Part:BBa_K823007
Anderson promoter J23103
Anderson promoter J23103 from the Anderson collection without RFP in backbone vector pSB1C3 to easily fuse the promoter with other reporters e.g. the lux operon or the lacZ reporter gene.
Usage and Biology
The first group of promoters evaluated are the promoters of the Anderson collection ("Anderson promoters"). They have already been measured in Escherichia coli where they all showed a constitutive behavior with different strength. In this project, eleven Anderson promoters were characterized in B. subtilis with the lux operon as a reporter. In B. subtilis these promoters show quiet low activity. To confirm these results some Anderson promoters were also evaluated with the reporter gene lacZ by doing β-galactosidase assays. For more details visit the [http://2012.igem.org/Team:LMU-Munich/Data/Anderson Data] page of the LMU-Munich Team 2012 or get an overview of our whole project [http://2012.igem.org/Team:LMU-Munich Beadzillus].
Here you can see the list with the Anderson promoters characterized in B. subtilis:
- J23100 (BioBrick:BBa_K823004)
- J23101 (BioBrick:BBa_K823005)
- J23102 (BioBrick:BBa_K823006)
- J23103 (BioBrick:BBa_K823007)
- J23106 (BioBrick:BBa_K823008)
- J23107 (BioBrick:BBa_K823009)
- J23113 (BioBrick:BBa_K823010)
- J23114 (BioBrick:BBa_K823011)
- J23115 (BioBrick:BBa_K823012)
- J23117 (BioBrick:BBa_K823013)
- J23118 (BioBrick:BBa_K823014)
Evaluation
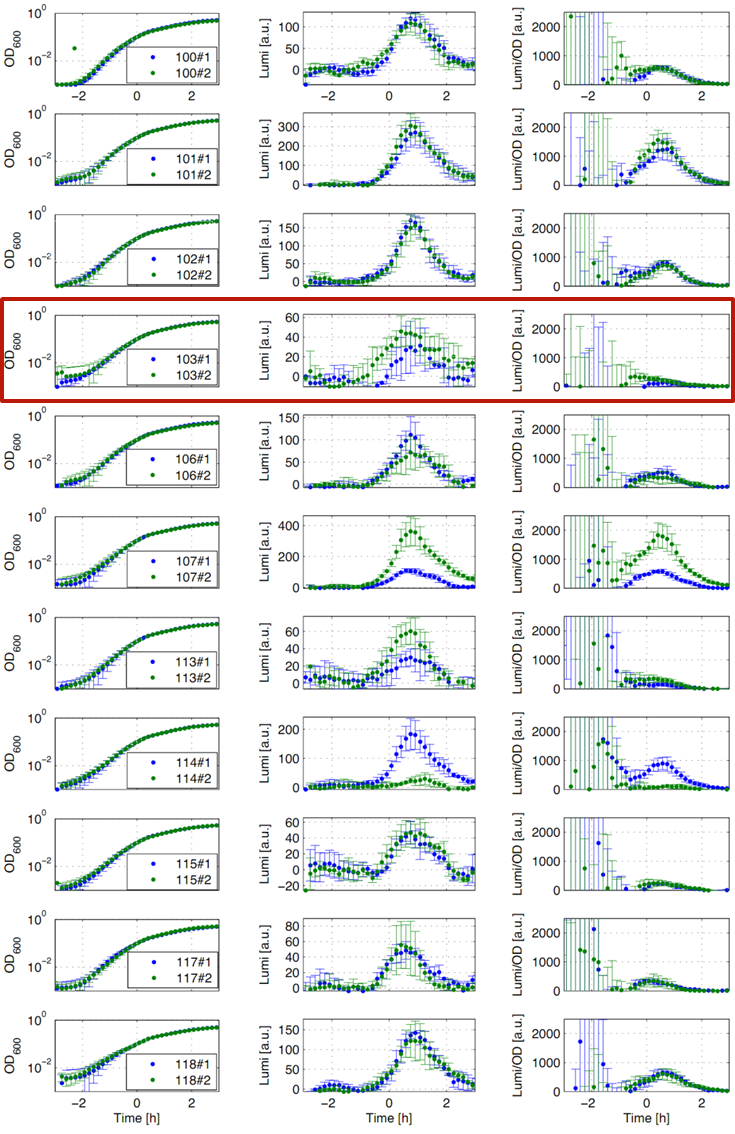
Eleven of the nineteen promoters of the Anderson collection (J23100, J23101, J23102, J23103, J23106, J23107, J23113, J23114, J23115, J23117, J23118) were evaluated. Therefore, we used the reporter vector pSBBs3C-luxABCDE from the BioBrickBox containing the lux operon as a reporter for promoter activity. The promoter activity leads to the expression of the lux operon and to the production of the enzyme luciferase. The luminescence, which is produced by the luciferase, can be measured with the plate reader Synergy2 (BioTek) (Fig.1).
|
All clones show a usual growth behaviour. The activity of the promoters increases during transition from log to stationary phase. The maximum (t=1h) reaches from 200Lumi/OD600 (promoter J23115) to a maximum of 1500 Lumi/OD600 for the strongest promoter (J23101). Afterwards the activity goes down to the beginning level (t=2h). The oscillation of luminescence (Lumi/OD600) in the beginning of the curves are due to the small OD600 values and do not mean high promoter activity. One clone of J23107 and J23114 shows significantly lower promoter activity. Therefore, additional clones should be measured. In comparison to all the other evaluated Bacillus promoters, these Anderson promoters showed a very low acitivity in B. subtilis.
β-galactosidase assay
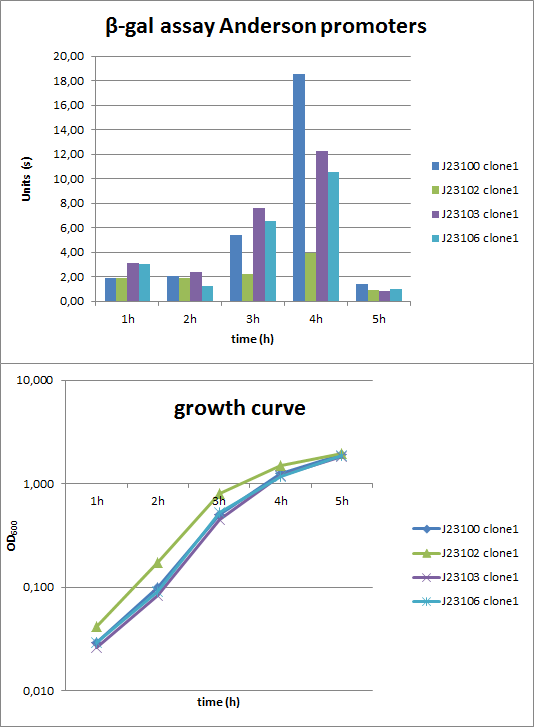
To evaluate the activity not only with the lux reporter operon, four promoters of the Anderson collection were cloned into the reporter vector pSBBs1C-lacZ to do β-galactosidase assays and then to compare the results of the strength of these promoters in B. subtilis (Fig. 2). The results were compared to the results from the luminescence measurements.
|
Verification of the promoter activity by β-galactosidase assays revealed that the Anderson promoters do not seem to be as weak as measured by luminescence (Fig. 1). For direct comparison we should measure a constitutive promoter, e.g. PliaG, in the same experiment. But we think the luminescence measurements are more reliable because they were repeated three times with two independent clones.
Sequence and Features
This part is a modified BioBrick. We got the DNA from the Partsregistry, removed RFP and send it in again in pSB1C3.
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
//chassis/prokaryote/subtilis
//promoter/anderson
//regulation/constitutive
//rnap/prokaryote/ecoli/sigma70
| None |