|
This BioBrick was designed to be used as an integration backbone for the ‘’B. subtilis’’ by integrating at the ‘’sacA’’ region of the endogenous chromosome via double crossover.
The Groningen iGEM 2012 team has showed that this BioBrick can be replicated in ‘’E. coli’’, however have not showed any results/characterisation that this backbone can integrate correctly in ‘’B. subtilis’’.
We transformed this backbone into ‘’B. subtilis’’ but as can be seen in Figure 1, no colonies were found on the ‘’B. subtilis’’ str. 168 + pSac-Cm derived integration plasmid, however the positive control ‘’B. subtilis’’ str.168 + pGFPrrnB (integrates at amyE) did work, which suggested that this backbone was not integrated.
 Plate 1: B. Subtilis str. 168 transformed with pGFPrrnB. |
 Plate 2: B.subtilis str. 168 transformed with H20(negative control). |
 Plate 3: B. subtilis str. 168 with pSac + Cm derived plasmid.
|
Figure 1. Plates of B. Subtilis str. 168 transformed with the pSac+Cm derived plasmid, pGFPrrnB (positive control) and water (negative control). |
We repeated the transformations using higher concentration of plasmids 5ug, 10ug, and 15ug and plated them onto the LB + 5ug/ml Chloramphenicol plates. The results Figure 2, shows that there were colonies growing on the 10ug and 15ug plates.
 B. subtilis str. 168 with 5ug pSac + Cm derived plasmid. |
 B. subtilis str. 168 with 10ug pSac + Cm derived plasmid. |
 B. subtilis str. 168 with 15ug pSac + Cm derived plasmid. |
 B.subtilis str. 168 transformed with H20(negative control) |
 B. Subtilis str. 168 transformed with pGFPrrnB. |
Figure 1. Plates of B. Subtilis str. 168 transformed with 5,10 and 15ug pSac+Cm derived plasmid, pGFPrrnB (positive control) and water (negative control). |
To test for integration, we used the Phenol red Sucrose test; the media where the transformants from the 10ug and 15ug plates were inoculated showed the same results as the control (‘’B. subtilis’’ + pGFPrrnB) with pH ranging between 4.6 – 5.2 suggesting that they could utilized sucrose as their carbon source and produced the acid by products, Figure 3.
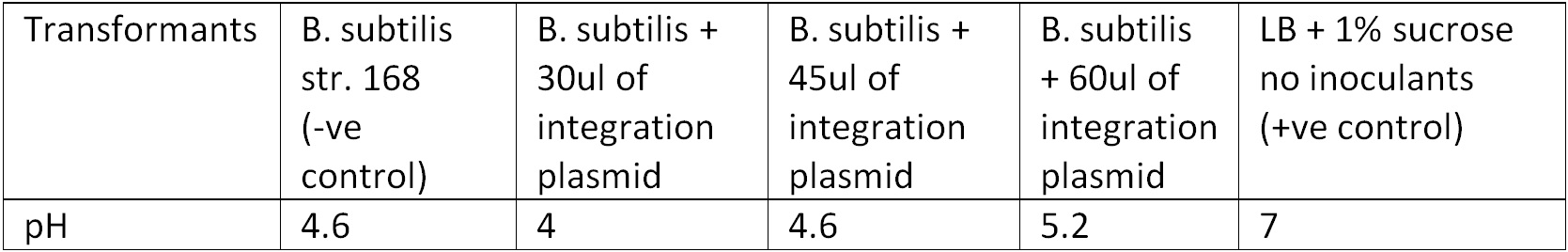

Figure 3. Display the pH of each samples following the Phenol Red Sucrose test on overnight grown culture in LB + 1% Sucrose media.
We then sequenced the backbone and found out that there were 49 mutations which include SNV, deletions and insertions.Table 1 display the mutations found on the sequence.
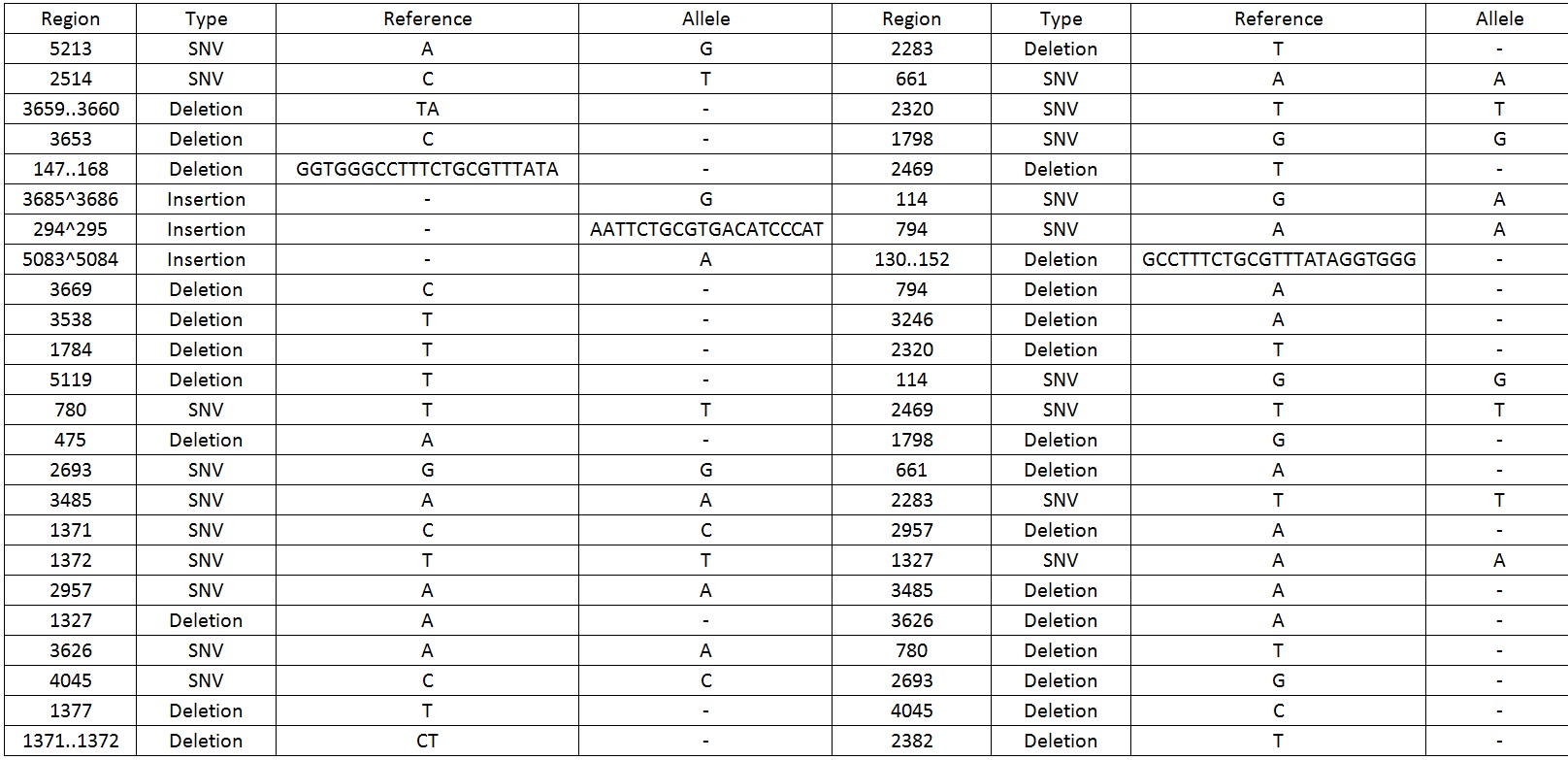 Table 1. Shows all the mutations found in the BBa_K818000 backbone.
Table 1. Shows all the mutations found in the BBa_K818000 backbone.
Table 2. Shows list of mutations found in the BBa_K818000 backbone, including the position, region and type of mutations after analysing the initial sequencing and the sequencing of highly mutated regions.
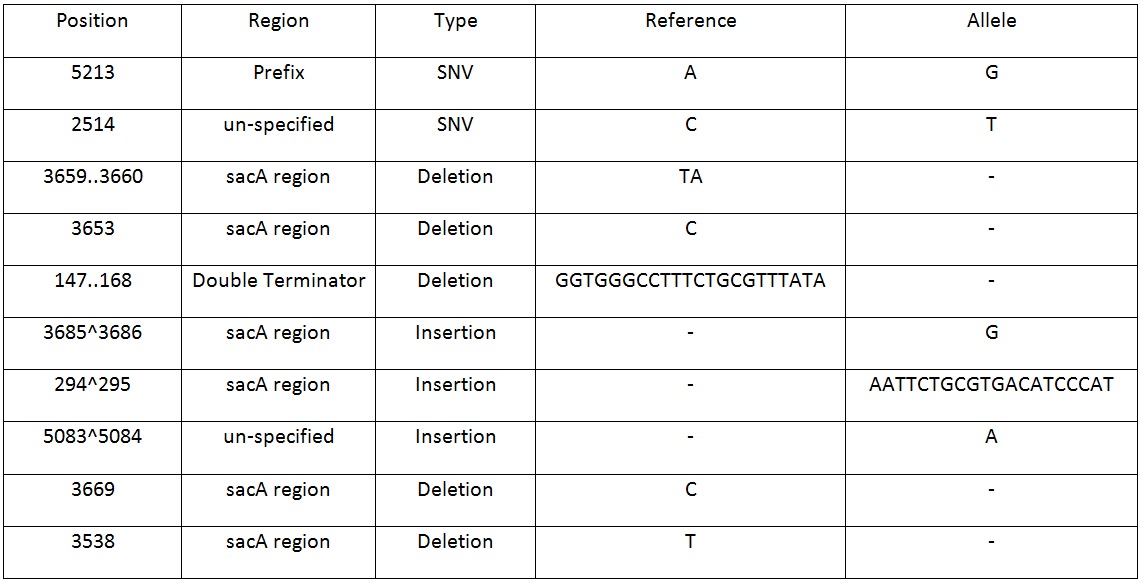 Table 2. Shows list of mutations found in the BBa_K818000 backbone, including the position, region and type of mutations after analysing the initial sequencing and the sequencing of highly mutated regions.
Table 2. Shows list of mutations found in the BBa_K818000 backbone, including the position, region and type of mutations after analysing the initial sequencing and the sequencing of highly mutated regions.
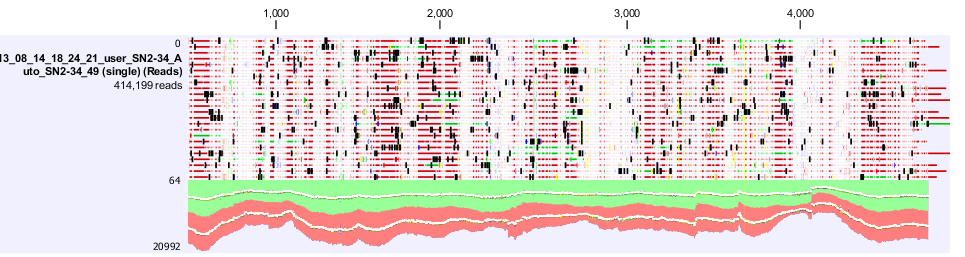
Figure 4. shows a screen shot of the pSac-Cm derived integration plasmid sequence analysis that we performed. It shows the amount of coverage, where the mutations are.
The results from both sequencing run proved to show similar mutations were found on this backbone, most of the mutations occurred in the sacA integration regions. These results explained the reason why this pSac-Cm derived integration backbone for ‘’B.subtilis’’ were not working.
To get round this problem, we align the sequence of this BioBrick to the Integration vector pSac-Cm sequence from the (Middleton, R., Hofmeister, A. New shuttle vectors for ectopic insertion of genes into Bacillus subtilis. Plasmid Volume 51, Issue 3, May 2004, Pages 238–245). The results showed that the sequence that the Groningen team 2012 put up on the registry was correct. This suggests that the plasmid that they have submitted and the sequence they provided did not match. By using the correct sequence to generate this integration plasmid we will be able to make this part functional not just in ‘’E. coli’’ but also ‘’B. subtilis.
| 










