Difference between revisions of "Part:BBa K2926089"
| (One intermediate revision by one other user not shown) | |||
| Line 3: | Line 3: | ||
<partinfo>BBa_K2926089 short</partinfo> | <partinfo>BBa_K2926089 short</partinfo> | ||
| − | + | <html> | |
| + | {{Template:Bielefeld-CeBiTec/Menu}} | ||
| − | <!-- | + | <!-- Achtung immer erst bild Links dann rechts sonst passt die nummerierung nicht!--> |
| − | + | ||
| − | < | + | <html> |
| − | + | Nowadays it is common for two plasmid systems to be used in the laboratory. In this case it is needed to work with at least two antibiotics to keep the plasmids into the cell. However, it is known, that antimicrobial resistance is becoming a growing threat, as a continued rise in resistance by 2050 would lead to 10 million people dying every year (AMR Review Paper—Tackling a crisis for the health and wealth of nations_1, 2014) and the use of high levels of antibiotics is cost intensive. What if we had an alternative on how to bypass working with two antibiotics? To enable this, we splitted the antibiotic resistance gene for chloramphenicol acetyl transferase (CAT) and cloned each part into a different vector. After co transforming cells with both vectors, through intein-mediated trans-splicing active CAT was produced. </div> | |
| − | < | + | </div> |
| + | </div> | ||
| − | <!-- | + | <div class="contentBlock"> |
| − | === | + | <div> |
| − | < | + | |
| − | <!-- --> | + | <a class="anchor" id="h1"></a> |
| + | <h1 id="a1"> Background </h1> | ||
| + | <hr> <!--Blaue Linie--> | ||
| + | |||
| + | |||
| + | <div class="quarter right"> | ||
| + | <figure class="figure large"> | ||
| + | <img style="width:400px" class="figure image" src="https://2019.igem.org/wiki/images/6/6d/T--Bielefeld-CeBiTec--Hygromycin_Bl.png"> | ||
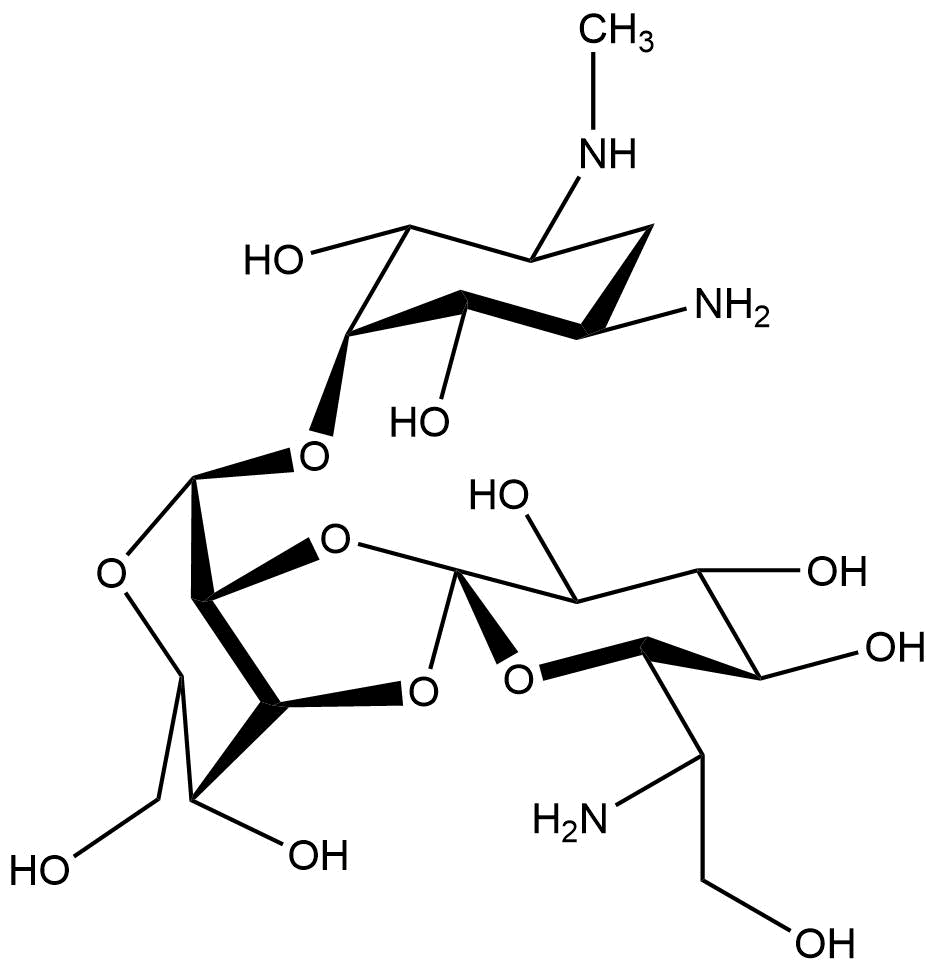
| + | <figcaption>Chemical structure of hygromycin B | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | </div> | ||
| + | |||
| + | <h2>Hygromycin B</h2> | ||
| + | |||
| + | <div> | ||
| + | Hygromycin B is an aminoglycoside that can kill prokaryotic and eukaryotic cells (Figure 1). It was developed in 1950 and used for animals as an anti-worming agent. This antibiotic is produced by Streptomyces hygroscopicus and later in 1980 the resistance gene was discovered (Gritz & Davies, 1983; Kaster, Burgett, Rao, & Ingolia, 1983). The mechanism of hygromycin B is to inhibit the enzymic translocation of peptidyl-tRNA and thus blocking the cytoplasmic protein synthesis (González, Jiménez, Vázquez, Davies, & Schindler, 1978). | ||
| + | </div> | ||
| + | |||
| + | <h2>Resistance</h2> | ||
| + | |||
| + | <div> | ||
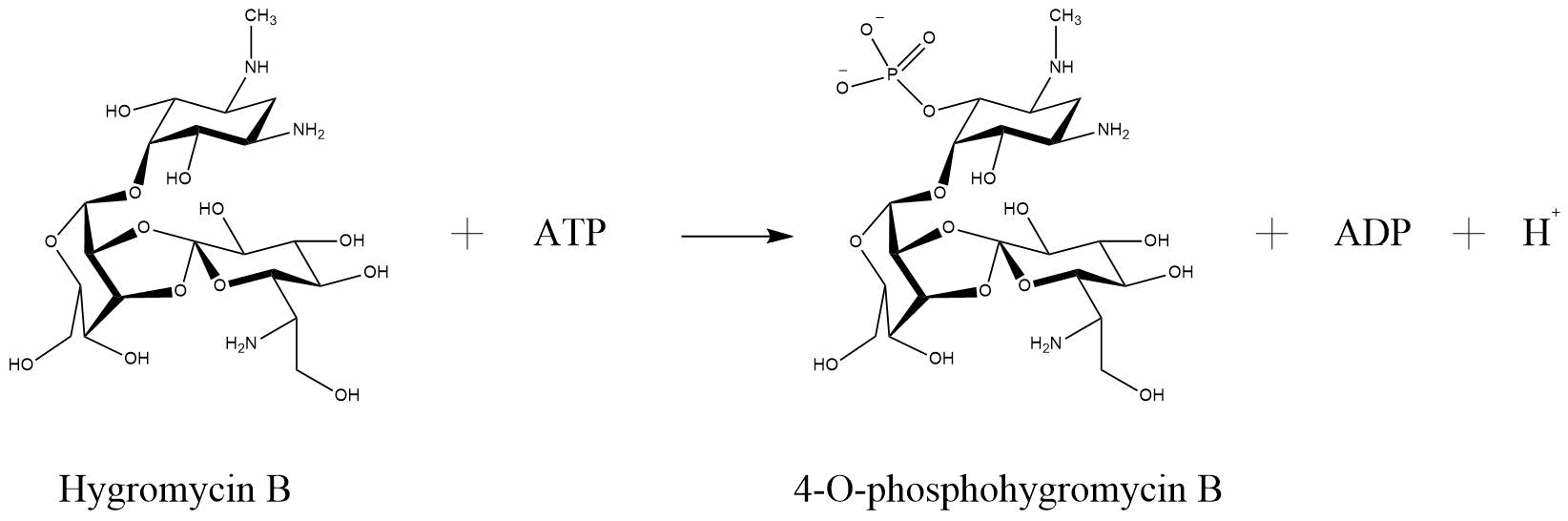
| + | One of the enzymes that mediates resistance against hygromycin B (hygB) is called hygromycin-B 4-O-kinase and was found in E. coli (Hph—Hygromycin-B 4-O-kinase, Uniprot). It has two substrates, both ATP and Hygromycin B. It inactivates the antibiotic through phosphorilation, so that it has as products phospho-hygromycin B and ADP (Figure 2). The same enzyme with the same characteristics, but from the organism Streptomyces fradiae named APH(4)-Ia, was tested for numerous aminoglycoside antibiotics, that perform possible substrates (Stogios et. al., 2011). The substrates that were tested included 4,6-disubstituted 2-deoxystreptamine-based, 4,5-disubstituted 2-deoxystreptamine-based and atypical aminoglycosides. The only antibiotic that was susceptible for phosphorylation by APH(4)-Ia was hygB. Such a specificity is not common for APHs in generall (Stogios et. al., 2011). | ||
| + | </div> | ||
| + | |||
| + | <div class="middle"> | ||
| + | <figure class="figure large"> | ||
| + | <img class="figure image" src="https://2019.igem.org/wiki/images/0/03/T--Bielefeld-CeBiTec--Hygromycin_B_Phosphorylation.png"> | ||
| + | <figcaption> Reaction of hygromycin B become inactivated through phosphorylation via hygromycin-B 4-O-kinase (Hph—Hygromycin-B 4-O-kinase, Uniprot) | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | </div> | ||
| + | |||
| + | <h2>Npu DnaE Intein</h2> | ||
| + | |||
| + | <div> | ||
| + | |||
| + | <div class="half left"> | ||
| + | <figure class="figure large"> | ||
| + | <img style="width:400px" class="figure image" src="https://2019.igem.org/wiki/images/4/47/T--Bielefeld-CeBiTec--basicpart_Intein3d.png" alt=""> | ||
| + | <figcaption>sNMR solution structure of NpuDnaE (PDB code: 2KEQ) (Oeemig et al. 2009)</figcaption> | ||
| + | </figure> | ||
| + | </div> | ||
| + | |||
| + | An intein is a part of a protein that excises itself from the precursor protein and fuses its exteins with a peptide bond through protein trans-splicing (Anraku, Mizutani, & Satow, 2005). For more information about the difference between split and non-split intein visit our <a href="https://parts.igem.org/Part:BBa_J31005"target="_blank"<>part collection</a>. The Npu DnaE intein is a subunit of the DNA polymerase III (DnaE) (Figure 3) from nostoc punctiforme and prefers specific extein sequences as flanks. The nativ N-exteins consists of the amino acids A E Y and the C-exteins of C F N (Cheriyan, Pedamallu, Tori, & Perler, 2013; Zettler, Schütz, & Mootz, 2009). However, Npu DnaE can splice a bride range of sequences, due to the variable extein types. Some result in faster splice rates in comparison to the native ones (Cheriyan et al., 2013). | ||
| + | </div> | ||
| + | |||
| + | <h2>Improved Part </h2> | ||
| + | |||
| + | <div> | ||
| + | For our project we improved the part <a href="https://parts.igem.org/Part:BBa_K678021"target="_blank"<>BBa_K678021</a> coding the Hygromycin-B 4-O-kinase. Since we use two plasmids to produce our Troygenics, we noticed that E. coli grows slower when containing two plasmids with diverse resistance genes. Additionally, it is widely known that multi resistant bacteria are becoming a growing threat as in agriculture and medicine, the usage of many antibiotics is common. However, this poses a problem, because a continued rise in resistance by 2050 would lead to 10 million people dying every year (AMR Review Paper—Tackling a crisis for the health and wealth of nations_1, 2014). By using many antibiotics simultaneously, the risk of bacteria becoming multi-resistant grows and can lead to massive crop loss and humans and animal deaths. For this reason, we thought that using only one resistance gene split in two plasmids would be a better approach. Therefore, we constructed plasmids the plasmids BBa_K2926036 and BBa_K2926037. | ||
| + | <br> | ||
| + | <br> | ||
| + | Our aim was to construct two plasmids that express the hygromycin B resistance gene, only by the presence of both plasmids into one E. coli cell. To achieve this, we had to use intein-mediated trans-splicing and choose the optimal split point. Latest, we did orientated on the paper of Jillette, Du, & Cheng, 2018, taking into consideration the amino acids relevant for the splicing process. The information about the flanking sequences the Npu DnaE intein prefers we took from Cheriyan et al., 2013. | ||
| + | For hygromycin we chose the split point 240-241 on the amino acid raster. | ||
| + | </div> | ||
| + | |||
| + | <h2>Mechanism</h2> | ||
| + | |||
| + | <div> | ||
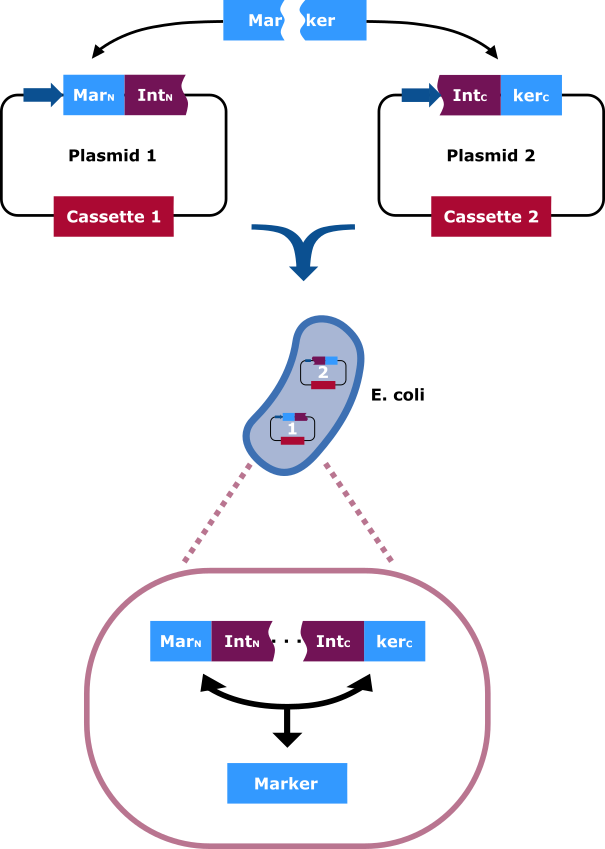
| + | The selection marker is split in two halfs. Each part of it is cloned into a different vector carrying distinct cassettes (Figure 4). The marekers N-terminal (MarN) and C-terminal (MarC) fragment is fused downstream and upstream respectively with the N-terminal (IntN) and C-terminal (IntC) fragment of the split Npu DnaE Intein. Only cells that contain both plasmids can grow on hygromycin rich medium. This occurs due to the intein fragments, because after the fusions being separately expressed, the split intein sections find one another and through trans splicing cut themselves out and fuse their flanking sites via peptide bond. Thereby an intact hygromycin-B 4-O-kinase is released and the cells gain the selection ability. </div> | ||
| + | </div> | ||
| + | <br> | ||
| + | <div class="middle"> | ||
| + | <figure class="figure large"> | ||
| + | <img style="width:400px" class="figure image" src="https://2019.igem.org/wiki/images/d/da/T--Bielefeld-CeBiTec--Marker_Intein_Schema.png"> | ||
| + | <figcaption>The selection marker is split in two halfs. The N-terminal and C-terminal fragment if it is fused upstream and downstream respectively of the N-terminal and C-terminal fragment of the Npu DnaE intein. Each fusion is cloned in a different backbone, so that the oris are compatible. On the protein level the split intein fragments find to each other and through protein trans-splicing they cut themselves out. Thereby an active hygromycin-B 4-O-kinase is assembled and the cells can grow on hygromycin B rich medium. | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | </div> | ||
| + | |||
| + | <h2>Plasmid structure</h2> | ||
| + | |||
| + | <div> | ||
| + | As previously mentioned for the hygromycin resistance (hygR) we constructed two plasmids. The N-terminal part of the resistance gene and intein, respectively is located on the first plasmid. On the second one the remaining parts were cloned (Figure 5,6). Upstream of the N-terminal Npu DnaE, the flanking amino acids from position -3 to -1 were W L A and downstream of the C-terminal intein from +1 to +3 C M E. These amino acids ensure a quicker slicing that the native ones. Each fusion has a different length. To keep the concentration of produced fusions at a close level, we cloned the longer one into the plasmid with the higher copy number and the shorter fusion into the lower copy number plasmid. | ||
| + | </div> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <div class="half left"> | ||
| + | <figure class="figure large"> | ||
| + | <img style="width:400px" class="figure image" src="https://2019.igem.org/wiki/images/a/a1/T--Bielefeld-CeBiTec--Plasmide_Hyg_Exteine1.png"> | ||
| + | <figcaption>The fusion on the first plasmid is illustrated. The N-terminal part of the HygR (HygRN) is shown in blue. The N-extein follows with the amino acids tryptophan, leucine and alanine (W L A) on positions -3, -2 and -1, respectively upstream of the N-terminal intein. | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | </div> | ||
| + | |||
| + | <div class="half right"> | ||
| + | <figure class="figure large"> | ||
| + | <img style="width:400px" class="figure image" src="https://2019.igem.org/wiki/images/8/82/T--Bielefeld-CeBiTec--Plasmide_Hyg_Exteine2.png"> | ||
| + | <figcaption>The fusion on the second plasmid is illustrated. The C-terminal part of the intein is shown. The C-extein follows with the amino acids cysteine, methionine and glutamic acids (C M E) on positions +1, +2 and +3, respectively upstream of the C-terminal HygR (HygRC). | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | </div> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <div class="third left"> | ||
| + | <figure class="figure large"> | ||
| + | <img style="width:400px" class="figure image" src="https://2019.igem.org/wiki/images/4/44/T--Bielefeld-CeBiTec--Single_Hyg.png"> | ||
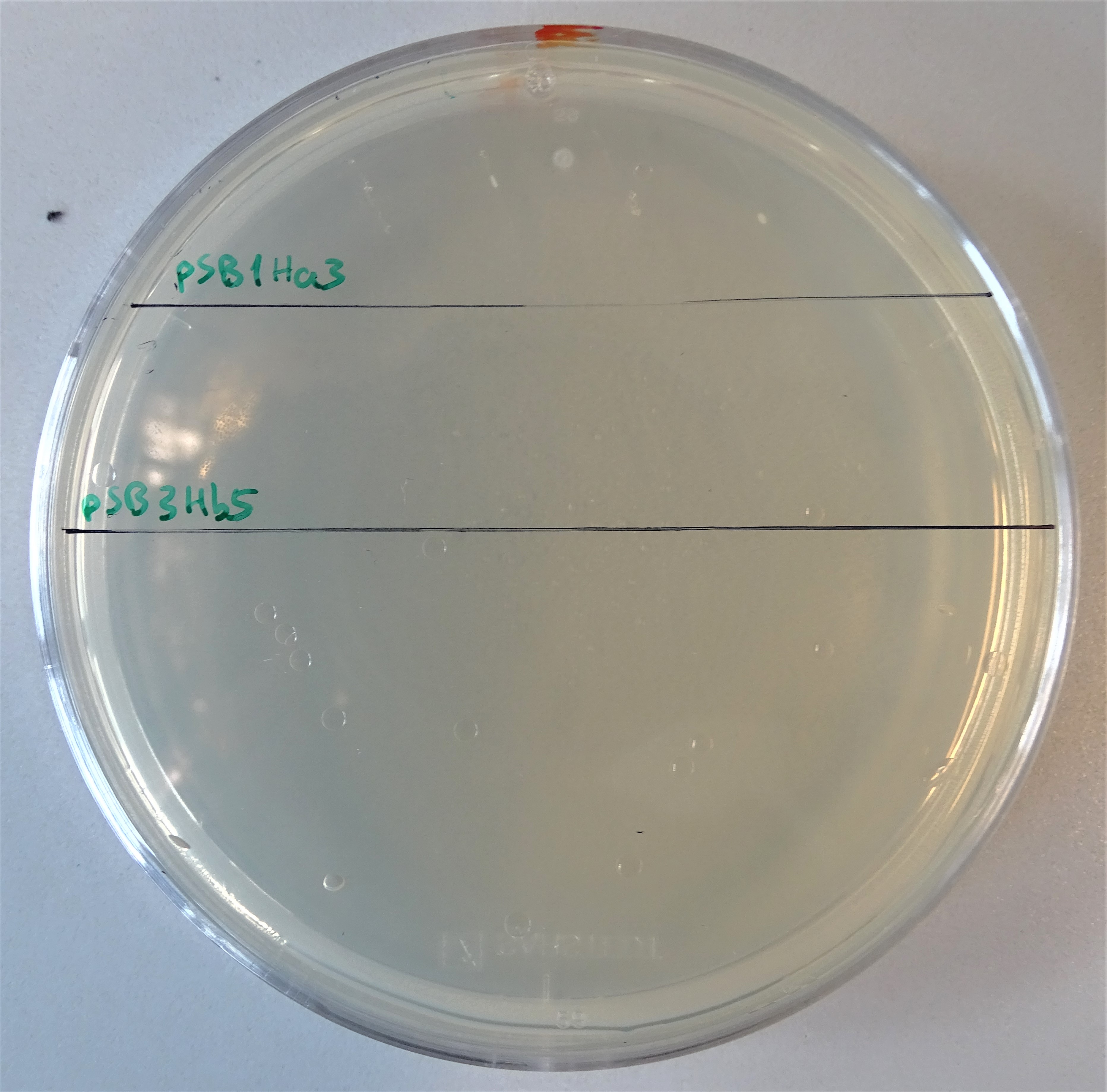
| + | <figcaption>Plate with a hygromycin B concentration of 50 μg/mL. On the upper half, cells were plated that contain only the pSB1Ha3 plasmid. On the lower half, cells were plated that contain only the pSB3Hb3 plasmid. | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | </div> | ||
| + | |||
| + | <div> | ||
| + | <h2 style="margin-top:1.8em;">Single plasmid control</h2> | ||
| + | |||
| + | After assembling the two plasmids required for expressing an active hph, we plated some kolonies on hygromycin B plates. On the one hand colonies containing the plasmid pSB1Ha3 and on the other hand pSB3Hb5. Through this way we wanted prove, that the half resistance gene fused to an intein is not able to to grow on hygromycin B (Figure 7). Normal working concentrations for hygromycin B is between 200 and 500 µg/mL (Hygromycin B, Thermofisher). Since no colonies grew on the plate it is proven, that cells that contain only one plasmid do not grow even in low hygromycin B concentrations. | ||
| + | </div> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <h2>Growth control</h2> | ||
| + | |||
| + | <div class="half right"> | ||
| + | <figure class="figure large"> | ||
| + | <img style="width:400px" class="figure image" src="https://2019.igem.org/wiki/images/4/4f/T--Bielefeld-CeBiTec--Split_Hyg.png"> | ||
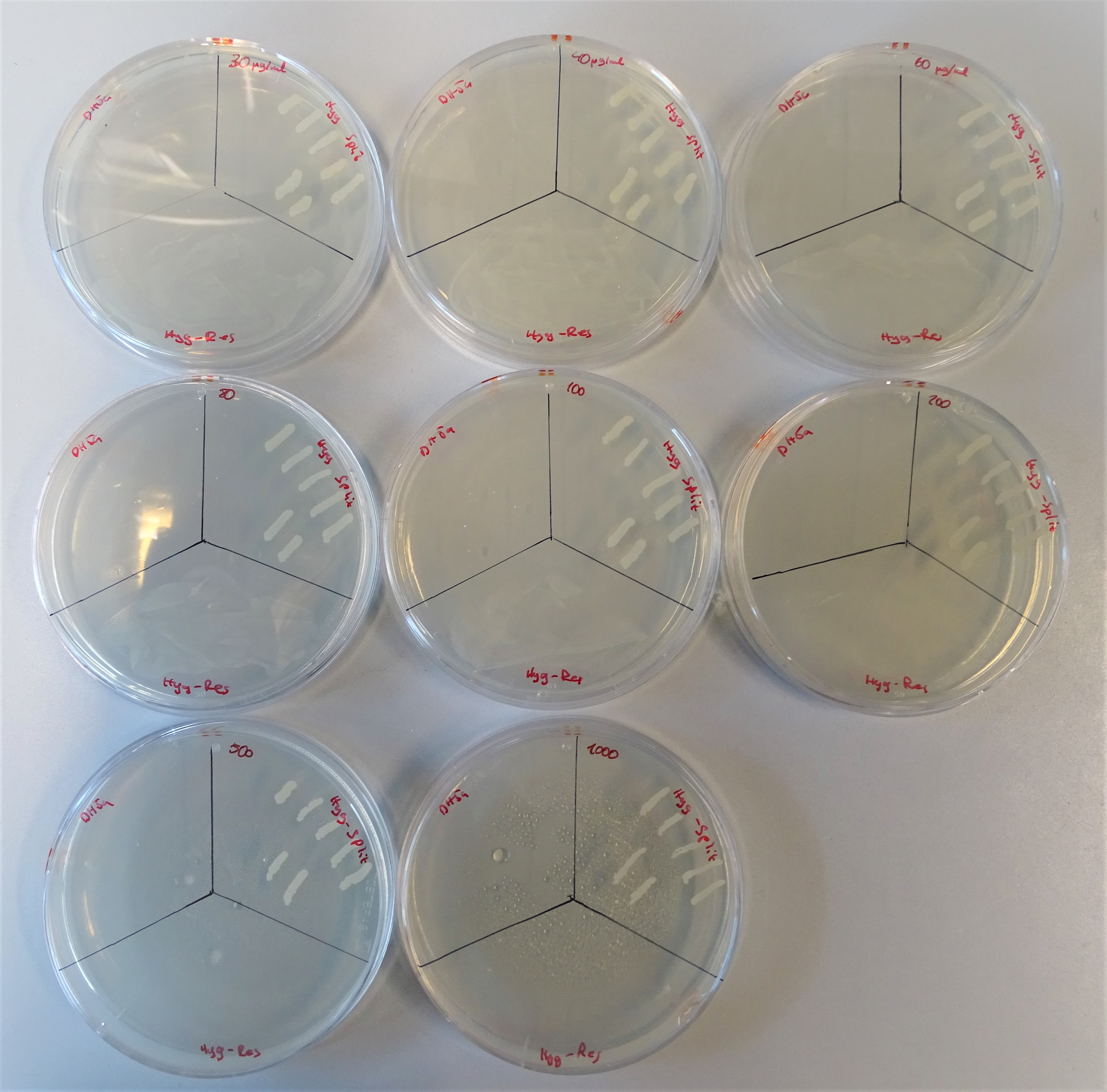
| + | <figcaption>From the upper left site to, to the right till the lower right site, plates are lined up with the hygromycin concentrations 40, 30, 60 80, 100, 200, 500, 1000μg/mL. Every plate is divided into three sections. On the upper left site DH5α was plated representing the negative control, on the upper right site colonies containing both plasmids and on the lower site a colony containing one plasmid with a hygromycin resistance gene. | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | </div> | ||
| + | |||
| + | <div> | ||
| + | Proving that the cells containing one plasmid do not grow was the first part of the characterisation of our new backbones. The next step was to co-transform cells with both plasmids and test their efficiency. To perform this, we poured plates with diverse concentrations of the hygromycin B antibiotic (Figure 8). The concentrations that we prepared were 30, 40, 60 80, 100, 200, 500 and 1000μg/mL. Each plate was divided into three sections. On the upper left site, DH5α was plated as the negative control, on the upper right site the colonies containing both plasmids and on the lower site a colony containing one plasmid with a hygromycin resistance gene. For DH5α these concentrations were to high to grow. The positive control did not have any problems growing on hygromycin B. The cells with the split antibiotic resistance plasmids easily grew on the plates till a concentration of 500 μg/mL and with some difficulties on 1000 μg/mL. | ||
| + | </div> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | </div> <!-- End content block --> | ||
| + | |||
| + | </div> <!-- End Wrapper--> | ||
| + | |||
| + | |||
| + | |||
| + | </body> | ||
| + | </html> | ||
Latest revision as of 03:52, 22 October 2019
Hygomycin resistance protein Part 1
{{Template:Bielefeld-CeBiTec/Menu}} Nowadays it is common for two plasmid systems to be used in the laboratory. In this case it is needed to work with at least two antibiotics to keep the plasmids into the cell. However, it is known, that antimicrobial resistance is becoming a growing threat, as a continued rise in resistance by 2050 would lead to 10 million people dying every year (AMR Review Paper—Tackling a crisis for the health and wealth of nations_1, 2014) and the use of high levels of antibiotics is cost intensive. What if we had an alternative on how to bypass working with two antibiotics? To enable this, we splitted the antibiotic resistance gene for chloramphenicol acetyl transferase (CAT) and cloned each part into a different vector. After co transforming cells with both vectors, through intein-mediated trans-splicing active CAT was produced.
Background
Hygromycin B
Resistance
Npu DnaE Intein

Improved Part
Our aim was to construct two plasmids that express the hygromycin B resistance gene, only by the presence of both plasmids into one E. coli cell. To achieve this, we had to use intein-mediated trans-splicing and choose the optimal split point. Latest, we did orientated on the paper of Jillette, Du, & Cheng, 2018, taking into consideration the amino acids relevant for the splicing process. The information about the flanking sequences the Npu DnaE intein prefers we took from Cheriyan et al., 2013. For hygromycin we chose the split point 240-241 on the amino acid raster.
Mechanism
Plasmid structure


Single plasmid control
After assembling the two plasmids required for expressing an active hph, we plated some kolonies on hygromycin B plates. On the one hand colonies containing the plasmid pSB1Ha3 and on the other hand pSB3Hb5. Through this way we wanted prove, that the half resistance gene fused to an intein is not able to to grow on hygromycin B (Figure 7). Normal working concentrations for hygromycin B is between 200 and 500 µg/mL (Hygromycin B, Thermofisher). Since no colonies grew on the plate it is proven, that cells that contain only one plasmid do not grow even in low hygromycin B concentrations.Growth control

