Chassis/Cell-Free Systems/Commercial E.coli S30/Product Stability
Contents
Commercial E. coli S30
| Cell-Free Systems | Chassis description | Temperature dependence | DNA quantity dependence | Product stability | Peak time | Expression lifespan | Expression capacity |
Product stability
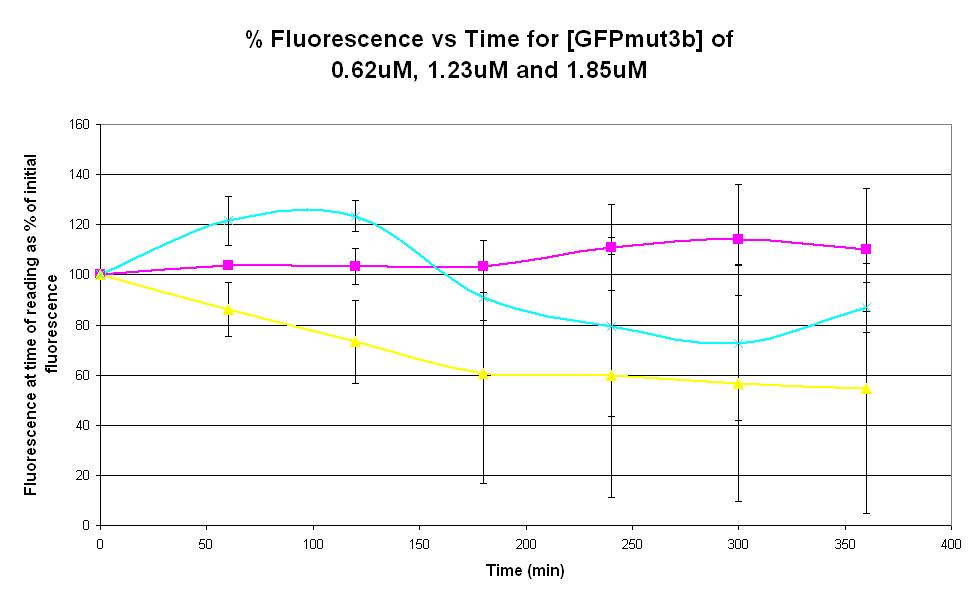
Graph 3. Graph of % fluorescence against time (minutes) for [GFPmut3b] of 0.62uM (pink), 1.23uM (yellow) and 1.85uM (light blue). Purified GFPmut3b was used. Please click [http://2007.igem.org/Imperial/Wet_Lab/Protocols/Prot1.6 here] for the protocol for purification of GFPmut3b. Each of the three coloured lines show average measurements based on two replicates of the experiment. The error bars represent the standard deviation of the measurements.
Experimental Protocol
Aims
- To determine the product stability of the commercial E. coli S30 chassis.
- This is done by measuring the fluorescence of GFPmut3b in the chassis over a 6 hour period, at GFPmut3b concentrations of 0.62uM, 1.23uM and 1.85uM.
Equipment
- Fluorometer + PC
- 1 Fluorometer Plate (Black)
- 25°C water bath
- Sticky plate lid
- Eppendorf tubes
- Gilson p20,p200,p1000
- Stop watch
Reagents
- Commercial S30 E.coli extract. Including:
- 175µl Amino Acid Mixture Minus Cysteine, 1mM
- 175µl Amino Acid Mixture Minus Methionine, 1mM
- 175µl Amino Acid Mixture Minus Leucine, 1mM
- 450µl S30 Extract, Circular (3 × 150µl)
- 750µl S30 Premix Without Amino Acids
- Nuclease Free water
- GFPmut3b Recombinant Protein (1mg/ml)
Preparation
- First collect all equipment and reagents and ensure that the fluorometer and that the PC connected has a data collection protocol installed.
- For the cell extract, get the following out of the cell extract kit:
- A.A's from kits
- Premix tube
- S30 tubes
- For each reaction we will have 40ul Cell extract +17ul DNA (2ug DNA) + 3 ul GFPmut3b dilution.
- To prepare the commercial E.coli Cell Extract (for 10 reactions), carry out the following Procedure:
- First prepare a complete amino acid mixture for the extract solution: Add the 25µl volume of two amino acid minus mixtures into an labeled eppendorf to give a volume of 50µl. Each amino acid minus mixture is missing one type of amino acid.
- Add 200µl of S30 Premix (Without Amino Acid) into the eppendorf tube.
- Then add 150µl of S30 Extract Circular too.
- This mixture is for all the samples. Label the tube.
- Any left over premix or cell extract should be returned to the freezer (biochemistry level 5) and labeled with new volumes.
- Combine the Cell Extract and AA to give a final volume : 400µl
- Now Prepare the different DNA concentration, we are using an empty vector (330ng/µl) because we do not want any expression but do want to have DNA present to simulate the in vitro chassis
- Remove 12.1µl of DNA in 170ul of total volume (make up with nuclease free water)
- This can be added to cell extract to give a final volume of 570ul, this is enough to make 10 reactions of 57ul, each of which will be made up to 60ul when GFPmut3b dilution is added.
- Finally we need to prepare the GFPmut3b dilutions, we use a commercial recombinant protein of 1mg/ml. We need enough for 2 reactions and some excess so we will prepare 9ul dilutions.
- 1.85uM = 9ul GFPmut3b into an eppendorf tube
- 1.23uM = 6ul GFPmut3b + 3ul of Nuclease Free WATER
- 0.62uM = 3ul GFPmut3b + 6ul of Nuclease Free Water
- 0uM = 9ul Nuclease Free Water
Plate loading
- First read the background fluorescence of the 96-well plate using the fluorometer.
- Choose suitable wells, with minimum fluorescence (30-40 au) to put the samples in. Do not use the wells at the edges and avoid putting samples in consecutive wells.
- Add 57ul of cell extract+DNA solution to the correct wells following the schematic. When pipetting it is best to pipette straight down into the base of the wells and avoide getting solution on the sides of the wells.
- Then add 3ul of the correct GFPmut3b dilution to the correct well following the schematic. Ensure that this is done as quickly as possible to avoid degradation.
- Remove lid off the 96 well plate and place in the fluorometer.
- Measure the plate in the fluorometer. This is the first reading.
- Place the plate in the fluorometer to measure its initial fluorescent reading.
- After the measurement, place the sticky tape across the plate, and put the plate in the 25oC water bath.
- Before placing in the water bath, wrap aluminium foil around the plate to prevent photobleaching.
- Repeat the reading every 1 hour, until 2hours have elapsed.
- Plot a graph of % fluorescence against time for the 3 protein concentrations.
Results
Our raw data and data analysis can be found here: File:S30 Product Data.xls
