Difference between revisions of "Part:BBa K1172905"
| (One intermediate revision by the same user not shown) | |||
| Line 13: | Line 13: | ||
For more details about the seperate genes in this Biosafety-System and their function, click [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S here] | For more details about the seperate genes in this Biosafety-System and their function, click [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S here] | ||
<br><br> | <br><br> | ||
| − | For more details about the function of the Biosafety-System in general, click [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System here] | + | For more details about the function of the Biosafety-System in general, click [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System here]<br> |
| − | Moreover the BioBrick is not limited to this Biosafety-System, as it can be used for the regulated trancription of any coding sequence behind the P<sub>''BAD''</sub> promoter, opening the possibility of an antibiotic-free selection in the [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain Biosafety-Strain].< | + | Moreover the BioBrick is not limited to this Biosafety-System, as it can be used for the regulated trancription of any coding sequence behind the P<sub>''BAD''</sub> promoter, opening the possibility of an antibiotic-free selection in the [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain Biosafety-Strain].<br> |
| + | <br> | ||
<span class='h3bb'>'''Sequence and Features'''</span> | <span class='h3bb'>'''Sequence and Features'''</span> | ||
| Line 25: | Line 26: | ||
<partinfo>BBa_K1172905 parameters</partinfo> | <partinfo>BBa_K1172905 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
| + | |||
| + | Following, the characterization of this part in the Biosafety-System ''AraCtive'' are shown, but this part could be used for any other gene behind the P<sub>''BAD''</sub> promoter.<br> | ||
==='''Biosafety-System araCtive'''=== | ==='''Biosafety-System araCtive'''=== | ||
| Line 42: | Line 45: | ||
<br> | <br> | ||
| + | |||
| + | ==='''Characterization of the Biosafety-System araCtive'''=== | ||
| + | <br> | ||
| + | <p align="justify"> | ||
| + | The Biosafety-System araCtive was characterized on M9 minimal medium using glycerol as carbon source. As for the characterization of the pure arabinose promoter P<sub>''BAD''</sub> above, the bacterial growth and the fluorescence of GFP <bbpart>BBa_E0040</bbpart> was measured. Therefore, the wild type and the Biosafety-Strain ''E. coli'' K-12 ∆''alr'' ∆''dadX'', both containing the Biosafety-Plasmid <bbpart>BBa_K1172909</bbpart>, were cultivated once with the induction of 1% L-rhamnose and once only on glycerol.<br> | ||
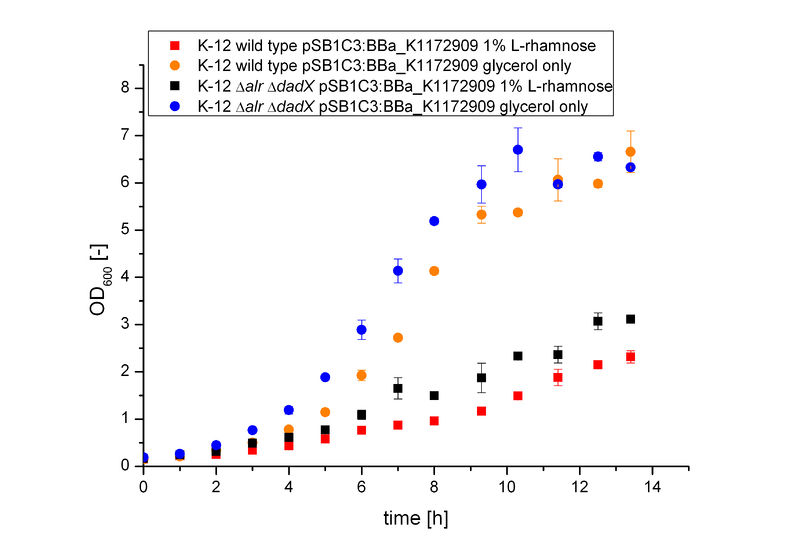
| + | It becomes obvious (Figure 6) that the bacteria, induced with 1 % L-rhamnose (red and black curve) grow significantly slower than on pure glycerol (orange and blue curve). This is attributed to the high metabolic burden encountered by the induced bacteria. The expression of the repressor AraC and the alanine racemase (Alr) simultaneously causes a high strain on the cells, so that they grow slower than the uninduced cells, which express only GFP.<br> | ||
| + | Comparing the bacterial growth with the fluorescence in Figure 7, it can be seen that the fluorescence seems to follow the same trend than the bacterial growth. The uninduced cells show approximately an exponential rise of fluorescence, while in comparision the fluorescence of the induced bacteria increases only slowly.</p><br> | ||
| + | |||
| + | [[File:Team-Bielefeld-Biosafety-System-araCtive-ODALL.jpg|600px|thumb|center|'''Figure 6:''' Characterization of the bacterial growth of the Biosafety-System on M9 minimal medium with glycerol. The Figure compares the wild type K-12 and the Biosafety-Strain K-12 ∆''alr'' ∆''dadX'' containing the Biosaftey-Plasmid <bbpart>BBa_K1172909</bbpart> and the induction by 1% L-rhamnose to pure glycerol.]] | ||
| + | <br> | ||
| + | [[File:Team-Bielefeld-Biosafety-System-araCtive-FlourescenceALL.jpg|600px|thumb|center|'''Figure 7:''' Characterization of the fluorescence of the Biosafety-System araCtive. The Figure compares the wild type K-12 and the Biosafety-Strain K-12 ∆''alr'' ∆''dadX'' containing the Biosafety-Plasmid <bbpart>BBa_K1172909</bbpart> and the induction by 1% L-rhamnose to pure glycerol..]] | ||
| + | <br> | ||
| + | <p align="justify"> | ||
| + | From the data presented above it can not be determined if the expression of the repressor AraC does affect the transcription of GFP or not. The slower growth of the bacteria is a first indication that the repressor AraC and the alanine racemase (Alr) are highly expressed, but the growth of the bacteria shows nearly the same kinetics as the fluorescence. So it could be possible that the repressor does not affect the expression level of GFP under the control of the arabinose promoter P<sub>''BAD''</sub>. This becomes more clear by the calculation of the specific production rate of GFP by equation (1) . As shown in Figure 8 below the specific production rate differs clearly between the uninduced Biosafety-System and the Biosafety-System induced by 1% L-rhamnose. The production of GFP in the presence of L-rhamnose (red curve) is always lower than in its absence (orange curve), so that the expression of GFP is repressed in the presence of L-rhamnose.<br> | ||
| + | |||
| + | Because the specific production rate of GFP was calculated between every single measurement point, the curve in Figure 8 is not smoothed and so the fluctuations have to be ignored, as they do not stand for are real fluctuations in the expression of GFP. They are caused by measurement variations in the growth curve and the fluorescence curve. But there is a clear tendency that the production of GFP is significantly lower when the bacteria are induced with 1% L-rhamnose. So the Biosafety-System araCtive works.</p><br> | ||
| + | |||
| + | [[File:Team-Bielefeld-Biosafety-System-araCtive-spezProductSara.jpg|600px|thumb|center|'''Figure 8:''' Specific production rate of GFP for the Biosafety-System araCtive, calculated via equation (1). The production rate of GFP of the uninduced bacteria is significantly higher compared to the bacteria induced with 1% L-rhamnose. The Biosafety-System AraCtive works.]] | ||
Latest revision as of 15:29, 30 October 2013
Part 1 of the Biosafety-System araCtive
Usage and Biology
These biobrick is the first part of the Biosafety-System AraCtive BBa_K1172909, which is an improvement of the Biobrick BBa_K914014 by replacing:
- the first promoter arabinose PBAD by the rhamnose promoter PRha to obtain a lower basal transcription.
- intgration of the alanine racemase BBa_K1172901(alr) for gaining higher plasmid stability and for taking advantage of the double-kill switch mechanism.
- the repressor LacI BBa_C0012 to AraC and the lac promoter to the arabinose promoter PBAD for a higher repression and lower basal transcription of the second part.
For more details about the seperate genes in this Biosafety-System and their function, click [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S here]
For more details about the function of the Biosafety-System in general, click [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System here]
Moreover the BioBrick is not limited to this Biosafety-System, as it can be used for the regulated trancription of any coding sequence behind the PBAD promoter, opening the possibility of an antibiotic-free selection in the [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain Biosafety-Strain].
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 1374
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 1298
Illegal BamHI site found at 2000 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 1416
Illegal AgeI site found at 1716 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 1173
Following, the characterization of this part in the Biosafety-System AraCtive are shown, but this part could be used for any other gene behind the PBAD promoter.
Biosafety-System araCtive
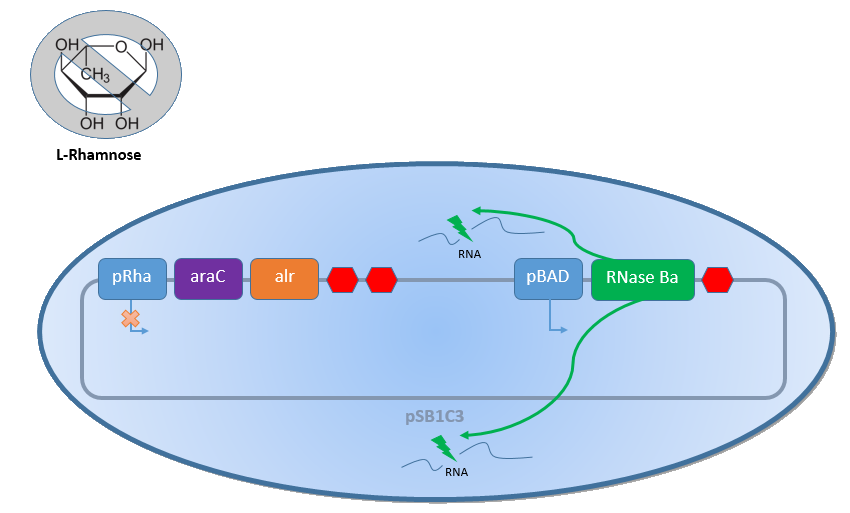
Combining the [http://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S genes] described above with the Biosafety-Strain K-12 ∆alr ∆dadX results in a powerful device, allowing us to control the bacterial cell division. The control of the bacterial growth is possible either active or passive. Active by inducing the PBAD promoter with L-arabinose and passive by the induction of L-rhamnose. The passive control makes it possible to control the bacterial cell division in a defined closed environment, like the MFC, by continuously adding L-rhamnose to the medium. As shown in the Figure 1 below, this leads to an expression of the essential alanine racemase (alr) and the AraC regulator, so that the expression of the RNase Ba is repressed.
In the event that bacteria exit the defined environment of the MFC or L-rhamnose is not added to the medium any more, both the expression of the alanine racemase (Alr) and the AraC regulatordecrease, so that the expression of the toxic RNase Ba (Barnase) begins. The cleavage of the intracellular RNA by the Barnase and the lack of synthesized D-alanine, caused by the repressed alanine racemase inhibit the cell division and makes sure that the bacteria can only grow in the defined environment or the device of choice respectively.
Characterization of the Biosafety-System araCtive
The Biosafety-System araCtive was characterized on M9 minimal medium using glycerol as carbon source. As for the characterization of the pure arabinose promoter PBAD above, the bacterial growth and the fluorescence of GFP BBa_E0040 was measured. Therefore, the wild type and the Biosafety-Strain E. coli K-12 ∆alr ∆dadX, both containing the Biosafety-Plasmid BBa_K1172909, were cultivated once with the induction of 1% L-rhamnose and once only on glycerol.
It becomes obvious (Figure 6) that the bacteria, induced with 1 % L-rhamnose (red and black curve) grow significantly slower than on pure glycerol (orange and blue curve). This is attributed to the high metabolic burden encountered by the induced bacteria. The expression of the repressor AraC and the alanine racemase (Alr) simultaneously causes a high strain on the cells, so that they grow slower than the uninduced cells, which express only GFP.
Comparing the bacterial growth with the fluorescence in Figure 7, it can be seen that the fluorescence seems to follow the same trend than the bacterial growth. The uninduced cells show approximately an exponential rise of fluorescence, while in comparision the fluorescence of the induced bacteria increases only slowly.

From the data presented above it can not be determined if the expression of the repressor AraC does affect the transcription of GFP or not. The slower growth of the bacteria is a first indication that the repressor AraC and the alanine racemase (Alr) are highly expressed, but the growth of the bacteria shows nearly the same kinetics as the fluorescence. So it could be possible that the repressor does not affect the expression level of GFP under the control of the arabinose promoter PBAD. This becomes more clear by the calculation of the specific production rate of GFP by equation (1) . As shown in Figure 8 below the specific production rate differs clearly between the uninduced Biosafety-System and the Biosafety-System induced by 1% L-rhamnose. The production of GFP in the presence of L-rhamnose (red curve) is always lower than in its absence (orange curve), so that the expression of GFP is repressed in the presence of L-rhamnose.
Because the specific production rate of GFP was calculated between every single measurement point, the curve in Figure 8 is not smoothed and so the fluctuations have to be ignored, as they do not stand for are real fluctuations in the expression of GFP. They are caused by measurement variations in the growth curve and the fluorescence curve. But there is a clear tendency that the production of GFP is significantly lower when the bacteria are induced with 1% L-rhamnose. So the Biosafety-System araCtive works.
References
- Baldoma L and Aguilar J (1988) Metabolism of L-Fucose and L-Rhamnose in Escherichia coli: Aerobic-Anaerobic Regulation of L-Lactaldehyde Dissimilation [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC210658/pdf/jbacter00179-0434.pdf|Journal of Bacteriology 170: 416 - 421.].
- Carafa, Yves d'Aubenton Brody, Edward and Claude (1990) Thermest Prediction of Rho-independent Escherichia coli Transcription Terminators - A Statistical Analysis of their RNA Stem-Loop Structures [http://ac.els-cdn.com/S0022283699800059/1-s2.0-S0022283699800059-main.pdf?_tid=ede07e2a-2a92-11e3-b889-00000aab0f6c&acdnat=1380629809_2d1a59e395fc69c8608ab8b5aea842f7|Journal of molecular biology 216: 835 - 858].
- Cass, Laura G. and Wilcoy, Gary (1988) Novel Activation of araC Expression and a DNA Site Required for araC Autoregulation in Escherichia coli B/r [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC211425/pdf/jbacter00187-0394.pdf|Journal of Bacteriology 170: 4174 - 4180].
- Hamilton, Eileen P. and Lee, Nancy (1988) Three binding sites for AraC protein are required for autoregulation of araC in Escherichia coli [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC279856/pdf/pnas00258-0029.pdf|Proc Natl Acad Sci U S A 85: 1749 - 53].
- Mossakowska, Danuta E. Nyberg, Kerstin and Fersht, Alan R. (1989) Kinetic Characterization of the Recombinant Ribonuclease from Bacillus amyloliquefaciens (Barnase) and Investigation of Key Residues in Catalysis by Site-Directed Mutagenesis [http://pubs.acs.org/doi/pdf/10.1021/bi00435a033|Biochemistry 28: 3843 - 3850.].
- Paddon, C. J. Vasantha, N. and Hartley, R. W. (1989) Translation and Processing of Bacillus amyloliquefaciens Extracellular Rnase [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC209718/pdf/jbacter00168-0575.pdf|Journal of Bacteriology 171: 1185 - 1187.].
- Schleif, Robert (2010) AraC protein, regulation of the L-arabinose operon in Escherichia coli, and the light switch mechanism of AraC action [http://gene.bio.jhu.edu/Ourspdf/127.pdf|FEMS microbial reviews 34: 779 - 796.].
- Voss, Carsten Lindau, Dennis and Flaschel, Erwin (2006) Production of Recombinant RNase Ba and Its Application in Downstream Processing of Plasmid DNA for Pharmaceutical Use [http://onlinelibrary.wiley.com/doi/10.1021/bp050417e/pdf|Biotechnology Progress 22: 737 - 744.].
- Walsh, Christopher (1989) Enzymes in the D-alanine branch of bacterial cell wall peptidoglycan assembly. [http://www.jbc.org/content/264/5/2393.long|Journal of biological chemistry 264: 2393 - 2396.]
- Wickstrum, J.R., Santangelo, T.J., and Egan, S.M. (2005) Cyclic AMP receptor protein and RhaR synergistically activate transcription from the L-rhamnose-responsive rhaSR promoter in Escherichia coli. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1251584/?report=reader|Journal of Bacteriology 187: 6708 – 6719.].


