Difference between revisions of "Part:BBa J23114"
Sanshikagarg (Talk | contribs) |
|||
| (6 intermediate revisions by 5 users not shown) | |||
| Line 1: | Line 1: | ||
| − | |||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_J23114 short</partinfo> | <partinfo>BBa_J23114 short</partinfo> | ||
| + | <!--[[Template:Berk2006iGEM_JCA_TCPromoters]]--> | ||
| + | {{Berk2006iGEM_JCA_TCPromoters}} | ||
| − | |||
| − | |||
===Usage and Biology=== | ===Usage and Biology=== | ||
| + | The Yale iGEM team has assembled this promoter to citrine (an improved version of YFP, with excitation peak at 514nm and emission peak at 527nm) and a T7-terminator to quantify the level of expression in E. coli and in non-model organism hosts. This construct is available as the biobrick [https://parts.igem.org/Part:BBa_K1856002 K1856002]. | ||
| + | This construct has been successfully cloned into E. coli using the broad-host range vector pKT230, a RSF1010 derived plasmid, as well as using the pPZP200 plasmid which can be transformed into agrobacterium and rhizobium. Leaky expression of citrine was observed. | ||
| + | |||
| + | ==Characterization== | ||
| + | [[Image:Expression_Level_for_anderson-citrine_(pKT230-Lic).jpg|300px|right]] | ||
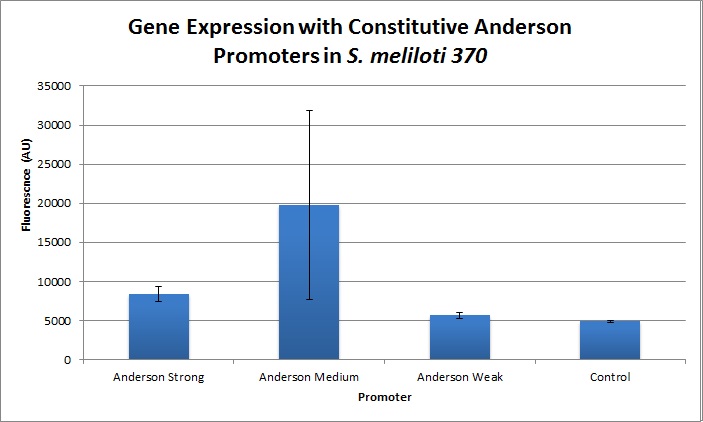
| + | [[Image:Expression_Level_for_constitutive_promoters_in_S_meliloti_(pPZP200-LIC).jpg|300px|right]] | ||
| + | ===Measured strength=== | ||
| + | The Yale team found fluorescence levels over fifteen times above the baseline level in E. coli cells transformed with the promoter-citrine construct. However, fluorescence levels were not significantly over the baseline when the construct was transformed into S. meliloti. | ||
| + | |||
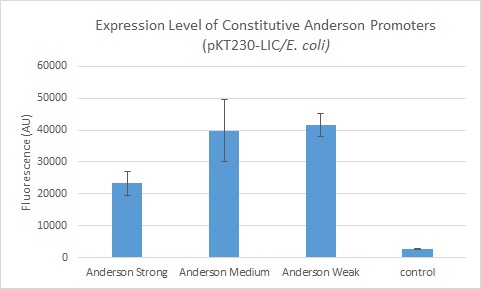
| + | We further compared expression level of this promoter to two of the other promoters in the Anderson collection. The Anderson lab found relatively expression levels of 1, 0.33, and 0.10 respectively when comparing the promoters [https://parts.igem.org/Part:BBa_J23100 J23100 (Anderson Strong)], [https://parts.igem.org/Part:BBa_J23110 J23110 (Anderson Medium)] and [https://parts.igem.org/Part:BBa_J23114 J23114 (Anderson Weak)]. We found the relative expressions to be 1, 1.71 and 1.78 when we transformed our constructs into E.coli: [https://parts.igem.org/Part:BBa_K1856004 K1856004 (Anderson Strong-Citrine)], [https://parts.igem.org/Part:BBa_K1856003 K1856003 (Anderson Medium-Citrine)], [https://parts.igem.org/Part:BBa_K1856002 K1856002 (Anderson Weak-Citrine)]. This highlights that even as all three Anderson promoters are effective in driving gene expression in E. coli, the relative expression level may vary depending on the gene fused to the promoter and on the E. coli strain used. The Yale iGEM team's readings were obtained in E. coli DH10B. | ||
| + | |||
| + | Comparing expression levels in S. meliloti (a rhizobium strain), the relative expression values were 1, 2.36, and no significant expression for the Anderson Strong, Anderson Medium, and Anderson Weak promoters respectively. | ||
<!-- --> | <!-- --> | ||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
| Line 17: | Line 28: | ||
<partinfo>BBa_J23114 parameters</partinfo> | <partinfo>BBa_J23114 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
| + | |||
| + | ==Added by KEYSTONE_A 2020 Team== | ||
| + | |||
| + | J23114 can be used as constitutive promoter in bacterial cellulose producing strain [[Part:BBa_K1321306|K. rhaeticus iGEM]], but it is of low strength. | ||
| + | |||
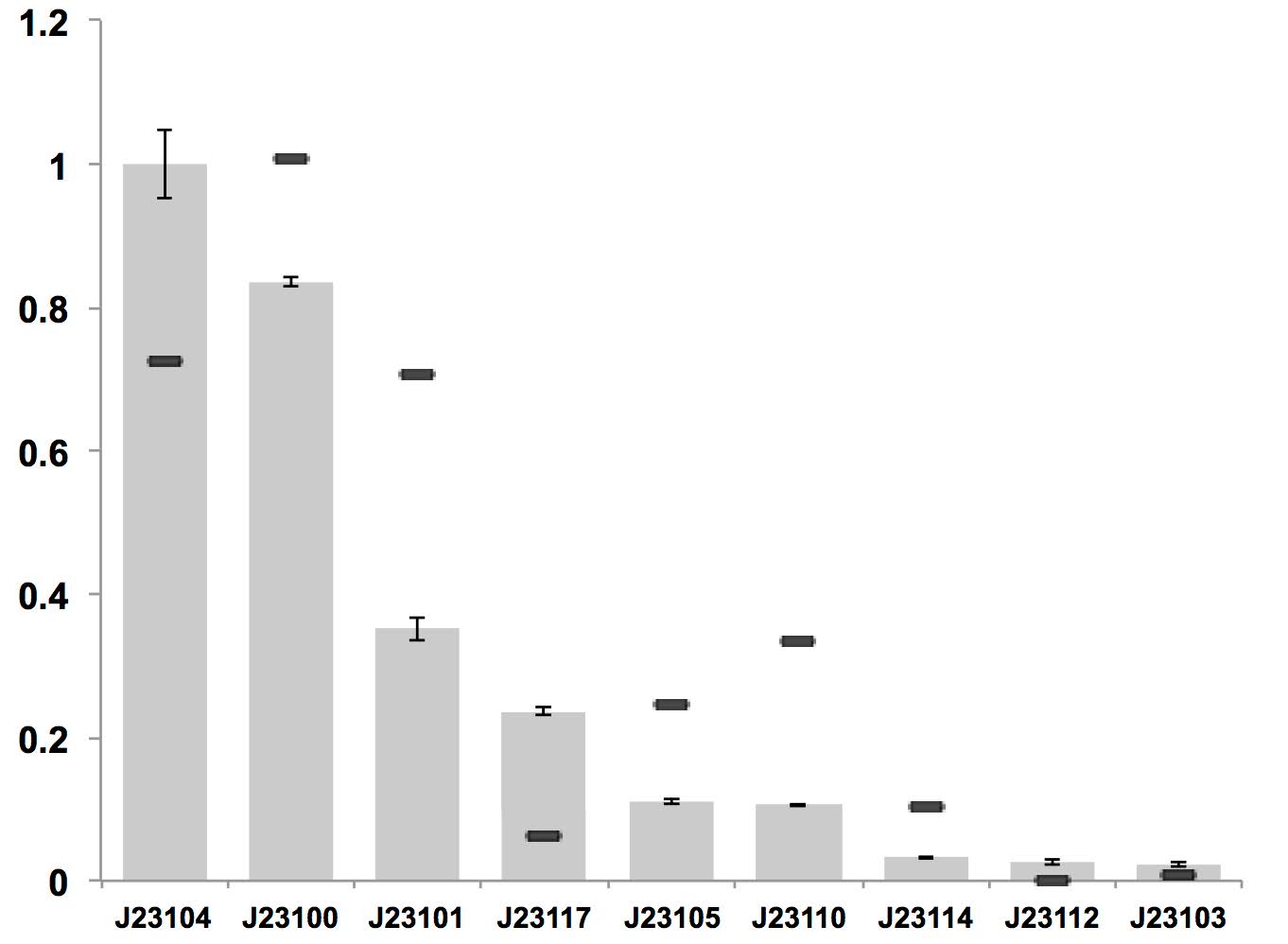
| + | [[File:KEYSTONE A constitutive promoter strength.jpg|600px|thumb|center|Visual Results as Normally Open Switches]] | ||
| + | |||
| + | '''Figure above: Constitutive promoter average strengths in K. rhaeticus iGEM and E. coli, normalized against J23104.''' | ||
| + | Although all promoters are functional, their relative strengths differ between K. rhaeticus and E. coli. For K. rhaeticus, data is shown as grey bars, with standard deviation of N=3 biological replicates, characterized in liquid HS-medium containing cellulase, measured 3 h post-inoculation. Relative promoter strengths in E. coli are superimposed as black stripes. | ||
| + | '''References:''' | ||
| + | |||
| + | Florea, M., Hagemann, H., Santosa, G., Abbott, J., Micklem, C. N., Spencer-Milnes, X., ... & Chughtai, H. (2016). Engineering control of bacterial cellulose production using a genetic toolkit and a new cellulose-producing strain. Proceedings of the National Academy of Sciences, 113(24), E3431-E3440. | ||
| + | |||
| + | {|width='100%' style='border:1px solid gray' | ||
| + | |- | ||
| + | |width='10%'| | ||
| + | <partinfo>BBa_J23100 AddReview 5</partinfo> | ||
| + | <I>University of Texas at Austin iGEM 2019</I> | ||
| + | |width='60%' valign='top'| | ||
| + | |||
| + | <h3>UT Austin iGEM 2019: Characterization of metabolic burden of the Anderson Series</h3> | ||
| + | |||
| + | <h4>Description</h4> | ||
| + | The 2019 UT Austin iGEM team transformed the Anderson Series promoters into our 'burden monitor' DH10B strain of E. coli, which contains a constitutive GFP cassette in the genome of the cell. GFP expression fluctuates depending on the number of ribosomes available. Using this strain, we characterized the relative burden (percent reduction in growth rate) of each Anderson Series part. Our results showed a range of growth rate reductions for each of these parts due to ribosomal reallocation from the genome of the host cell, towards the expression of RFP. Anderson Series parts with strong promoters are depicted with darker red colors and Anderson Series parts with weak promoters are depicted with lighter pink colors to show relative RFP expression. | ||
| + | We saw a positive correlation between relative promoter strength and metabolic burden; parts with stronger promoters expressed less GFP and had a lower growth rate than parts with weaker promoters. The regression line for the graph below was constructed by measuring the burden of 5 parts that were created by the 2019 UT Austin iGEM team that each contained an Anderson Series promoter (<partinfo>J23104</partinfo> or <partinfo>J23110</partinfo>), an RBS of varying strength, and a BFP reporter. For more information on characterization of these parts through the burden monitor, visit our team’s wiki page: [https://https://2019.igem.org/Team:Austin_UTexas] | ||
| + | |||
| + | <html> | ||
| + | <figure> | ||
| + | <div class = "left"> | ||
| + | <img src = "https://static.igem.org/mediawiki/parts/a/a0/AndersonCharacterization.jpg" style = "width:550px;height:500px"> | ||
| + | </div> | ||
| + | <figcaption><b>Fig.1:</b>Growth vs GFP Expression graph showing the relative burden positions of the Anderson Series promoters. The parts with strong promoters are depicted in dark red and are clustered near the bottom of the graph because they have lower growth rates and express lower levels of GFP as a result of high cellular burden. The parts with weaker promoter are depicted in light pink ad are clustered near the top of the graph because they have higher growth rates and express higher levels of GFP as a result of low cellular burden.</figcaption> | ||
| + | </figure> | ||
| + | </html> | ||
| + | <br><br> | ||
| + | <html> | ||
| + | <figure> | ||
| + | <div class = "left"> | ||
| + | <img src = "https://static.igem.org/mediawiki/parts/8/80/T--Austin_Utexas--andersontable.png" style = "width:545px;height:375px"> | ||
| + | </div> | ||
| + | <figcaption><b>Table.1:</b> Burden measurements for the Anderson Series promoters measured as percent reduction in growth rate ± 95% confidence interval. </figcaption> | ||
| + | </figure> | ||
| + | </html> | ||
| + | |||
| + | <h4>Importance of Characterizing Burden</h4> | ||
| + | <p> Although often we cannot avoid using a specific burdensome part, knowing in advance that it is burdensome, and that it has a high chance of mutating into a non-functional genetic device, can help with troubleshooting and coming up with alternatives. In the specific case of fluorescent protein-expressing devices, Fluorescence-activated cell sorting (FACS) can be used to filter out individual cells that meet a certain fluorescence threshold. This way, the cells expressing lower levels of the fluorescent protein are weeded out of the population.</p> | ||
| + | |||
| + | |||
| + | ==Characterization: Jilin_China 2019== | ||
| + | <h3>'''Group:'''</h3> Jilin_China 2019 | ||
| + | |||
| + | <h5> | ||
| + | <P style="text-indent:2em;"> | ||
| + | We hope that when we use an unrepresentated promoter, we can easily and accurately get its relative strength in the already characterized promoter family. | ||
| + | </P> | ||
| + | |||
| + | <P style="text-indent:2em;"> | ||
| + | Therefore, we constructed the promoter strength measurement vector BBa_K3078100 and characterized the promoter with a large intensity gradient of the Anderson promoter family, BBa_J23119, BBa_J23104, BBa_J23108, BBa_J23105, BBa_J23114, and it is expected to establish a standard curve with fluorescence intensity. | ||
| + | </p> | ||
| + | |||
| + | </h5> | ||
| + | |||
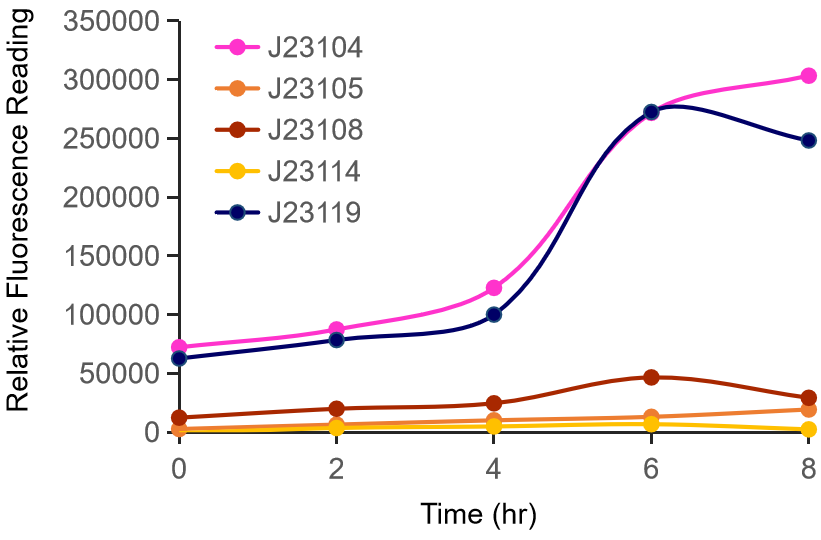
| + | [[File:表征1.png|600px|center|表征1]] | ||
| + | <center style="text-align:left;"> | ||
| + | Figure 1. The fluorescence of each promoter changes with time (fluorescence / OD600). Different elements are connected to the same measurement carrier, cultured overnight, diluted to OD = 0.02 the next day, and the fluorescence at absorption wavelength 528nm was measured under excitation wavelength 485nm light. | ||
| + | </center> | ||
| + | |||
| + | |||
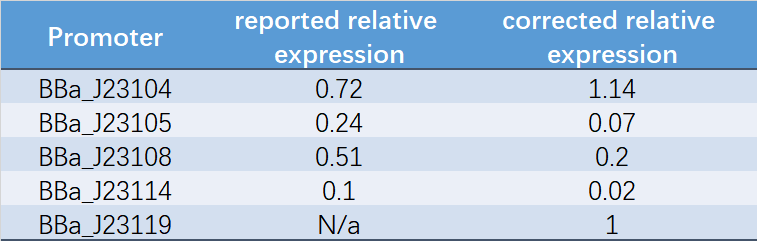
| + | <center style="text-align:left;"> | ||
| + | Table 1. Reported relative expression and corrected relative expression. We calculated the relative fluorescence intensity of each promoter for it based on J23119. | ||
| + | </center> | ||
| + | |||
| + | [[File:表征2.png|600px|center|表征2]] | ||
| + | |||
| + | <h5> | ||
| + | <P style="text-indent:2em;"> | ||
| + | The fluorescence data showed the intensity of BBa_J23104 and BBa_J23119 is much too high to constitute a linear relationship with the other three promoters. The reason of which might be the fusion genes influenced the relatve expression level of the promoter. | ||
| + | </P> | ||
| + | |||
| + | <P style="text-indent:2em;"> | ||
| + | We invited JiangnanU_China to repeat, and the result of its characterization was not consistent with reported relative expression .Therefore, we speculated that the relative strength might be affected by experimental strains, plasmids and other factors. Under our experimental conditions, the relative strength of Anderson promoter was not consistent with reported relative expression. | ||
| + | </p> | ||
| + | |||
| + | </h5> | ||
| + | </center> | ||
| + | ==NYU Abu Dhabi 2024 Characterization== | ||
| + | <h3>'''Group:'''</h3> NYU-Abu-Dhabi 2024 <h5> <P style="text-indent:2em;"> BBa_J23114 was tested as a promoter driving eYFP expression in bacterial colonies under two temperature conditions: 28°C (ambient) and 37°C (heat stress). According to the promoter activity graph, BBa_J23114 exhibited distinct trends at each temperature. The promoter activity was measured using fluorescence quantification. At 28°C, the promoter showed moderate activity that remained fairly consistent over time, without significant fluctuations. In contrast, at 37°C, the activity started higher than at 28°C but then gradually decreased over time. These trends suggest that BBa_J23114 may perform better initially at higher temperatures, but it demonstrates more stable performance under ambient conditions (28°C) in the long term. </P> </h5> src="https://static.igem.wiki/teams/5125/whatsapp-image-2024-10-01-at-11-00-13-am.jpeg" class= "center" width="50px"></html> | ||
| + | |||
| + | <b><i>Figure 1. Promoter activity of BBa_J23114 at 28°C and 37°C. </center> | ||
Latest revision as of 06:50, 2 October 2024
constitutive promoter family member
Variant RFP (au) J23112 1 J23103 17 J23113 21 J23109 106 J23117 162 J23114 256 J23115 387 J23116 396 J23105 623 J23110 844 J23107 908 J23106 1185 J23108 1303 J23118 1429 J23111 1487 J23101 1791 J23104 1831 J23102 2179 J23100 2547 |
Constitutive promoter family
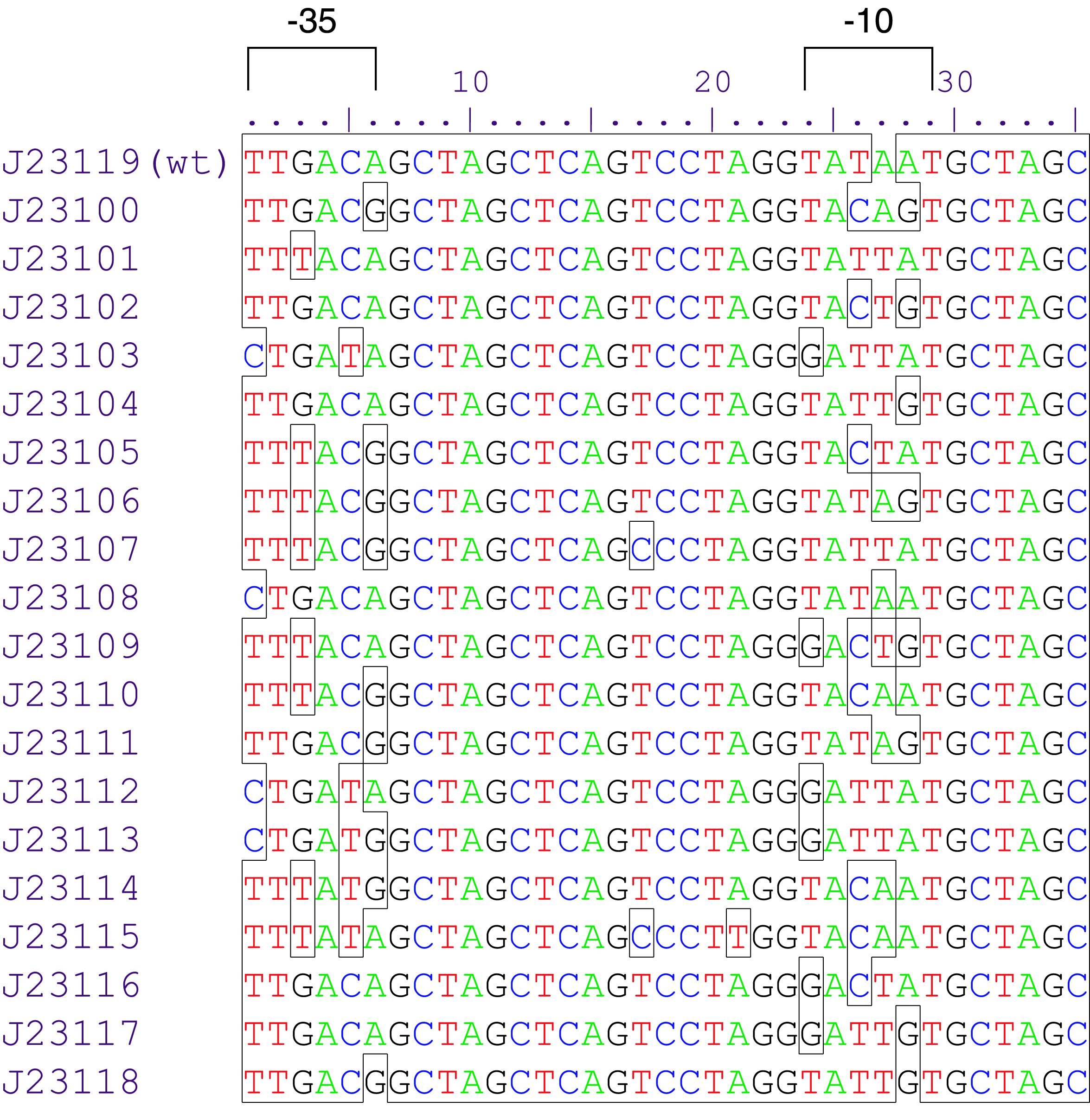
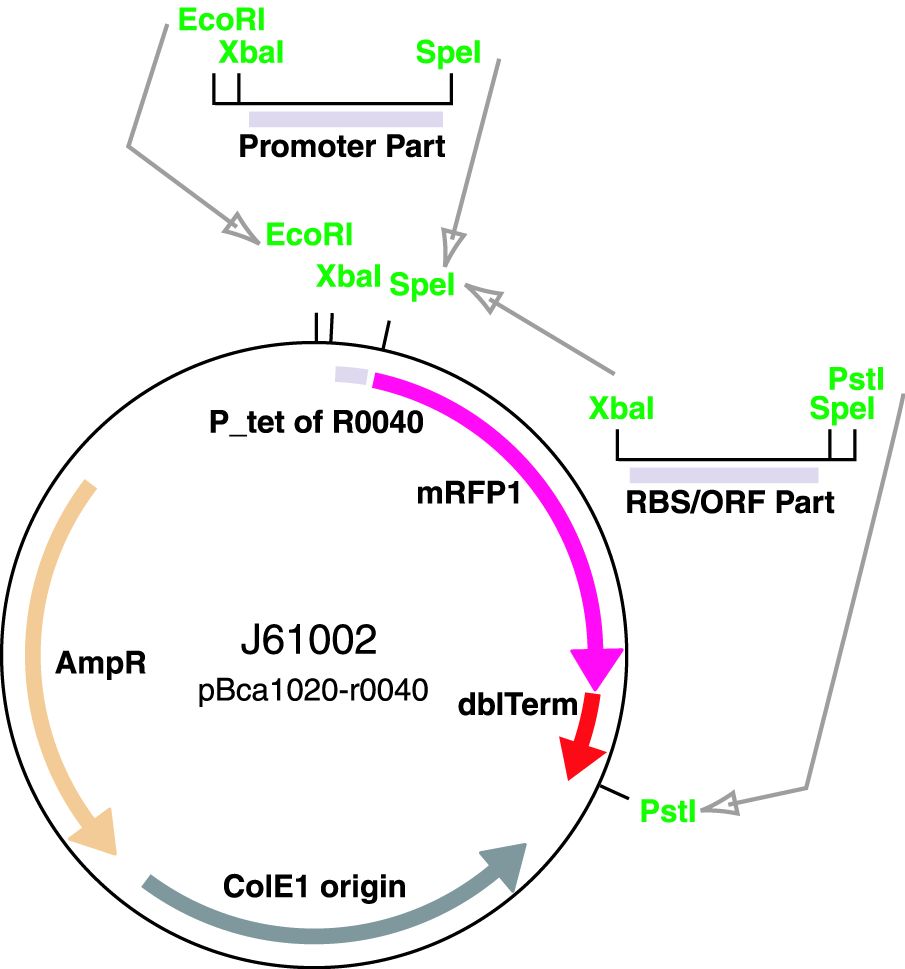
Parts J23100 through J23119 are a family of constitutive promoter parts isolated from a small combinatorial library. J23119 is the "consensus" promoter sequence and the strongest member of the family. All parts except J23119 are present in plasmid J61002. Part J23119 is present in pSB1A2. This places the RFP downstream of the promoter. Reported activities of the promoters are given as the relative fluorescence of these plasmids in strain TG1 grown in LB media to saturation. See part BBa_J61002 for details on their use.
These promoter parts can be used to tune the expression level of constitutively expressed parts. The NheI and AvrII restriction sites present within these promoter parts make them a scaffold for further modification. JCAraw
Usage and Biology
The Yale iGEM team has assembled this promoter to citrine (an improved version of YFP, with excitation peak at 514nm and emission peak at 527nm) and a T7-terminator to quantify the level of expression in E. coli and in non-model organism hosts. This construct is available as the biobrick K1856002.
This construct has been successfully cloned into E. coli using the broad-host range vector pKT230, a RSF1010 derived plasmid, as well as using the pPZP200 plasmid which can be transformed into agrobacterium and rhizobium. Leaky expression of citrine was observed.
Characterization
Measured strength
The Yale team found fluorescence levels over fifteen times above the baseline level in E. coli cells transformed with the promoter-citrine construct. However, fluorescence levels were not significantly over the baseline when the construct was transformed into S. meliloti.
We further compared expression level of this promoter to two of the other promoters in the Anderson collection. The Anderson lab found relatively expression levels of 1, 0.33, and 0.10 respectively when comparing the promoters J23100 (Anderson Strong), J23110 (Anderson Medium) and J23114 (Anderson Weak). We found the relative expressions to be 1, 1.71 and 1.78 when we transformed our constructs into E.coli: K1856004 (Anderson Strong-Citrine), K1856003 (Anderson Medium-Citrine), K1856002 (Anderson Weak-Citrine). This highlights that even as all three Anderson promoters are effective in driving gene expression in E. coli, the relative expression level may vary depending on the gene fused to the promoter and on the E. coli strain used. The Yale iGEM team's readings were obtained in E. coli DH10B.
Comparing expression levels in S. meliloti (a rhizobium strain), the relative expression values were 1, 2.36, and no significant expression for the Anderson Strong, Anderson Medium, and Anderson Weak promoters respectively. Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Added by KEYSTONE_A 2020 Team
J23114 can be used as constitutive promoter in bacterial cellulose producing strain K. rhaeticus iGEM, but it is of low strength.
Figure above: Constitutive promoter average strengths in K. rhaeticus iGEM and E. coli, normalized against J23104. Although all promoters are functional, their relative strengths differ between K. rhaeticus and E. coli. For K. rhaeticus, data is shown as grey bars, with standard deviation of N=3 biological replicates, characterized in liquid HS-medium containing cellulase, measured 3 h post-inoculation. Relative promoter strengths in E. coli are superimposed as black stripes. References:
Florea, M., Hagemann, H., Santosa, G., Abbott, J., Micklem, C. N., Spencer-Milnes, X., ... & Chughtai, H. (2016). Engineering control of bacterial cellulose production using a genetic toolkit and a new cellulose-producing strain. Proceedings of the National Academy of Sciences, 113(24), E3431-E3440.
|
•••••
University of Texas at Austin iGEM 2019 |
UT Austin iGEM 2019: Characterization of metabolic burden of the Anderson SeriesDescriptionThe 2019 UT Austin iGEM team transformed the Anderson Series promoters into our 'burden monitor' DH10B strain of E. coli, which contains a constitutive GFP cassette in the genome of the cell. GFP expression fluctuates depending on the number of ribosomes available. Using this strain, we characterized the relative burden (percent reduction in growth rate) of each Anderson Series part. Our results showed a range of growth rate reductions for each of these parts due to ribosomal reallocation from the genome of the host cell, towards the expression of RFP. Anderson Series parts with strong promoters are depicted with darker red colors and Anderson Series parts with weak promoters are depicted with lighter pink colors to show relative RFP expression. We saw a positive correlation between relative promoter strength and metabolic burden; parts with stronger promoters expressed less GFP and had a lower growth rate than parts with weaker promoters. The regression line for the graph below was constructed by measuring the burden of 5 parts that were created by the 2019 UT Austin iGEM team that each contained an Anderson Series promoter (BBa_J23104 or BBa_J23110), an RBS of varying strength, and a BFP reporter. For more information on characterization of these parts through the burden monitor, visit our team’s wiki page: [1]
Importance of Characterizing BurdenAlthough often we cannot avoid using a specific burdensome part, knowing in advance that it is burdensome, and that it has a high chance of mutating into a non-functional genetic device, can help with troubleshooting and coming up with alternatives. In the specific case of fluorescent protein-expressing devices, Fluorescence-activated cell sorting (FACS) can be used to filter out individual cells that meet a certain fluorescence threshold. This way, the cells expressing lower levels of the fluorescent protein are weeded out of the population.
Characterization: Jilin_China 2019Group:Jilin_China 2019
|



 " class= "center" width="50px"></html>
" class= "center" width="50px"></html>