Part:BBa_K525303
S-Layer SgsE from Geobacillus stearothermophilus with T7/lac
S-Layer SgsE from Geobacillus stearothermophilus
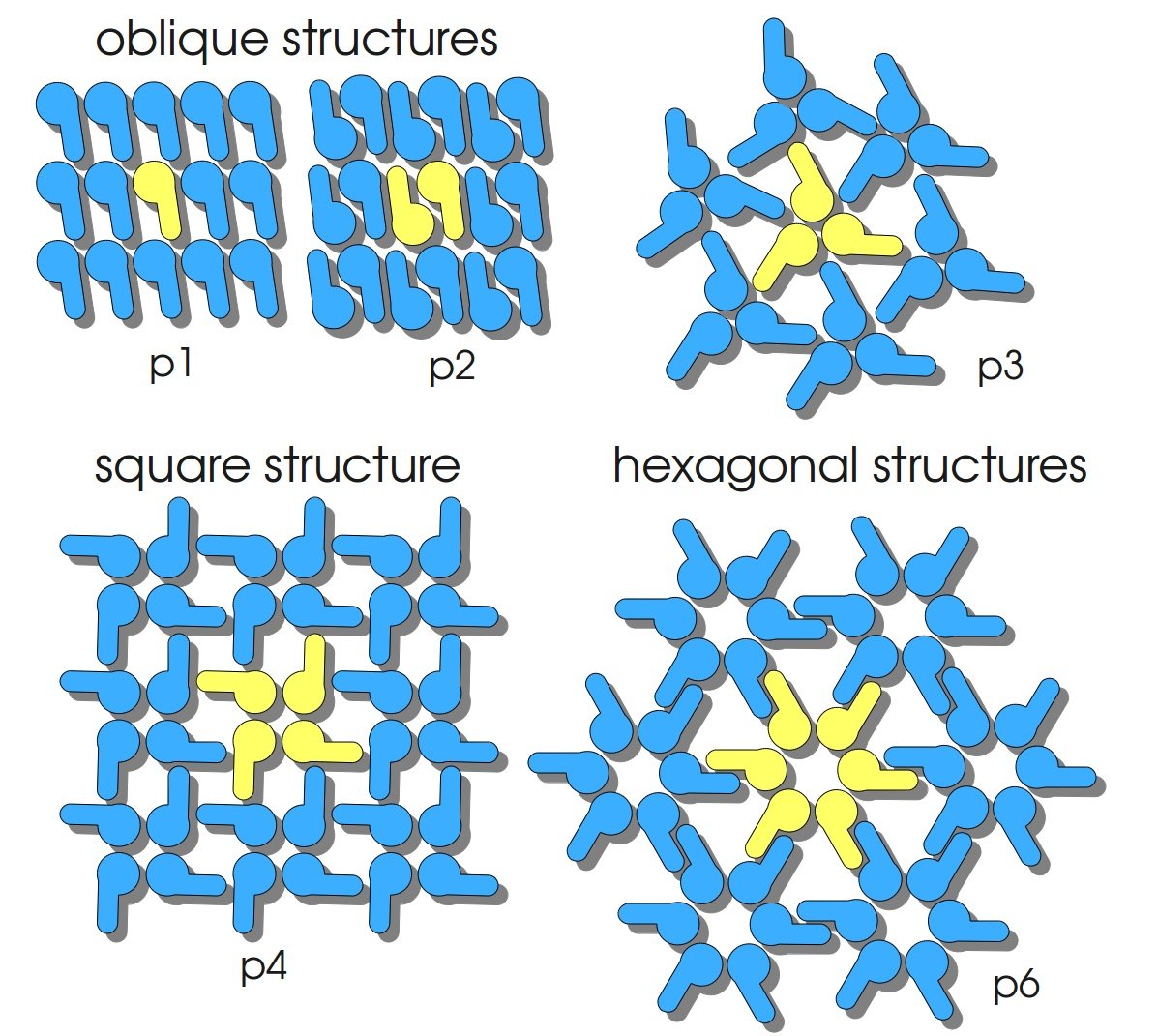
S-layers (crystalline bacterial surface layer) are crystal-like layers consisting of multiple protein monomers and can be found in various (archae-)bacteria. They constitute the outermost part of the cell wall. Especially their ability for self-assembly into distinct geometries is of scientific interest. At phase boundaries, in solutions and on a variety of surfaces they form different lattice structures. The geometry and arrangement is determined by the C-terminal self assembly-domain, which is specific for each S-layer protein. The most common lattice geometries are oblique, square and hexagonal. By modifying the characteristics of the S-layer through combination with functional groups and protein domains as well as their defined position and orientation to eachother (determined by the S-layer geometry) it is possible to realize various practical applications ([http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.2006.00573.x/full Sleytr et al., 2007]).
Usage and Biology
S-layer proteins can be used as scaffold for nanobiotechnological applications and devices by e.g. fusing the S-layer's self-assembly domain to other functional protein domains. It is possible to coat surfaces and liposomes with S-layers. A big advantage of S-layers: after expressing in E. coli and purification, the nanobiotechnological system is cell-free. This enhances the biological security of a device.
This S-layer gene is downsteam of a T7 / lac fusion promoter and in the Freiburg BioBrick Assembly standard so functional protein domains can easily be fused to its N-terminus and expressed and purified afterwards. SgsE was characterized with BBa_K525305.
Important parameters
| Experiment | Characteristic | Result |
|---|---|---|
| Expression (E. coli) | Localisation | Inclusion body |
| Compatibility | E. coli KRX and BL21(DE3) | |
| Induction of expression | expression of T7 polymerase + IPTG or lactose | |
| Purification | Molecular weight | 83.2 kDa |
| Theoretical pI | 5.75 | |
| Immobilization behaviour | Immobilization time | 4 h |
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 167
Illegal BglII site found at 1022 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 76
Illegal AgeI site found at 2401 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 1657
//proteindomain/internal
//rnap/bacteriophage/t7
| None |