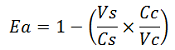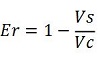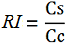Difference between revisions of "Part:BBa B0010:Experience"
AptamersBoi (Talk | contribs) |
|||
| (5 intermediate revisions by 3 users not shown) | |||
| Line 2: | Line 2: | ||
This experience page is provided so that any user may enter their experience using this part.<BR>Please enter | This experience page is provided so that any user may enter their experience using this part.<BR>Please enter | ||
how you used this part and how it worked out. | how you used this part and how it worked out. | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
===Applications of BBa_B0010=== | ===Applications of BBa_B0010=== | ||
| + | |||
| + | *'''PCR Problems:''' Using primers VR and VF2 to PCR B0010 results in excess bands. VR can anneal to B0010, resulting in shorter bands than expected. A full description of the problem is available [[Problems with PCR using VR/VF2|here]]. | ||
===User Reviews=== | ===User Reviews=== | ||
| Line 22: | Line 19: | ||
|}; | |}; | ||
<!-- End of the user review template --> | <!-- End of the user review template --> | ||
| + | |||
| + | {|width='80%' style='border:1px solid gray' | ||
| + | |- | ||
| + | |width='10%'| | ||
| + | <partinfo>BBa_B0015 AddReview 5</partinfo> | ||
| + | <I>iGEM Athens 2019</I> | ||
| + | |width='60%' valign='top'| | ||
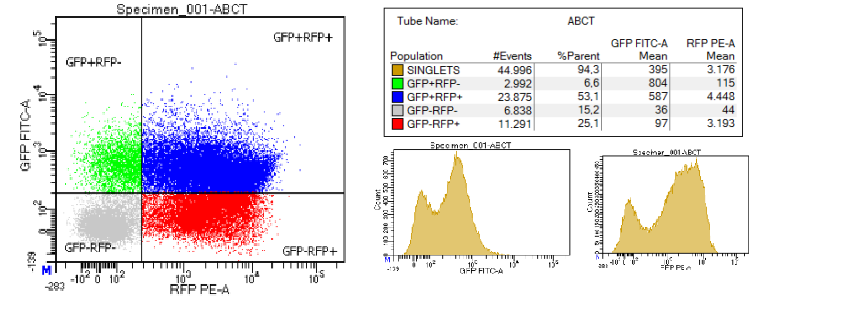
| + | We attempted to measure the efficiency of the terminator by cloning it into the pSB1A10 plasmid and culturing the bacteria for 24 hours (37 degrees Celsius, 200RPM, 0.1% arabinose, 100 µg/mL ampicillin). The culture was pelleted and the pellet was resuspended in 1mL of 1X PBS. From that, 10 μL were diluted in 990 mL of 1X PBS to yield approximately 5 million cells. For FACS, we used BD FACSCanto™ II. During FACS, a low flow rate was used. We used the following excitation and emission wavelengths: GFP: excitation = 488 nm, emission = 510 nm ; RFP: excitation = 488 nm, emission = 584 nm. | ||
| + | [[File:T--Athens--ABCT.png|400px|thumb|center|FACS measurement. The terminator was named ABC terminator for our project.]] | ||
| + | |||
| + | The efficiency is measured through the following formula: Termination efficiency = 1 -{[RFPterm/GFPterm]/[(RFPcontrol/GFPcontrol)mean]}. | ||
| + | This measurement showed that the efficiency of the terminator is 0%. | ||
| + | We also measured the terminator efficiency using the GFP+RFP+ population and the formula: Terminator efficiency = 1 - (Intensity of GFP+RFP+term/Intensity of GFP+RFP+control). | ||
| + | This measurement showed that the terminator efficiency of the terminator is 7%. | ||
| + | |}; | ||
| + | |||
{|width='80%' style='border:1px solid gray' | {|width='80%' style='border:1px solid gray' | ||
|- | |- | ||
| Line 28: | Line 41: | ||
<I>Antiquity</I> | <I>Antiquity</I> | ||
|width='60%' valign='top'| | |width='60%' valign='top'| | ||
| − | This review comes from the old result system and indicates that this part did not work in some test. | + | This review comes from the old result system and indicates that this part did not work in some test. |
|} | |} | ||
| + | |||
| + | |||
| + | {|width='80%' style='border:1px solid gray' | ||
| + | |- | ||
| + | |width='10%'| | ||
| + | <partinfo>BBa_B0010 AddReview 0</partinfo> | ||
| + | <I>Trento iGEM team 2012</I> | ||
| + | |width='60%' valign='top'| | ||
| + | This part was successfully extracted from the distribution kits 2012 and 2011. However, it was not confirmed by restriction digestion nor PCR. We were able to amplify it by PCR from BBa_E0840, that contains the double terminator BBa_B0015, subcloned it into pSB1C3 and confirmed by sequencing. We re-deposited rrnBT1 terminator well-characterized and well-functioning as [[Part:BBa_K731722|BBa_K731722]]. | ||
| + | |||
| + | |||
| + | The characterization of this part was done using the platforms for terminators' characterization [[Part:BBa_K731700|BBa_K731700]], [[Part:BBa_K731710|BBa_K731710]] with both T7 and ''E. coli'' RNA polymerases ''in vivo'' and with T7 RNA polymerase ''in vitro''. | ||
| + | |||
| + | |||
| + | We made some preliminary analyses to define the best procedure to characterize this terminator using [[Part:BBa_K731700|BBa_K731700]] and [[Part:BBa_K731710|BBa_K731710]]. We found that the emission from mVenus was much stronger than the mCherry emission, resulting in some spectral overlap. To improve the quality of the data, off-peak excitation and emission wavelengths were identified that minimized the effect. Therefore, mVenus was excitated at 485 nm (excitation peak is at 515 nm), and the emission of mVenus was collected at its maximum position, i.e. 528 nm. The maximum mCherry excitation wavelength was used (587 nm), but the emission of mCherry was read at 615 nm, as opposed to the maximum at 609 nm. Here is a summary of these wavelengths: | ||
| + | <div style="text-align:center"> | ||
| + | |||
| + | {|border = "1" cellpadding="5" cellspacing="0" align="centre" | ||
| + | | | ||
| + | |Standard Excitation (nm) | ||
| + | |Standard Emission (nm) | ||
| + | |Modified Excitation (nm) | ||
| + | |Modified Emission (nm) | ||
| + | |- | ||
| + | |mCherry | ||
| + | |587 | ||
| + | |609 | ||
| + | |587 | ||
| + | |615 | ||
| + | |- | ||
| + | |mVenus | ||
| + | |515 | ||
| + | |528 | ||
| + | |485 | ||
| + | |528 | ||
| + | |} | ||
| + | </div> | ||
| + | <html> | ||
| + | <p style="width:900px; text-align:justify "><em><strong>TABLE 1.</strong> <b>Standard and modified excitation and emission wavelengths. </b> We adopted these modified excitation and emission wavelengths for both in vivo and iv vitro measurements.<br/> | ||
| + | </em> </p> </html> | ||
| + | |||
| + | |||
| + | The activity of this terminator was measured as the ratio between the two proteins' levels using the equation found in literature for termination efficiency. | ||
| + | Doing this measurements, however, we realized that terminators can have an effect also on mCherry expression, enhancing it. This effect was previously described <cite>Abe</cite>. As a consequence of this unexpected outcome, we defined other two parameters that are here summarized in addition to the literature definition of termination efficiency. | ||
| + | |||
| + | The parameters used to analyze the data are: | ||
| + | |||
| + | apparent termination efficiency, calculated with the equation found in literature <cite>Nojima</cite>, [[Image:FormulaEa.jpg]] | ||
| + | |||
| + | raw termination efficiency, that does not consider the mCherry contribution [[Image:FormulaEr.jpg]] | ||
| + | |||
| + | relative increase in the upstream gene expression, [[Image:RIequation.png]] | ||
| + | |||
| + | |||
| + | where | ||
| + | |||
| + | -''Vs'' is the A206K Venus peak’s intensity of the construct with the terminator of interest inserted in the prefix-suffix linker | ||
| + | |||
| + | -''Vc'' is the A206K Venus peak’s intensity of the control construct without intervening terminator | ||
| + | |||
| + | -''Cs'' is the mCherry peak’s intensity of the construct with the terminator inserted | ||
| + | |||
| + | -''Cc'' is the mCherry peak’s intensity of the control construct | ||
| + | |||
| + | |||
| + | '''''In vivo'' measurements:''' | ||
| + | |||
| + | Our experiments exploited an ''E. coli'' lysogen strain carrying T7 RNA polymerase and lacIq. Additionally, the cells, i.e. ''E. coli'' BL21(DE3) pLysS, also contained a plasmid encoding T7 lysozyme and chloramphenicol resistance. T7 lysozyme is a natural inhibitor of T7 RNA polymerase activity, thus reducing background expression of the target genes. The T7 RNA polymerase is behind a lacUV5 promoter. | ||
| + | |||
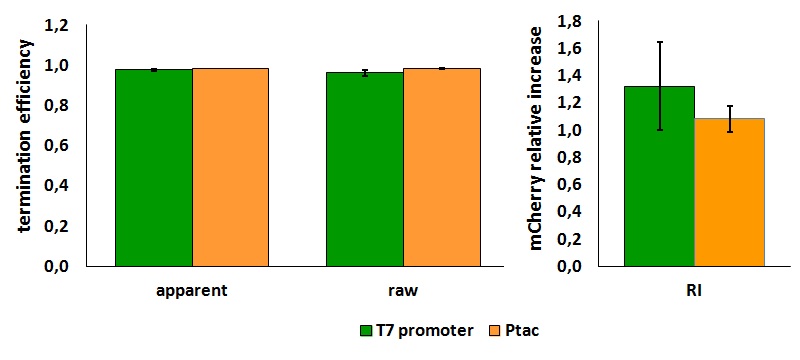
| + | <div style="text-align:center">[[Image:Ecoliterminatorinvivo.jpg]]</div> | ||
| + | |||
| + | |||
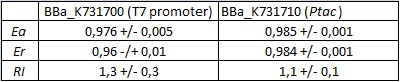
| + | <div style="text-align:center">[[Image:Ecoliterminatortabinvivo.jpg]]</div> | ||
| + | <p style="width:600px; margin-left:150px; margin-bottom:60px; | ||
| + | text-align:justify "><em><strong>FIGURE 1.</strong> '''''E.coli'' terminator's effect on protein expression with two different RNA polymerases'''<br/>The data shown in figure 1 were acquired in two different days. For each day 4 different replicates were measured at different times. | ||
| + | |||
| + | Briefly, BL21(DE3)pLysS were grown in 10 mL of LB until an OD of 0.6 was reached and induced with 0.5 mM IPTG. After 3 hours of induction, 4 separate aliquots of 1 mL were taken and sonicated 3 times for 10 seconds at intervals of 30 seconds. After sonication the samples were diluted 1:2 with PBS 1X directly into a cuvette and incubated overnight at 4°C. Fluorescence measurements were taken with a Cary Eclipse Varian fluorimeter with a window ranging from 450 nm to 700 nm, a slit of 5nm, a voltage of 570V for the characterization with T7 polymerase and of 520V for the measurements with ''E. coli'' RNA polymerase. | ||
| + | |||
| + | </em></p> | ||
| + | |||
| + | '''In vitro measurements:''' | ||
| + | |||
| + | We performed the in vitro analysis only with the T7 RNA polymerase. | ||
| + | |||
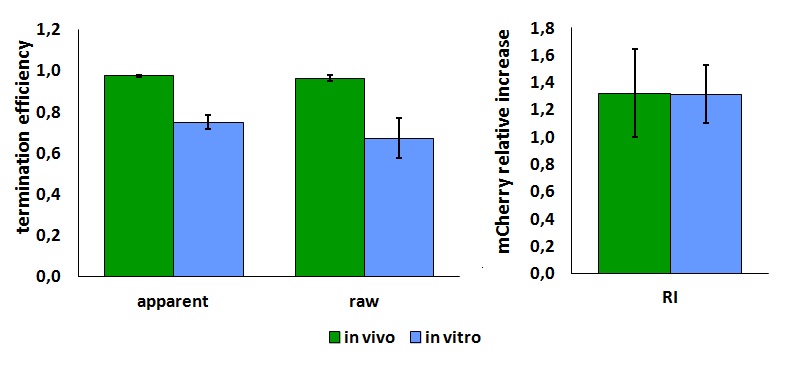
| + | <div style="text-align:center">[[Image:Ecoliterminatorinvitro.jpg]]</div> | ||
| + | |||
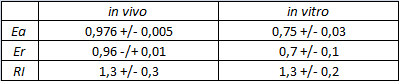
| + | <div style="text-align:center">[[Image:Ecoliterminatortabinvitro.jpg]]</div> | ||
| + | <p style="width:600px; margin-left:150px; margin-bottom:60px; | ||
| + | text-align:justify "><em><strong>FIGURE 2.</strong> '''E.coli terminator's effect on in vitro protein synthesis with the T7 RNA polymerases'''<br/>Cell free measurements were done with the PurExpress kit from New England Biolabs, using 250 ng of DNA previously purified by ethanol precipitation, following the protocol suggested by the manufacturer. Measurements were done with a PTI fluorimeter. | ||
| + | |||
| + | </em> </p> | ||
| + | We submitted also this Part inside both [[Part:BBa_K731700|BBa_K731700]] (T7 promoter backbone) and [[Part:BBa_K731710|BBa_K731710]] (''tac'' promoter backbone) as, respecitively, [[Part:BBa_K731701|BBa_K731701]] and [[Part:BBa_K731702|BBa_K731702]]. We hope that this could be helpful for anyone who want to verify and further delve into our results. | ||
| + | |||
| + | More information can be found in the iGEM Trento 2012 wiki page. | ||
| + | |||
| + | ==References== | ||
| + | <biblio> | ||
| + | #Nojima pmid=16379390 | ||
| + | #Abe pmid=9150882 | ||
| + | </biblio> | ||
| + | |}; | ||
| + | <!-- End of the user review template --> | ||
| + | |||
<!-- DON'T DELETE --><partinfo>BBa_B0010 EndReviews</partinfo> | <!-- DON'T DELETE --><partinfo>BBa_B0010 EndReviews</partinfo> | ||
Latest revision as of 18:18, 21 October 2019
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_B0010
- PCR Problems: Using primers VR and VF2 to PCR B0010 results in excess bands. VR can anneal to B0010, resulting in shorter bands than expected. A full description of the problem is available here.
User Reviews
UNIQ43a1c6e7d56afa5c-partinfo-00000000-QINU
|
•••••
iGEM Athens 2019 |
We attempted to measure the efficiency of the terminator by cloning it into the pSB1A10 plasmid and culturing the bacteria for 24 hours (37 degrees Celsius, 200RPM, 0.1% arabinose, 100 µg/mL ampicillin). The culture was pelleted and the pellet was resuspended in 1mL of 1X PBS. From that, 10 μL were diluted in 990 mL of 1X PBS to yield approximately 5 million cells. For FACS, we used BD FACSCanto™ II. During FACS, a low flow rate was used. We used the following excitation and emission wavelengths: GFP: excitation = 488 nm, emission = 510 nm ; RFP: excitation = 488 nm, emission = 584 nm. The efficiency is measured through the following formula: Termination efficiency = 1 -{[RFPterm/GFPterm]/[(RFPcontrol/GFPcontrol)mean]}. This measurement showed that the efficiency of the terminator is 0%. We also measured the terminator efficiency using the GFP+RFP+ population and the formula: Terminator efficiency = 1 - (Intensity of GFP+RFP+term/Intensity of GFP+RFP+control). This measurement showed that the terminator efficiency of the terminator is 7%. |
|
Antiquity |
This review comes from the old result system and indicates that this part did not work in some test. |
|
Trento iGEM team 2012 |
This part was successfully extracted from the distribution kits 2012 and 2011. However, it was not confirmed by restriction digestion nor PCR. We were able to amplify it by PCR from BBa_E0840, that contains the double terminator BBa_B0015, subcloned it into pSB1C3 and confirmed by sequencing. We re-deposited rrnBT1 terminator well-characterized and well-functioning as BBa_K731722.
TABLE 1. Standard and modified excitation and emission wavelengths. We adopted these modified excitation and emission wavelengths for both in vivo and iv vitro measurements.
The parameters used to analyze the data are: apparent termination efficiency, calculated with the equation found in literature Nojima, raw termination efficiency, that does not consider the mCherry contribution relative increase in the upstream gene expression,
-Vs is the A206K Venus peak’s intensity of the construct with the terminator of interest inserted in the prefix-suffix linker -Vc is the A206K Venus peak’s intensity of the control construct without intervening terminator -Cs is the mCherry peak’s intensity of the construct with the terminator inserted -Cc is the mCherry peak’s intensity of the control construct
Our experiments exploited an E. coli lysogen strain carrying T7 RNA polymerase and lacIq. Additionally, the cells, i.e. E. coli BL21(DE3) pLysS, also contained a plasmid encoding T7 lysozyme and chloramphenicol resistance. T7 lysozyme is a natural inhibitor of T7 RNA polymerase activity, thus reducing background expression of the target genes. The T7 RNA polymerase is behind a lacUV5 promoter.
FIGURE 1. E.coli terminator's effect on protein expression with two different RNA polymerases In vitro measurements: We performed the in vitro analysis only with the T7 RNA polymerase. FIGURE 2. E.coli terminator's effect on in vitro protein synthesis with the T7 RNA polymerases We submitted also this Part inside both BBa_K731700 (T7 promoter backbone) and BBa_K731710 (tac promoter backbone) as, respecitively, BBa_K731701 and BBa_K731702. We hope that this could be helpful for anyone who want to verify and further delve into our results. More information can be found in the iGEM Trento 2012 wiki page. References<biblio>
</biblio> |
UNIQ43a1c6e7d56afa5c-partinfo-00000005-QINU