Difference between revisions of "Part:BBa K404165"
| Line 304: | Line 304: | ||
alt="Image:Freiburg10 NLS mVenus mCherry.png"></p> | alt="Image:Freiburg10 NLS mVenus mCherry.png"></p> | ||
<p class="MsoNormal" style="line-height: normal;"><b><span | <p class="MsoNormal" style="line-height: normal;"><b><span | ||
| − | style="font-size: 10pt;" lang="EN-US">Figure | + | style="font-size: 10pt;" lang="EN-US">Figure 1: Fluorescence |
Microscopy. A) </span></b><span style="font-size: 10pt;" lang="EN-US">Brightfiled | Microscopy. A) </span></b><span style="font-size: 10pt;" lang="EN-US">Brightfiled | ||
picture. <b>B)</b> Excitation | picture. <b>B)</b> Excitation | ||
Revision as of 00:20, 28 October 2010
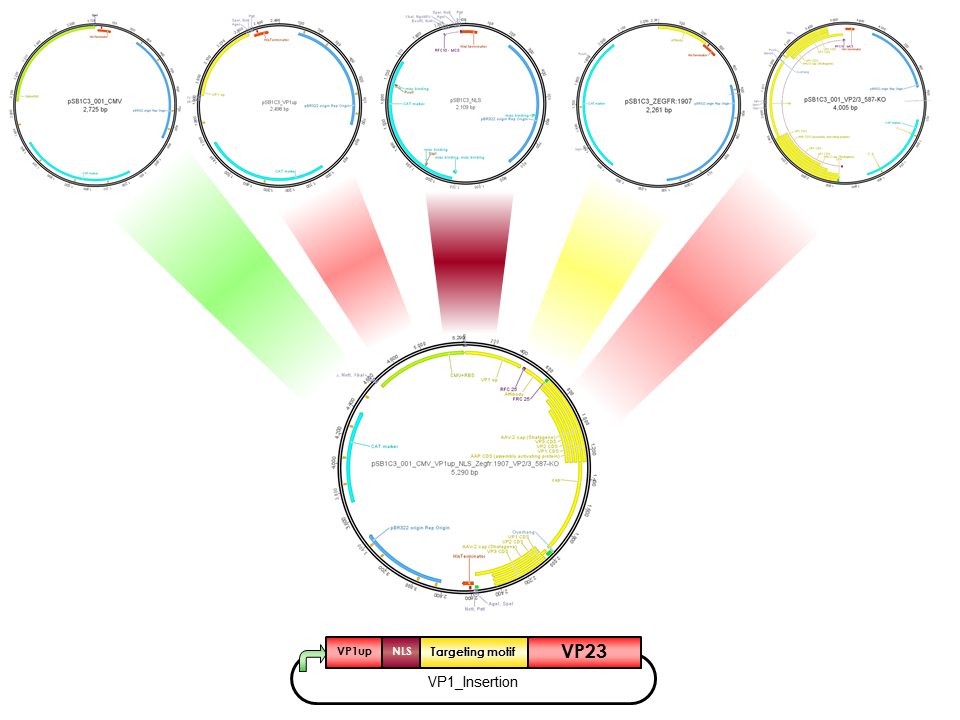
pCMV_VP1up_NLS_mVenus_[AAV2]-VP23 (ViralBrick-587KO-Empty)
| pCMV_VP1up_NLS_mVenus_(AAV2)-VP13(ViralBrick-587KO-Empty) | |
|---|---|
| BioBrick Nr. | BBa_K404165 |
| RFC standard | RFC 10 |
| Requirement | pSB1C3 |
| Source | |
| Submitted by | [http://2010.igem.org/Team:Freiburg_Bioware FreiGEM 2010] |
CMV
CMV promoter is derived from human Cytomegalovirus, which belongs to Herpesvirus group. All family members share the ability to remain in latent stage in the human body. CMV is located upstream of immediate-early gene. However, CMV promoter is an example of widely used promoters and is present in mammalian expression vectors. The advantage of CMV is the high-level constitutive expression in mostly all human tissues [Fitzsimons et al., 2002].
Capsid
(BBa_K404006)
The AAV capsid consists of 60 capsid protein subunits composed of the three cap proteins VP1, VP2, and VP3, which are encoded in an overlapping reading frame. Arranged in a stoichiometric ratio of 1:1:10, they form an icosahedral symmetry. The mRNA encoding for the cap proteins is transcribed from p40 and alternative spliced to minor and major products. Alternative splicing and translation initiation of VP2 at a nonconventional ACG initiation codon promote the expression of the VP proteins. VP1, VP2 and VP3 share a common C terminus and stop codon, but begin with a different start codon. The N termini of VP1 and VP2 play important roles in infection and contain motifs that are highly homologous to a phospholipase A2 (PLA2) domain and nuclear localization signals (NLSs). These elements are conserved in almost all parvoviruses. (Johnson et al., 2010a).
Whereas VP1 is translated from the minor spliced mRNA, while VP2 and VP3 are translated from the major spliced mRNA. The minor spliced product is approximately 10-fold less abundant than the major spliced mRNA. Thus, there is much less VP1 than VP2 and VP3 resulting in a capsid stoichiometric ratio of 1:1:10. The N terminus of VP1 has an extension of 65 amino acids including an additional extension of 138 N-terminal amino acids forming the unique portion of VP1. It contains a motif of about 70 amino acids that is highly homologous to phospholipase A2 (PLA2) domain. Furthermore, there are nuclear localization sequences (BR)(+), which are supposed to be necessary for endosomal escape and nuclear entry. (Bleker, Pawlita, & Kleinschmidt, 2006), (DiPrimio, Asokan, Govindasamy, Agbandje-McKenna, & Samulski, 2008), (Johnson et al., 2010a)
VP1up
VP1up protein is derived from the unique N-terminal region of VP1 protein. It contains a Phospholipase A2 motif which is essential for successful infection [Canaan et al., 2004; Zadori et al., 2001; Girod et al., 2002]
The Freiburg iGEM Team 2010 created this part in order integrate motifs, which are desired to be surface exposed, into the VP1 open reading frame. For this purpose VP1up needs to be fused to the N-terminus of the motif of interest, followed by coupling the resulting construct to VP2/3 (BBa_K404150, [AAV2]-VP23). For better infectivity a nuclear localization signal (Part:BBa_K404153, [AAV2]-NLS) (Grieger et al., 2007).
NLS
NLS are located in basic regions on the N terminus of VP2 (35 aa) and VP1 (172 aa) and mediate genome delivery into the nucleus and transduction [Hoque et al.,1999; Grieger et al., 2006]. Nuclear localisation sequence is hydrophilic and contains ß-turn and coil regions [Kalderon, et al, 1984]. It was also described in CPV and MVM viruses. Compared to CPV, MVM virus contains several NLS within the capsid, which are activated at different infection stages [Lombardo et al., 2000; Lombardo et al., 2002]
ViralBrick 587-KO empty
(BBa_K4004210)
The primary receptor of AAV-2 is the heparan sulfate proteoglycan (HSPG) receptor (Perabo et al. 2006). Its binding motif consists of five amino-acids located on the capsid surface: R484/R487, K532, R585/587. (Trepel et al. 2009). The positively charged arginine residues interact with the HSPGs' negatively charged acid residues. Opie et al. have shown that two point mutations (R585A and R588A) are sufficient to eliminate the heparin binding affinity in AAV2. (Opie et al. 2003). This ViralBrick has been created to introduce this knockout into other constructs. The biobricks with containing this knockout are annotated with „HSPG-ko“.
Characterization:
Nuclear Localization by Fluorescence Microscopy
AAV-293 cells were transfected with a 50:50 ratio of the Rep/Cap(VP1KO) ([AAV2]-Rep-VP23(ViralBrick-587KO-empty)_p5-TATAless, BBa_K404005) to this VP1-mVenus plasmid. We packaged mCherry, driven by the CMV promoter, into the virus capsids and followed protein expression via fluorescence microscopy. 30 hours post transfection mCherry fluorescence was detectable in the whole cytosol of the successfully transfected cells, demonstrating that DNA located between the AAV2 ITRs is already transcribed in the producer cell line. In contrast to that mVenus fluorescence signal could be observed only in the nuclei (Figure 1). This indicated that the nuclear localization sequence targets the single VP1_NLS_mVenus_VP2/3(587KO) proteins efficiently to the cell nucleus, where assembly and packaging of the virus particles takes place.

Figure 1: Fluorescence Microscopy. A) Brightfiled picture. B) Excitation at 555 nm showed mCherry signal in the cytosol. C) Excitation at 505 nm revealed mVenus fluorescence in the nuclei, indicating functionality of the nuclear localization sequence inserted – together with mVenus – into VP1. D) Merged image
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal XhoI site found at 698
Illegal XhoI site found at 884 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 665
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 3675
Illegal SapI site found at 2586
References
Johnson, J.S., 2010. Mutagenesis of adeno-associated virus type 2 capsid protein VP1 uncovers new roles for basic amino acids in trafficking and cell-specific transduction. Journal of virology, 84(17), 8888-902. doi: 10.1128/JVI.00687-10.
Lombardo, E, 2000. A beta-stranded motif drives capsid protein oligomers of the parvovirus minute virus of mice into the nucleus for viral assembly. Journal of Virology, 74(8), pp.3804-3814
Lombardo, E , 2002. Complementary roles of multiple nuclear targeting signals in the capsid proteins of the parvovirus minute virus of mice during assembly and onset of infection. Journal of Virology, 76(14), pp.7049-7059
Bleker, S., Pawlita, M., & Kleinschmidt, J., 2006. IImpact of capsid conformation and Rep-capsid interactions on adeno-associated virus type 2 genome packaging. Journal of virology, 80(2), 810-820. doi: 10.1128/JVI.80.2.810.
Canaan, 2004. Interfacial enzymology of parvovirus phospholipases A2. The Journal of Biological Chemistry, 279(15), pp.14502-14508.
DiPrimio, N., Asokan, A., Govindasamy, L., Agbandje-McKenna, M., & Samulski, R. J. , 2008. Surface loop dynamics in adeno-associated virus capsid assembly. Journal of virology, 82(11), 5178-89. doi: 10.1128/JVI.02721-07.
Girod, A., 2002. The VP1 capsid protein of adeno-associated virus type 2 is carrying a phospholipase A2 domain required for virus infectivity. The Journal of general virology, 83(Pt 5), pp.973-978. Available at: http://www.ncbi.nlm.nih.gov/pubmed/11961250.
Johnson, J.S., 2010. Mutagenesis of adeno-associated virus type 2 capsid protein VP1 uncovers new roles for basic amino acids in trafficking and cell-specific transduction. Journal of virology, 84(17), 8888-902. doi: 10.1128/JVI.00687-10.
Zadori, Z , 2001. A viral phospholipase A2 is required for parvovirus infectivity. Developmental Cell, 1(2), pp.291-302. Available at: http://www.ncbi.nlm.nih.gov/pubmed/11702787.
Grieger et al., (2007). Surface-exposed adeno-associated virus Vp1-NLS capsid fusion protein rescues infectivity of noninfectious wild-type Vp2/Vp3 and Vp3-only capsids but not that of fivefold pore mutant virions. J Virol. 2007 Aug;81(15):7833-43. Epub 2007 May 16.