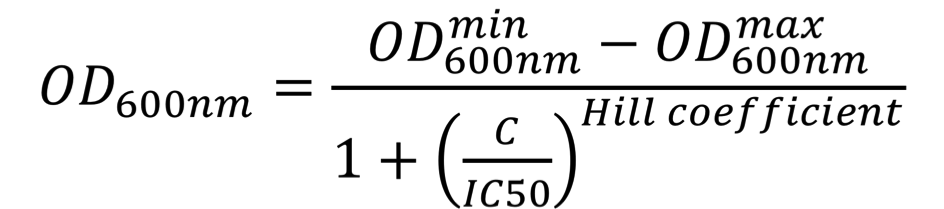Part:BBa_K2796014
AmpR
Ampicillin resistance gene is the most widely used selective marker in gene manipulation. Most plasmid vectors cloned in E. coli carry this gene. Penicillin can inhibit the synthesis of cell wall peptidoglycans, bind to related enzymes, inhibit their activities, and inhibit the transpeptidase reaction. The ampicillin-resistant gene (AmpR) encodes an enzyme that secretes into the periplasm of bacteria, inhibits transpeptidation and catalyzes the hydrolysis of beta-lactam cycles, thus eliminating the toxicity of ampicillin.
Contribution by iGEM Evry Paris-Saclay 2021
This part is AmpR, which is a derivative of the TEM-1 β-lactamase (Uniprot P62593) with two mutations V82I and V182A. Six sequences encoding it are found in the Parts Registry (BBa_K2082292, BBa_Y1002, BBa_I722014, BBa_K1927002, BBa_K2308011 and BBa_K2796014). BBa_K2082292 differs from all the other five by a synonymous mutation: T/C in position 132.
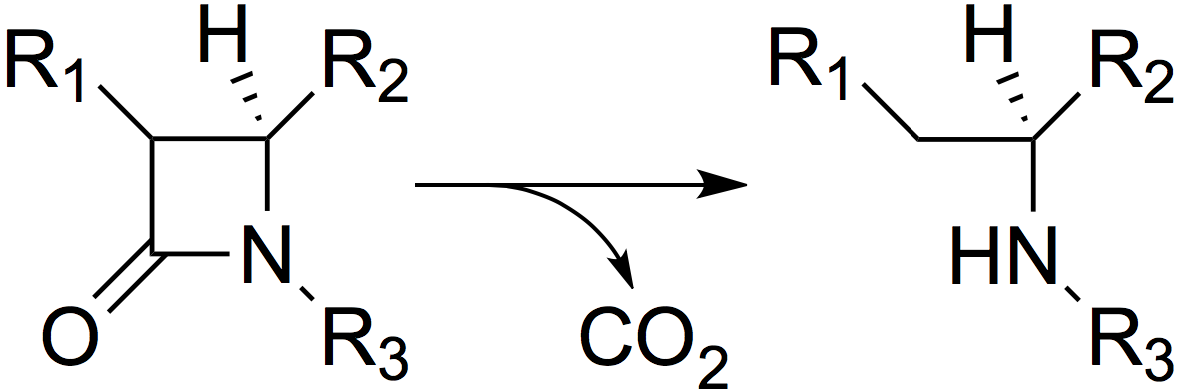
β-lactamases are enzymes secreted extracellularly by some bacteria. They allow the bacteria to resist β-lactam antibiotics by inactivating them before they can act (Figure 1). The best known β-lactamase to date is TEM-1, discovered in 1960, which allows resistance to penicillins and cephalosporins [1].
Figure 1. The mode of action of β-lactamases: hydrolysis of the β-lactam ring.
β-lactam antibiotics represent a large proportion of clinically used antibiotics. They are classified into different families based on their structure: penams (ex. penicillin), penems (ex. carbapenem), cephems (ex. cephalosporin) and monobactams (ex. aztreonam). These β-lactams have a bactericidal action [2]. Their mechanism of action involves crossing the peptidoglycan (PG) in the bacterial cell wall until they reach the enzymes called PBPs (penicillin binding proteins), which they inhibit thus preventing the synthesis of PG and promoting the release of an autolysin that will digest the PG. The mode of action of these β-lactams depends primarily on the presence of PBPs. Indeed, the efficacy of the antibiotic depends on the affinity it has to the PBP, the number of PBPs that the bacterium has on the surface of its peptidoglycan, but also their structure, which may differ, preventing the recognition of the PBP, as is the case with methicillin-resistant Staphylococcus aureus [3].
Ampicillin is a penam antibiotic that acts primarily on the cell-wall synthesis of Gram-positive bacteria, since the peptidoglycan is not protected by the outer membrane, and on some Gram-negative bacteria by using the process of passive diffusion through porins to reach the peptidoglycan. This broad spectrum antibiotic is widely used in molecular biology and has an application concentration of 100 µg/ml.
Carbenicillin, which belongs also to the penam family, is also used at a concentration of 100 µg/ml in molecular biology for selection in the laboratory.
The laboratory selection is based on the presence of a β-lactamase gene, the AmpR / bla, in the vector used to genetically modify bacteria, and it’s absence in the parental strain.
A panoply of vectors containing the AmpR gene are used to genetically modify bacteria. They have different origins of replications which define their copy number per cell and may influence the amount of β-lactamase produced and thus the amount of antibiotic a cell can resist.
We therefore were interested to know if the Gram-negative bacterium Escherichia coli could resist higher concentrations of Ampicillin or Carbenicillin, while taking into consideration the copy number of the plasmid harbouring the AmpR resistance gene.
To investigate this, E. coli NEB5ɑ cells were first grown overnight in 96-deep-well plates containing 1 mL of LB medium supplemented with 100 µg/mL ampicillin, then diluted by 40x into similar media. Upon reaching early log-phase, cells were further diluted 20x in 100 µL of LB medium supplemented with increasing concentrations of either ampicillin or carbenicillin (0 to 15000 µg/mL) in a transparent 96-well polystyrene microplate (Sarstedt). The plate was incubated at 37 °C at 200 rpm and ODsub>600nm</sub> was measured every 10 min for 24 hours in a CLARIOstar (BMGLabtech) plate reader. The ODsub>600nm</sub> values at the time point when the cultures performed in the absence of antibiotics reached the stationary phase were plotted as a function of the antibiotic concentration. These dose-response curves were fitted using the AAT Bioquest, Inc., Quest Graph™ IC50 Calculator to the sigmoidal équation:
where OD600nmmin represents the minimal OD600nm, OD600nmmax represents the maximal OD600nm, C is the concentration of the antibiotic, IC50 is the antibiotic concentration that inhibits 50% of the bacterial growth, Hill coefficient is describing the steepness of the curve. The OD600nm values were converted into equivalent particles using the calibration curves presented on the "Measurement" page of our wiki.
The results obtained (Figure 2) for the median inhibitory concentration IC50 for the two antibiotics tested show us that E. coli bacteria harbouring the AmpR gene can resist 10 to 100 times higher antibiotic concentrations than those usually used in molecular biology techniques. When we focus on the inhibitory concentration as a function of the copy number provided by the plasmid (Figure 3), we observe no correlation, suggesting the resistance efficacy of the bacteria to ampicillin or carbenicillin is not closely dependent on the copy number of this gene per cell.
Figure 2. IC50 values of ampicillin and carbenicillin in E. coli NEB5ɑ cells harboring the AmpR resistance gene in plasmids with various origins of replication. The data and error bars are the mean and standard deviation of at least three measurements on independent biological replicates.
Figure 3. IC50 values of ampicillin and carbenicillin as a function of the AmpR gene copy number.
References
[1] Salverda MLM, De Visser JAGM, Barlow M. Natural evolution of TEM-1 β-lactamase: experimental reconstruction and clinical relevance. FEMS microbiology reviews (2010) 34: 1015–1036.
[2] Bush K, Bradford PA. β-Lactams and β-Lactamase Inhibitors: An Overview. Cold Spring Harbor Perspectives in Medicine (2016) 6: a025247.
[3] Chambers HF. Methicillin-resistant Staphylococcus aureus. Mechanisms of resistance and implications for treatment. Postgraduate Medicine (2001) 109: 43–50.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 716
| None |