://parts.igem.org/wiki/index.php/Part:BBa_I719015 BBa_I719015] has been used. Each of the three coloured lines show average measurements based on three replicates of the experiment. The error bars represent the standard deviation of the measurements. |
|- | |- | ||
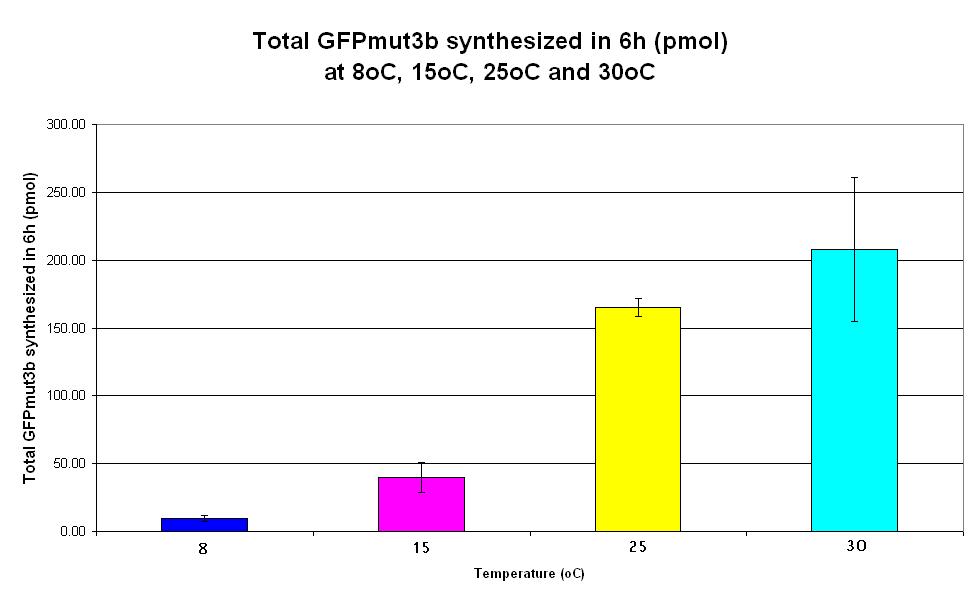
| − | |<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30/Expression_Capacity <font face=georgia color=#000066 size=4>Expression capacity</font>]</center> || [[Image:S30_Expression_Capacity.JPG|thumb|200px|center|]] '''Graph | + | |<center>[https://parts.igem.org/wiki/index.php/Chassis/Cell-Free_Systems/Commercial_E.coli_T7_S30/Expression_Capacity <font face=georgia color=#000066 size=4>Expression capacity</font>]</center> || [[Image:S30_Expression_Capacity.JPG|thumb|200px|center|]] '''Graph 4. Bar chart showing total GFPmut3b synthesized in 6 hours (pmol) at 4oC (dark blue), 25oC (pink) and 37oC (yellow).''' The simple constitutive gene expression device [https://parts.igem.org/wiki/index.php/Part:BBa_I719015 BBa_I719015] has been used. Each of the three coloured bars show average measurements based on three replicates of the experiment. The error bars represent the standard deviation of the measurements. |
|} | |} | ||
==References== | ==References== | ||
Please refer to the [http://www.promega.com/tbs/tb219/tb219.pdf Promega cell extract user guide] for detailed information about the Commercial ''E. coli'' T7 S30 extract. | Please refer to the [http://www.promega.com/tbs/tb219/tb219.pdf Promega cell extract user guide] for detailed information about the Commercial ''E. coli'' T7 S30 extract. | ||
Latest revision as of 19:10, 13 February 2009
| Cell-Free Systems (back to Intro) | Introduction |
| The Commercial E. coli T7 S30 extract was purchased from Promega. It was prepared by modifications of the method described by Zubay et al (1980). E. coli strain B deficient in OmpT endoproteinase and lon protease activity was used.
The simple constitutive gene expression device BBa_I719015 has been used to characterize this cell-free chassis | |
| Graph 1. Graph of GFPmut3b synthesized (pmol) against time (minutes) at 4oC (dark blue), 25oC (pink) and 37oC. The simple constitutive gene expression device BBa_I719015 has been used. Each of the three coloured lines show average measurements based on three replicates of the experiment. The error bars represent the standard deviation of the measurements. | |
| Graph 2. Graph of rate of GFPmut3b synthesis (pmol per min) against time (minutes) at 4oC (dark blue), 25oC (pink) and 37oC (yellow). The simple constitutive gene expression device BBa_I719015 has been used. Each of the three coloured lines show average measurements based on three replicates of the experiment. The error bars represent the standard deviation of the measurements. | |
| Graph 3. Graph of rate of GFP synthesis (mol per min) against time (minutes) at 4oC (blue), 25oC (red) and 37oC (green). The simple constitutive gene expression device BBa_I719015 has been used. Each of the three coloured lines show average measurements based on three replicates of the experiment. The error bars represent the standard deviation of the measurements. | |
| Graph 4. Bar chart showing total GFPmut3b synthesized in 6 hours (pmol) at 4oC (dark blue), 25oC (pink) and 37oC (yellow). The simple constitutive gene expression device BBa_I719015 has been used. Each of the three coloured bars show average measurements based on three replicates of the experiment. The error bars represent the standard deviation of the measurements. |
References
Please refer to the [http://www.promega.com/tbs/tb219/tb219.pdf Promega cell extract user guide] for detailed information about the Commercial E. coli T7 S30 extract.