Difference between revisions of "Part:BBa J06504"
(→Improvement by UCopenhagen 2022) |
(→Improvement by UCopenhagen 2022) |
||
| Line 45: | Line 45: | ||
From a plasmid expressing just the mCherry, we used two sets of primers (seq primers and USER primers) to amplify the mCherry fragment as a control for mCherry. However, the seq primers also amplified some base pairs that were upstream and downstream and hence, one of the control fragments - mCherry Control (seq primers) - seems slightly higher than the other mCherry control fragment. | From a plasmid expressing just the mCherry, we used two sets of primers (seq primers and USER primers) to amplify the mCherry fragment as a control for mCherry. However, the seq primers also amplified some base pairs that were upstream and downstream and hence, one of the control fragments - mCherry Control (seq primers) - seems slightly higher than the other mCherry control fragment. | ||
| − | [[File:mccfig2.jpg. | + | [[File:mccfig2.jpg.]] |
'''Conclusion - '''Hence, it is clear that our USER cloning was successful in adding the SnoopCatcher to the mCherry fragment as seen in the PCR. | '''Conclusion - '''Hence, it is clear that our USER cloning was successful in adding the SnoopCatcher to the mCherry fragment as seen in the PCR. | ||
Revision as of 18:52, 11 October 2022
monomeric RFP optimized for bacteria
mRFP1-derived, altered to be a BioBrick by removing a PstI site and adding BioBrick ends. [mRFP1 was itself a derived from DsRed (via 33 mutations!)]
mCherry is one of several "second-generation" monomeric fluorescent proteins developed in Roger Tsien's laboratory at UCSD (cf., Nature Biotechnology 22, 1567 - 1572 (2004). PMID 15558047
Usage and Biology
Some strange experimental results that have been seen could be explained by an internal RBS + start. The 10th amino acid is a Met which is preceded by AGGAGGA(NNNN). This is almost a perfect consensus RBS so it seems quite likely that translation can begin 10 amino acids in. Note that mCherry was designed by fusing the N and C terminal regions of EGFP on to a mRFP variant (to increase tolerance to protein fusions). Thus, removing the first several amino acids is not expected to have much effect on fluorescence. If this is truly a strong internal RBS, then the identity of any attached RBS may have little effect. Also, one should be careful when making protein fusions. --Austin
The copy as provided in the 2010 distribution is incorrect - it contains ~500 bp of something that is not mCherry between the VF2 and VR sites. You can get a functioning copy via PCR out of Part:BBa_J06702. --[http://openwetware.org/wiki/User:Joseph_T._Meyerowitz jmeyerow]
Improvement by SYSU-CHINA 2016
This part has been improved to be the mCHERRY UNIT including a degradation tag named DBOX, a fluorescent gene mCherry and an sv40 terminator flanked by homodromous recognition site of recombinase VIKA named Vox. The whole part can be cut off by VIKA. For more information of the improved part, please go to the page of Part:BBa_K1926013.
Improvement by USP_UNIFESP-Brazil
The reporter mCherry has been codon-optimized for Chlamydomonas reinhardtii and it was successfully tested as a reporter in this chassis. We have made several measurements to show its fluorescence in Chlamydomonas reinhardtii as well its properties (Fluorescence excitation/emission spectrum). For more information of the improved part, please go to the page of Part:BBa_K2136016.
Improvement by Evry_Paris-Saclay 2017
The reporter mCherry has been codon-optimized for Escherichia coli K12 (Part:BBa_K2448004) and it was successfully tested as a reporter in this chassis. We have used this part to build the Universal Biosensing Chassis (BBa_K2448023 and BBa_K2448024) and test our Psicose Biosensors (BBa_K2448025, BBa_K2448026, BBa_K2448027, BBa_K2448028, BBa_K2448029, BBa_K2448030 and BBa_K2448031) as well as our Fructose Biosensor (BBa_K2448032). For more information on the improved part, please go to the page of Part:BBa_K2448004.
Improvement by University of Minnesota 2017
The internal RBS-like sequence in this mCherry reporter was removed, and the overall sequence was codon-optimized for E. coli K12. This new part BBa_K2375001 should be more suitable for creating fusion proteins.
Improvement by TAU_Israel 2021
The codon sequence has been optimized for expression in Bacillus subtilis against E. coli using the Communique tool using several methods. For more information, refer to part pages BBa_K3871001, BBa_K3871002, BBa_K3871003, BBa_K3871004,BBa_K3871005 and the 2021 TAU_Israel wiki, available here.
Improvement by UCopenhagen 2022
We improved this mCherry protein by adding SnoopCatcher (part BBa_K4247009), so that it can now spontaneously form an irreversible isopeptide bond with any protein that contains the SnoopTag (part BBa_K4247008).
Addition of SnoopCatcher to mCherry
Aim - To attach the SnoopCatcher sequence (Part BBa_K4247009) to the original mCherry part (Part BBa_J06504).
Results - We designed 6 primers with Uracil to perform a USER cloning reaction. Two primers to amplify the mCherry fragment while swapping the 6-His tag’s location from the C-terminus to the N-terminus, two for amplifying the backbone of the expression vector (pBAD) and two for amplifying the SnoopCatcher fragment from a part we already produced (BBa_K4247021 - Mfp151_SnoopCatcher). We verified the success of the cloning via PCR and sequencing.
From the final USER cloned plasmid coding for the mCherry_SnoopCtacher, we amplified mCherry, SnoopCatcher and the whole mCherry_SnoopCatcher fragment via PCR to confirm that our USER cloning worked. We also used mCherry and SnoopCatcher fragments as controls.
From a plasmid expressing just the mCherry, we used two sets of primers (seq primers and USER primers) to amplify the mCherry fragment as a control for mCherry. However, the seq primers also amplified some base pairs that were upstream and downstream and hence, one of the control fragments - mCherry Control (seq primers) - seems slightly higher than the other mCherry control fragment.
Conclusion - Hence, it is clear that our USER cloning was successful in adding the SnoopCatcher to the mCherry fragment as seen in the PCR.
Protein expression of mCherry_SnoopCatcher
Aim - To express and purify mCherry_SnoopCatcher.
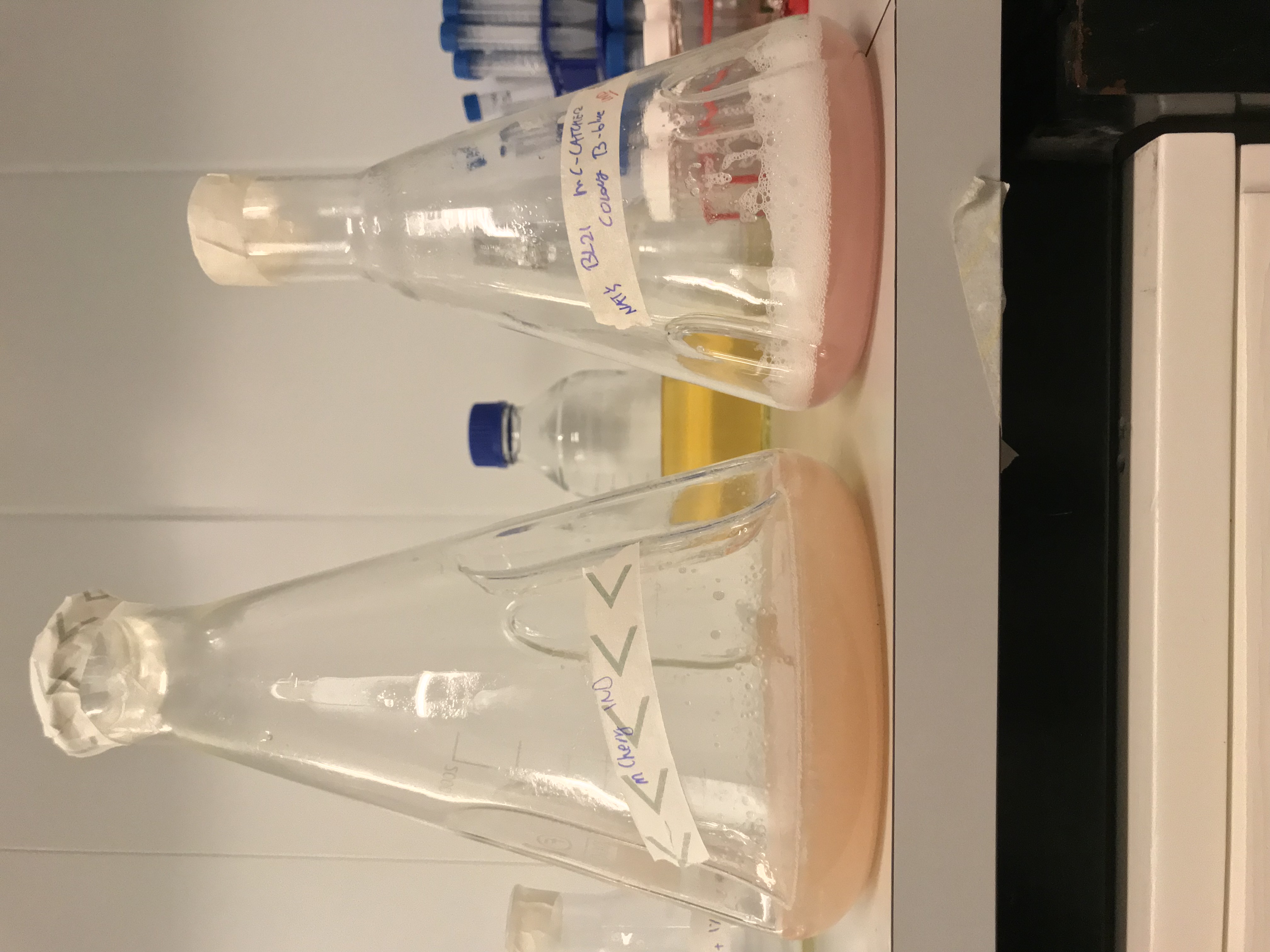
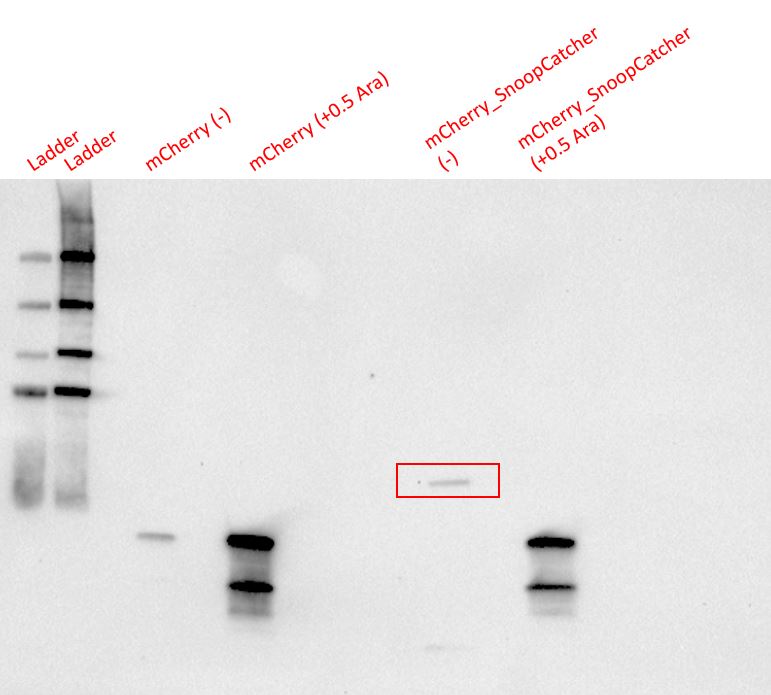
Results - We expressed the mCherry_SnoopCatcher and mCherry in E.coli BL21(DE3) and induced protein expression using 0.5% arabinose overnight. The next day, we obtained a bright red E.coli culture for both proteins. Then, we performed SDS-PAGE and a Western blot to check for protein expression. It was clear that both proteins were expressed since we found bands of appropriate size only in the induced lanes.
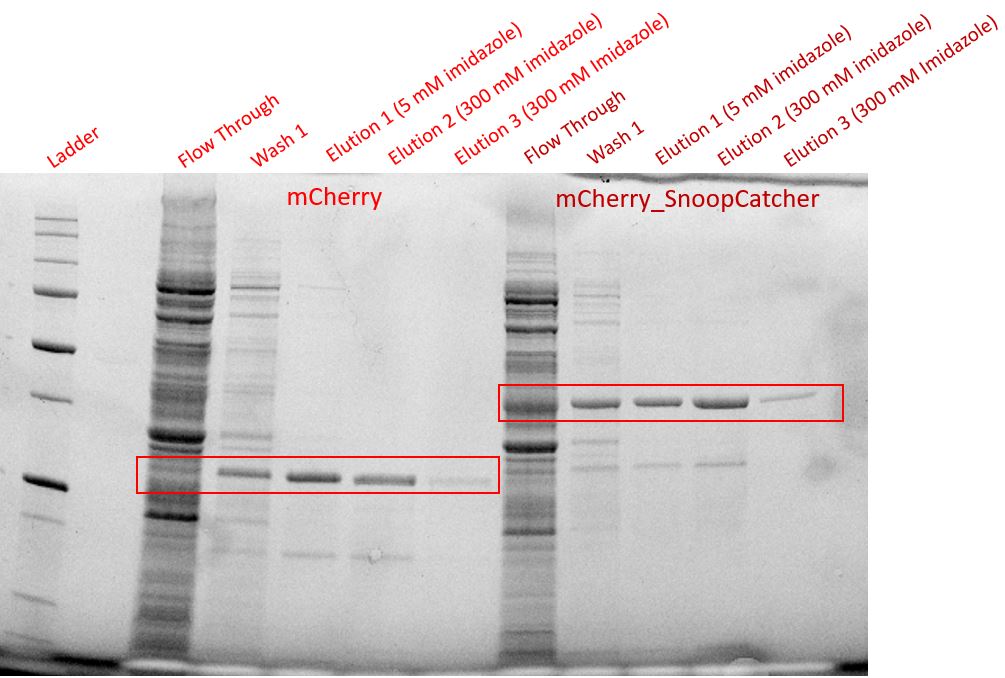
We then performed IMAC using Ni-NTA resin since the protein was expressed with a 6x His-tag and successfully purified mCherry_SnoopCatcher as well as mCherry. Then, we performed SDS-PAGE and a Western blot on the purification fractions of both proteins which confirmed that both mCherry_SnoopCatcher (40.89 KDa) and mCherry (26.7 KDa) were expressed since we found bands at the expected molecular weights. Further, we noticed that mCherry has a leaky expression (see mCherry control in the western blot) and that mCherry_Catcher does not stain as well as mCherry. The purification was not as efficient since an important portion of the protein was lost in the washes, probably due to saturation of the beads. However, we only needed small amounts of proteins for our assays.
Conclusion - Thus, mCherry and mCherry_SnoopCatcher were successfully expressed and purified.
Effect of the SnoopCtacher on mCherry’s fluorescence
Aim - To ensure the addition of the SnoopCatcher would not impair the main characteristic of mCherry, its fluorescence.
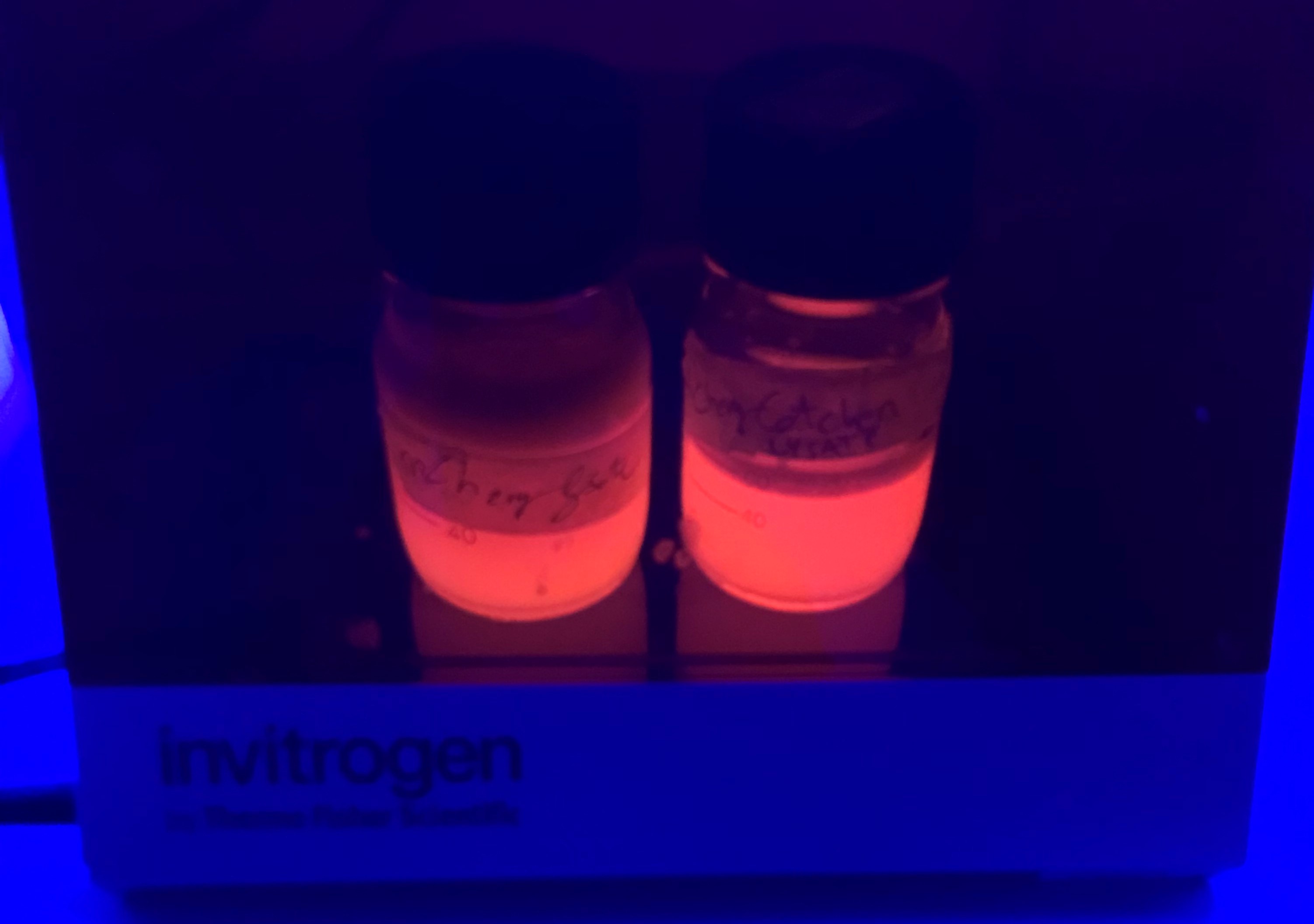
Results - We noticed that mCherry_SnoopCatcher also emits fluorescence in the red wavelengths. It was clear from the lysates that mCherry_SnoopCatcher fluoresces just as much, if not more than the original mCherry, considering that the lysate of mCherry comes from a 0.5L culture while the lysate of mCherry-SnoopCatcher comes from a 150mL culture. This leads us to believe that the addition of SnoopCatcher might improve the protein expression of mCherry or might even improve the fluorescence property of mCherry. However, these results need to be validated since we could not quantify and compare the protein yields or the fluorescence intensity due to time constraints.
Conclusion - Hence, the addition of SnoopCatcher does not affect the properties of mCherry.
Validation of SnoopCatcher’s function
Aim - To demonstrate the spontaneous isopeptide bond formation between our minispidroin protein displaying the SnoopTag and mCherry_SnoopCatcher.
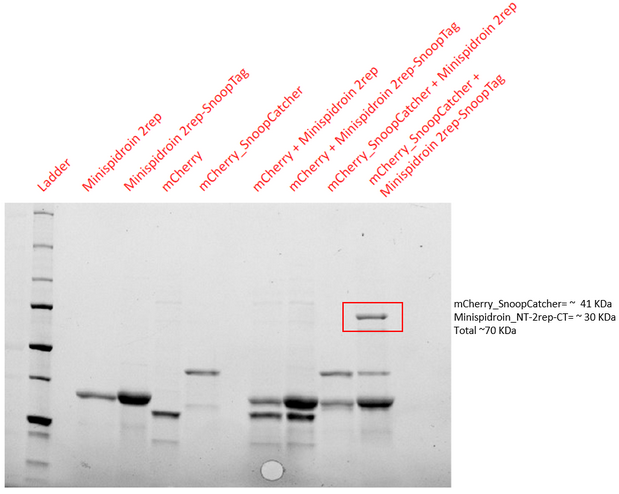
Results - The proteins were allowed to interact with each other by mixing equal amounts of both proteins. The reaction was carried out at 25°C for 40 minutes and the pH of the reaction was maintained at 8 using pH adjusted PBS. After 40 minutes, the reaction was stopped by adding SDS sample buffer. Then, we performed an SDS-PAGE and a Western blot to visualize the results. We would expect to see a band whose molecular weight is the sum of the molecular weights of both proteins to indicate that both proteins are bound to each via an isopeptide bond between the SnoopTag and the SnoopCatcher. We see such a band only in the reaction where mCherry_SnoopCatcher and Minispidroin 2rep with a SnoopTag were allowed to interact. The fact that we don't see this band in any of the controls confirms that the proteins were able to bind to each other solely due to the SnoopTag-Catcher system.
Conclusion - The SnoopCatcher enabled the mCherry_SnoopCatcher to bind to Minispidroin 2rep with a SnoopTag and hence, the function of the SnoopTag-Catcher system was validated.
By adding a functional SnoopCatcher to mCherry and validating the SnoopTag-Catcher system, we have successfully improved the part BBa_J06504. Thus, mCherry with the SnoopCatcher can bind to essentially any protein with a SnoopTag. Since mCherry is a widely used protein, we believe our improvement opens up many new uses of the protein among its repertoire of diverse biotechnological applications. For more information about our improved part, go to BBa_K4247025
Contribution: TU_Eindhoven 2019
In this contribution, we futher characterized mCherry (BBa_J06504). mCherry was cloned into a pET28a(+) vector (containing an N-terminal 6xHis-tag) and subsequently expressed in BL21 (DE3) E. coli and purified using immobilized metal affinity chromatography (IMAC). Different concentrations of mCherry were made and an increase in color intensity was visible (Figure 1).
Figure 1. Purified mCherry in different concentrations, showing a clear increase in color intensity.
Consequently, the purified protein was analyzed on an SDS-PAGE (Figure 2) which shows a clear blob above 25 kDa, corresponding with 6xHis-tagged mCherry’s molecular weight of 29.3 kDa. The blob in between 10 kDa and 15 kDa is a truncated version of mCherry as described by Tripathi [1]. There are other bands visible so the purity is not optimal. However, the contrast between the huge blobs and the other bands is very big.
Figure 2. SDS-PAGE of mCherry after purification.
Following protein purification, the protein functionality was analyzed. Firstly, the excitation and emission of mCherry were measured within its absorption and emission range (Figure 3). This shows a clear excitation maximum at 587 nm and an emission maximum at 610 nm. Secondly, the photostability of mCherry was measured following UV radiation up until thirty minutes to advance usage of mCherry in experiments (Figure 4). Exposure to UV light shows a decrease in fluorescence intensity. After 30 min, the intensity is almost halved compared to no UV exposure.
Figure 3. Excitation and emission spectrum of mCherry.AAAAAAAAAAFigure 4. Excitation spectrum of mCherry after different periods of UV radiation.
HUBU-WUHAN 2019-Characterization
We characterized the bleaching effect in zymomonas mobilis using fluorescent microscopy Leica DMi8. We obtained image information through fluorescence microscope and analyzed it with the Rleative Flourescense Calculator in LAS X.
It is expressed in pEZ-15A plasmid with a PlacUV5 promoter. LAS Z was used to analyze the images and calculate the rate of photobleaching. We obtained a half-life period as t1/2=40.07s
Characterization: SMMU-China 2019
In our project this year, we aim to develop a biomedical tattoo that could detect disease related molecules and output signals through accumulation of fluorescent proteins. We have also developed a software to analyze the intensity of the fluorescence so that this signal could reflect the disease conditions. To test whether this software could accurately measure the fluorescence and whether florescent proteins are sensitive enough to meet our conception, we carried out the following experiments: First, we cloned this part (BBa_J06504) into eukaryotic expression vector pcDNA3.1 and transfected it into HEK293T cells through Lipofectamine 2000 transfection reagent. Twenty-four and 48 hours after transfection, bright red light could be observed under florescence microscopy (Figure. 1)
The transfected and untransfected HEK293T cells were digested at 24h by trypsin and centrifugated in microtubes. The cells in microtubes were used to mimic the biomedical tattoo and were further examined. Whereas the cells did not show apparent difference with negative control under white light (Figure. 2a), they emitted red light when exposed to exciting light from a mercury lamp (Figure. 2b). This result suggested that fluorescent proteins were sensitive reporters and could be distinguished even by naked eyes under exciting light.
To demonstrate that the software was able to accurately measure fluorescent intensity of tattoo cells, we diluted mCherry-293 cells to a panel of concentrations by mixing with blank cells (Figure. 3), and made a standard curve (Figure. 4). The percentage of mCherry-293 cells in diluted cells were 20%, 40%, 60%, 80%, and 100%, respectively. The standard samples were taken photos by a camera and their fluorescence intensity was measured by using the software to analyze the photos. The standard curve was shown in Figure. 4.
We also made two samples whose concentrations were 45% and 70%, respectively. Their fluorescence intensity measured by the software were 41.810 and 60.634. Concentrations were calculated by putting the numbers into the standard curve and the results acquired were 44.5% and 69.8%, which were very close to the true value 45% and 70%.
These results demonstrated fluorescent proteins such as mCherry were sensitive and could be measured quantitively by our software.
References
[1] R. Tripathi, “Functional characterisation of LEA proteins from bdelloid rotifers,” 2012.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Contribution: iGEM Bielefeld-CeBiTec 2019
Additionally, we compared two different purification methods and characterized light-stability and pH sensitivity of mCherry.
Usage and Biology
Because of the wide range of applications for fluorescing proteins there was a great interest in finding and engineering improved variants and a wider colour spectrum. In the last few years red fluorescing proteins became more and more important. Common native red fluorescing proteins are often dimeric or tetrameric what makes their usage in experimental setups difficult. Directed mutation of dsRFP from the corallimorpharia Discosoma sp. Led to the first monomeric red fluorescing protein mRFP1 (Shaner et al. 2004). Unfortunately this mutations resulted in a lower quantum yield and decreased photostability (Shaner et al. 2004). During further protein engineering attempts, scientists were able to create the red fluorescent protein mCherry. mCherry is a 26.7 kDa protein that shows a very short maturation time of about 15 minutes and a low acid sensitivity. Its excitation maximum lies at 587 nm and it has its emission maxiumum at 610 nm (www.fpbase.org). In 2006 the crystal structure of mCherry was published (Shu and Remington 2006).

Expression
To further characterize purified mCherry we compared two different purification protocols. Purification via his-tag was compared to the IMPACT-purification protocol from NEB.
Expression cultures of mCherry in pTXB1 and mCherryHis in pSB1C3 in E. coli ER2566 were cultivated to an OD of around 0.6 at 37 °C in LB with 100 mg Ampicillin per L. Protein expression was induced by addition of IPTG to a final concentration of 0.4 mM. After additional 30 min at 37 °C cultures were transferred to 17 °C and protein expression was performed over night.

Expression cultures of mCherry and mCherryHis in E. coli ER2566 were harvested by centrifugation at 4 °C for 20 min and 4 000 rpm.
Purification
After cultivation and cell lysis via Ribolyzer the protein was purified using the His-purification kit from Macherey Nagel and the IMPACT-purification kit from NEB (Fig. 4).
Harvested cells were lysed using Zirconia metal beads (1 mm) in a Ribolyzer at 8 000 rpm for 15 s. The lysate was cleared by centrifugation at 4 °C for 1 h and 4 500 rpm. Cleared lysate was loaded onto a chitin column (IMPACT-purification) or a Ni-TED column (purification via his tag) and washed with wash buffer. Finally the protein was eluted, washed in PBS and concentrated.

E. coli lysate of the expression culture, flow-through- and wash-fraction as well as the purified protein were denatured by heating the samples to 98 °C for 10 min in SDS-PAGE loading buffer containing DTT and loaded on an polyacrylamide-gel (12 %). The proteins were separated through electrophoresis (25 mA). Suggested mCherry bands in the lane with purified proteins were marked in dark red.
In the last lane you can see that we were able to purify mCherry as well as mCherryHis. While the IMPACT-purification resulted in a higher yield, the purity of mCherryHis was higher as the protein lane in Fig. 5 indicated.
Following the SDS-PAGE we analyzed the purified protein via MALDI-ToF. For this purpose we excised the marked bands (Fig. 5) from the SDS-PAGE and started a tryptic digestion of the washed gel fragment. Analysis via MALDI-ToF confirmed that we were able to purify mCherry (Fig. 6).

Excised bands from the SDS-PAGE of mCherry and mCherryHis were washed, digested over night with trypsine and co-cristallyzed with a HCCA-matrix on a MALDI target. Mass spectrum was recorded in a MALDI-ToF MS from Bruker Daltronics and data was evaluated using the software BioTools.
Characterization
To gain some more knowledge about mCherry we analyzed different properties of the protein. First of all we measured its emission- and excitation spectra (Fig. 7).
Emission- (dashed lines) and excitation-spectra (solid lines) of mCherry purified via IMPACT-Kit (dark purple) and His-tag (pink) were measured (λEx=570 nm, λEm=600 nm to 850 nm) using the TECAN infinite M200 and normalized to their maximum.
Next we compared the fluorescence intensity of the two different mCherry-variants normalized to Texas Red (Fig. 8).

Fluorescence intensity of a dilution series of mCherry purified via IMPACT-Kit (dark purple) and mCherryHis (pink) was measured (λEx=570 nm, λEm=610 nm) using the TECAN infinite M200 and normalized to the fluorescence intensity of 0.5 µM Texas Red at the same wavelength.
Additionally we wanted to characterize the light tolerance and the pH-range of mCherry to get insight into the optimal handling procedures. To determine the stability of mCherry it was exposed to normal daylight in the lab at room temperature for a longer period of time (Fig. 9).

mCherry was exposed to normal daylight at room temperature. The remaining fluorescence intensity was measured at determinated time points (λEx=570 nm, λEm=610 nm, gain calculated from 2.5 µM Texas Red) using the TECAN infinite M200 and normalized to the intensity at t=0.
An often stated advantage of mCherry is its low acid sensitivity. To analyze the pH-range of mCherry we measured the remaining fluorescence intensity after incubation in buffers with different pH (Fig. 10).

mCherry was incubated for 5 minutes in buffers with different pH. The remaining fluorescence intensity was measured (λEx=570 nm, λEm=610 nm, gain calculated from 2.5 µM Texas Red) using the TECAN infinite M200 and normalized to the intensity at pH 7.

mCherry was incubated in buffers with different pH for 5 min. The fluorescence spectrum of mCherry at pH 11 and pH 6 was measured (λEx=570 nm, gain calculated from 2.5 µM Texas Red) using the TECAN infinite M200 and normalized to the maximum.
References
Prasher, D. C.; Eckenrode, V. K.; Ward, W. W.; Prendergast, F. G.; Cormier, M. J. (1992): Primary structure of the Aequorea victoria green-fluorescent protein. In: Gene 111 (2).
Shaner, Nathan C.; Campbell, Robert E.; Steinbach, Paul A.; Giepmans, Ben N. G.; Palmer, Amy E.; Tsien, Roger Y. (2004): Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. In: Nature biotechnology 22 (12).
Shu, X.; Remington, S. J. (2006): Crystal structure of mCherry.
Shu, Xiaokun; Shaner, Nathan C.; Yarbrough, Corinne A.; Tsien, Roger Y.; Remington, S. James (2006): Novel chromophores and buried charges control color in mFruits. In: Biochemistry 45 (32).
Inprovement by BNU-China 2021
edited by Qiuchen Gu
We construct the composite part consisting of two parts: OmpT signal peptide (Part:BBa_K3805532) and mCherry (BBa_J06504).Outer membrane protease T (OmpT) is a 33.5 kDa endoprotease located on the outer membrane of Escherichia coli.Ompt signal peptide is an excretion tag linked on the N-terminus of proteins.It can transport proteins across the outer membrane with the help of OmpT. In our part, Ompt signal peptide is fused with mCherry, which will enable mCherry to be secreted to extracellular space. For detailed information, please go to part Part:BBa_K3805886
Functional Parameters
| abs | -- |
| biology | -NA- |
| emission | 610 |
| emit | 610 |
| excitation | 587 |
| excite | 587 |
| lum | -- |
| protein | mCherry2 |
| tag | None |