Difference between revisions of "Part:BBa K2259091"
(→This device in SynORI) |
|||
| Line 56: | Line 56: | ||
===Constitutive promoter=== | ===Constitutive promoter=== | ||
| − | Once the RNA I promoter was [https://parts.igem.org/wiki/index.php?title=Part:BBa_K2259000#RNA_I_inactivation_in_wild_type_replicon disabled] in the ColE1 origin of replication, it could be moved to a different plasmid location and used as a separate unit. We have discovered the sequence of wild type RNA I promoter by using PromoterHunter and | + | Once the RNA I promoter was [https://parts.igem.org/wiki/index.php?title=Part:BBa_K2259000#RNA_I_inactivation_in_wild_type_replicon disabled] in the ColE1 origin of replication, it could be moved to a different plasmid location and used as a separate unit. We have discovered the sequence of wild type RNA I promoter by using PromoterHunter and removed it, thus creating a wild type RNA I gene [[part:BBa_K2259005]]. First series of Anderson promoters were cloned next to the RNA I gene ([[part:BBa_K2259021]] (0.15 Anderson), [[part:BBa_K2259023]] (0.36 Anderson), [[part:BBa_K2259027]] (0.86 Anderson), [[part:BBa_K2259028]] (1.0 Anderson)) and then placed next to RNA II ([[part:BBa_K2259067]] (0.15 Anderson), [[part:BBa_K2259068]] (0.36 Anderson), [[part:BBa_K2259069]] (0.86 Anderson), [[part:BBa_K22590671]] (1.0 Anderson)). |
[[File:RNA_I_anderson.png|thumb|centre|900px|<b>Figure 4. </b> RNA I and RNA II constructs, with RNA I constructs under different-strength Anderson promoters.]] | [[File:RNA_I_anderson.png|thumb|centre|900px|<b>Figure 4. </b> RNA I and RNA II constructs, with RNA I constructs under different-strength Anderson promoters.]] | ||
| Line 68: | Line 68: | ||
===Inducible promoter=== | ===Inducible promoter=== | ||
| − | + | We wanted to move one step forward and try to build an inducible copy number system. We first had to make sure that at least part of our construct is well characterized and to do so we chose the Rhamnose promoter from the biobrick registry [[Part:BBa_K914003]] | |
| − | For this experiment we have built a | + | For this experiment we have built a Rhamnose promoter and RNA I construct [[part:BBa_K2259065]] and then cloned this construct next to RNA II [[part:BBa_K2259091]]. We have used different concentration of Rhamnose in our media in order to see if this approach was possible and if so, to figure out the dependency between the plasmid copy number and rhamnose concentration. |
[[File:Rha_rnr.png|thumb|centre|900px|<b>Figure 5. </b> RNA I and RNA II constructs, with RNA I gene being under the Rhamnose promoter, inducided by different rhamnose concentrations.]] | [[File:Rha_rnr.png|thumb|centre|900px|<b>Figure 5. </b> RNA I and RNA II constructs, with RNA I gene being under the Rhamnose promoter, inducided by different rhamnose concentrations.]] | ||
| − | The first thing we noticed was that | + | The first thing we noticed was that Rhamnose promoter was very strong in terms of plasmid copy number reduction. It was also considerably leaky (promoter can be enabled even without any inducer). At zero induction there were approximately only 9 plasmids per cell and at 1 percent induction the number droped to approximately 1 plasmid per cell. |
RNA I rhamnose-induced promoter seemed to be working well, with higher concentrations of inductor giving lower plasmid copy number. | RNA I rhamnose-induced promoter seemed to be working well, with higher concentrations of inductor giving lower plasmid copy number. | ||
| + | |||
We called it the SynORI copy number induction device. | We called it the SynORI copy number induction device. | ||
| − | So now when we can flexibly control the copy number of a | + | |
| + | So now when we can flexibly control the copy number of a plasmids, the only question is - what will come next? | ||
==References== | ==References== | ||
<references /> | <references /> | ||
Revision as of 23:34, 1 November 2017
SynORI inducible plasmid copy number device
This device consists of replication initiator RNA II that is modulated by replication regulator RNA I under the control of Rhamnose promoter.
Increased rhamnose concentrations leads to increase in RNA I concentration and consequently - decrease in plasmid copy number.
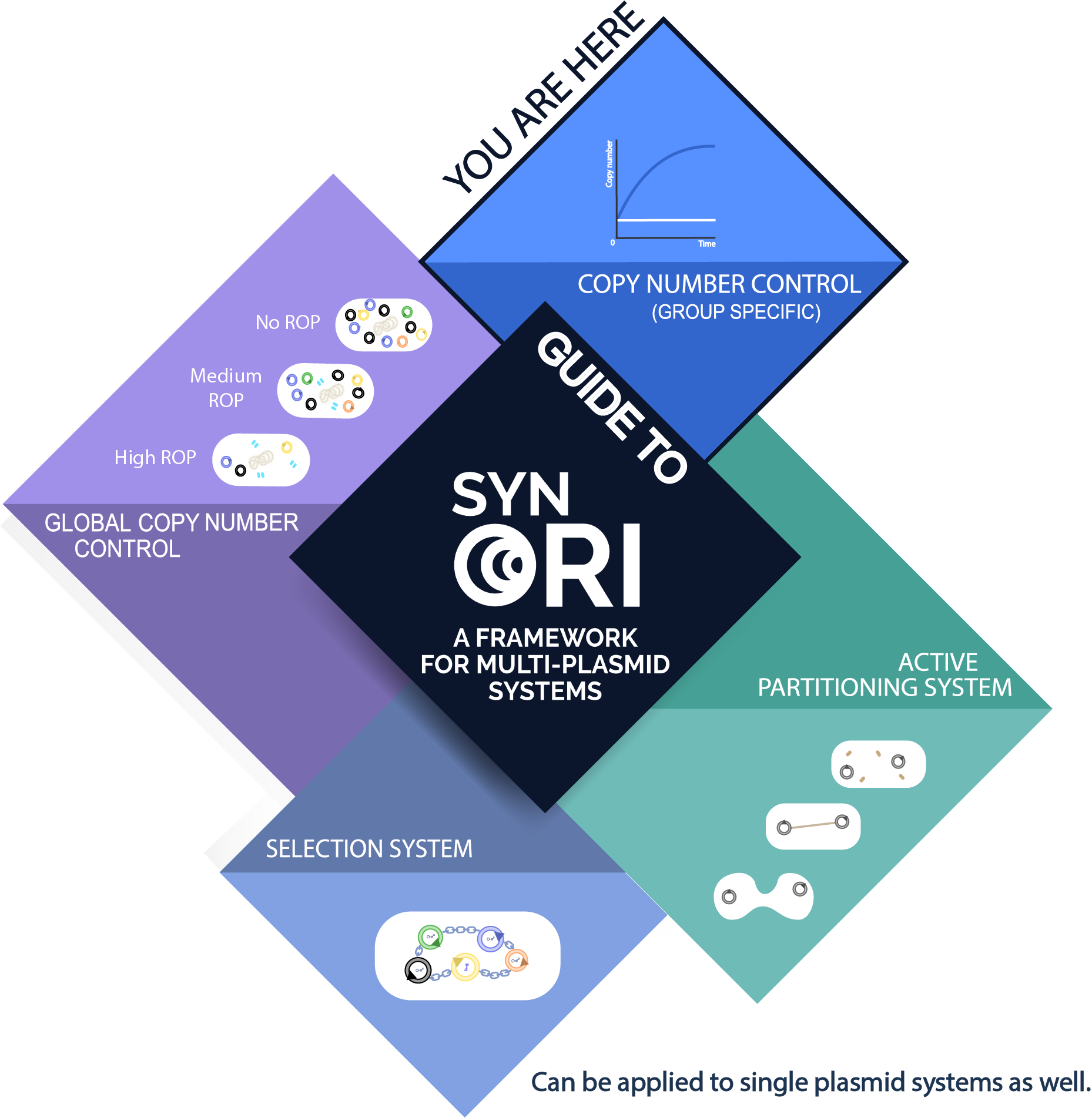
See how this part fits into the whole SynORI framework by pressing here!
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Introduction
Biology
ColE1 plasmid replication overview
ColE1-type plasmid replication begins with the synthesis of plasmid encoded RNA II</b> (also called primer transcript) by RNA polymerase which initiates transcription at a site 555bp upstream of origin of replication. The RNA transcript forms a RNA - DNA hybrid with template DNA near the origin of replication. Hybridized RNA is then cleaved at the replication origin by RNAse H and serves as a primer for DNA synthesis by DNA polymerase I (Figure 1. A).[1]
Initiation of replication can be inhibited by plasmid encoded small RNA, called RNA I . Synthesis of RNA I starts 445 bp upstream of the replication origin and proceeds in the direction opposite to that of RNA II synthesis and terminates near the RNA II transcription initiation site. RNA I binds to RNA II and thereby prevents the formation of a secondary structure of RNA II that is necessary for hybridization of RNA II to the template DNA (Figure 1. B).[2]
For RNA I to inhibit primer formation, it must bind before the nascent RNA II transcript extends to the replication origin. Consequently, the concentration of RNA I and the rate of binding of RNA I to RNA II is critical for regulation of primer formation and thus for plasmid replication. [3]
The interaction between RNA I and RNA II can be amplified by Rop protein, see part:BBa_K2259010.
Usage with SynORI (Framework for multi-plasmid systems)
About SynORI
SynORI is a framework for multi-plasmid systems created by Vilnius-Lithuania 2017 which enables quick and easy workflow with multiple plasmids, while also allowing to freely pick and modulate copy number for every unique plasmid group! Read more about [http://2017.igem.org/Team:Vilnius-Lithuania SynORI here]!
This device in SynORI
This device demonstrates SynORI framework ability to create an inducible plasmid copy number device. Different rhamnose concentration corresponds to different plasmid copy number, this relationship allows to manipulate copy number of plasmids when required without the need to construct new system or when changes needs to be made in a short period of time
Further details
For more background information and indepth insight on this part's design please see the individual parts page part:BBa_K2259000 and part:BBa_K2259005.
Characterization (Vilnius-Lithuania 2017)
Interaction between RNA I and RNA II groups
Constitutive promoter
Once the RNA I promoter was disabled in the ColE1 origin of replication, it could be moved to a different plasmid location and used as a separate unit. We have discovered the sequence of wild type RNA I promoter by using PromoterHunter and removed it, thus creating a wild type RNA I gene part:BBa_K2259005. First series of Anderson promoters were cloned next to the RNA I gene (part:BBa_K2259021 (0.15 Anderson), part:BBa_K2259023 (0.36 Anderson), part:BBa_K2259027 (0.86 Anderson), part:BBa_K2259028 (1.0 Anderson)) and then placed next to RNA II (part:BBa_K2259067 (0.15 Anderson), part:BBa_K2259068 (0.36 Anderson), part:BBa_K2259069 (0.86 Anderson), part:BBa_K22590671 (1.0 Anderson)).
In theory (see “Modelling” at [http://2017.igem.org/Team:Vilnius-Lithuania team Vilnius-Lithuania wiki]), lower-strength Anderson promoters should yield lower concentrations of RNA I, hence higher copy numbers of plasmids per cell. Our constitutive copy number device experiment results prove it to be true in practice as well. The stronger Anderson promoter is used, the less copy number per cell we get. With the strongest Anderson we get only 21+-6.84 plasmids per cell.
Worth to mention is that the closest to wild type ColE1 replicon is the 0.86 strength Anderson promoter (Part:BBa_J23102), measured by copy number alone.
We can state with certainty that we are now able to control the plasmid copy number in a constitutive manner, and we call it simply the SynORI constitutive copy number device.
Inducible promoter
We wanted to move one step forward and try to build an inducible copy number system. We first had to make sure that at least part of our construct is well characterized and to do so we chose the Rhamnose promoter from the biobrick registry Part:BBa_K914003
For this experiment we have built a Rhamnose promoter and RNA I construct part:BBa_K2259065 and then cloned this construct next to RNA II part:BBa_K2259091. We have used different concentration of Rhamnose in our media in order to see if this approach was possible and if so, to figure out the dependency between the plasmid copy number and rhamnose concentration.
The first thing we noticed was that Rhamnose promoter was very strong in terms of plasmid copy number reduction. It was also considerably leaky (promoter can be enabled even without any inducer). At zero induction there were approximately only 9 plasmids per cell and at 1 percent induction the number droped to approximately 1 plasmid per cell.
RNA I rhamnose-induced promoter seemed to be working well, with higher concentrations of inductor giving lower plasmid copy number.
We called it the SynORI copy number induction device.
So now when we can flexibly control the copy number of a plasmids, the only question is - what will come next?
References
- ↑ Itoh, T. and Tomizawa, J. (1980). Formation of an RNA primer for initiation of replication of ColE1 DNA by ribonuclease H. Proceedings of the National Academy of Sciences, 77(5), pp.2450-2454.
- ↑ Tomizawa, J. (1984). Control of cole 1 plasmid replication: The process of binding of RNA I to the primer transcript. Cell, 38(3), pp.861-870.
- ↑ Tomizawa, J. (1984). Control of cole 1 plasmid replication: The process of binding of RNA I to the primer transcript. Cell, 38(3), pp.861-870.