Part:BBa_K2278022
Lecrocin I antimicrobial peptide with Alpha-Factor Secretion Signal
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal XhoI site found at 244
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Introduction
This DNA biobrick was designed in order to produce Leucrocin I antimicrobial peptide.
1- Biological background
Antimicrobial peptides (AMP) are phylogenetically ancient components of the innate defense of both invertebrates and vertebrates. In the context of growing bacterial antibiotic-resistance, these AMP are considered as potential new therapeutical candidates.Leucrocin I from Siamese crocodile white blood cells shows a good antibacterial activity towards Vibrio cholerae. Leucrocin I action has been observed with fluorescence and electron microscopy. The molecule is cationic and can target bacterial membranes. It creates pores in these membranes leading to the cell lysis. The peptide is a 7 amino acid residue : NGVQPKY with a molecular mass around 806.99 Da.
2- Usage in iGEM projects
The part was designed during the Croc’n Cholera project (team INSA-UPS-France 2017). It produces the leucrocin I AMP when associated with a yeast promoter. The α-factor (BBa_K1800001) sequence contains a RBS and a signal sequence to secrete the produced peptides.Experiments
1- Molecular biology
The gene was placed under the control of an alpha factor signal. IDT performed the DNA synthesis and delivered the part as gBlock. The construct was cloned by conventional ligation into the pSB1C3 plasmid.
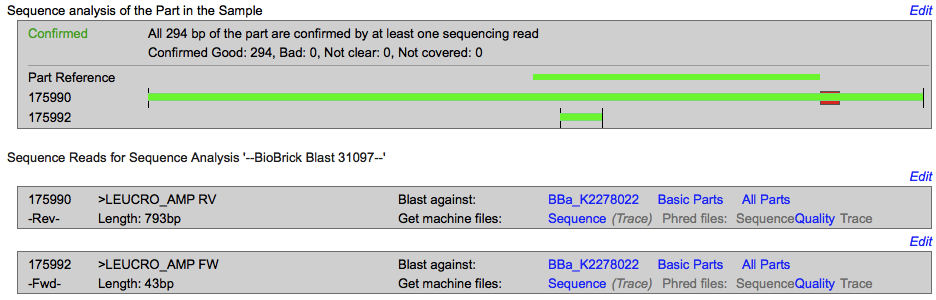
Analysis of the restriction map
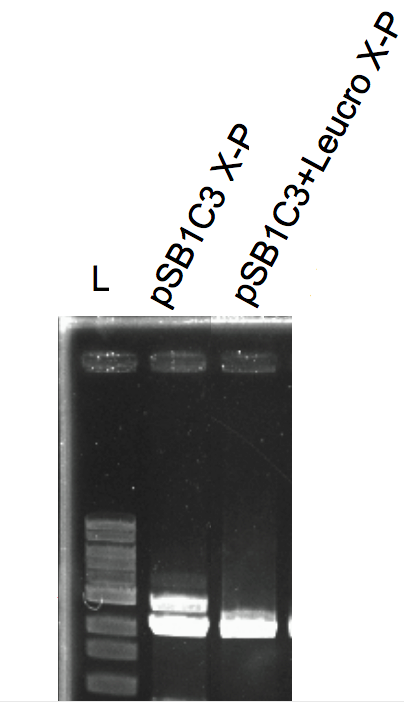
Sequencing
The sequencing successfully validated the sequence of the biobrick.
2- Integration in Pichia pastoris
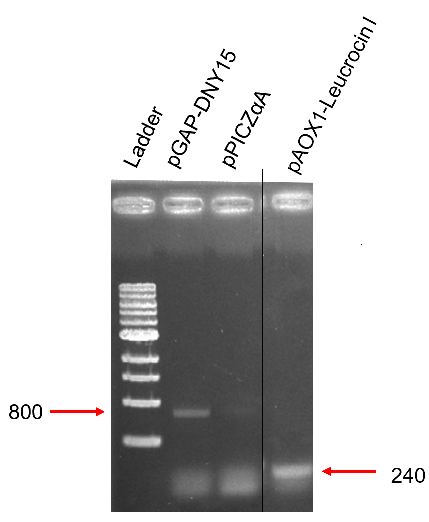
The biobrick was placed under the control of the methanol inducible pAOXI promoter (BBa_K431007) and was cloned in the pPICZalpha vector, an expression vector for the yeast Pichia pastoris. The plasmid was then linearized and transferred in Pichia pastoris by electroporation. The integration is done by using pAOXI homology region (present on pPICZalpha). Indeed, the pAOXI promoter made genome recombination easier in Pichia pastoris.
Correct amplifications were observed for the 2 tested colonies A & B (in spite of lower size non-specific fragments). Negative control with pPICZalpha did not present the correct size fragment, as expected.
Characterization
2. Toxicity assay
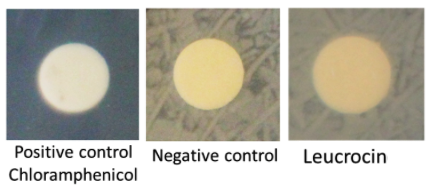
Leucrocin production was performed with Pichia pastoris rown for 4 days at 30 °C with shacking in YPD 40 g/L glucose plus methanol to trigger the pAOX1 promoter. Supernatants from yeasts with or without the leucrocin encoding gene were sampled. The supernatants were used in a halo assay against V. harveyi as the target of Leucrocin. Briefly, 35mL of supernatants were freeze-dried and then resuspended in 3.5mL of water. A paper cut was soaked with one of these solutions and placed on a Petri plate inoculated with V. harveyi (figure 3).
Conclusion :
No inhibition halo was observed around the yeast patch. The Leucrocin I cytoxicity can not be demonstrated.Perspectives:
Higher concentration of yeast supernatants could be tried.Design Notes
BBa_K1800001 : Alpha-Factor Secretion Signal
Source
The peptides DNA sequence has been obtained by reverse translate the amino acid sequence of the Leucrocin I proposed by Pata et al., 2011. Pata et al., 2011 had determinated the amino acid sequence by mass spectrometry analysis.
References
Pata, S., Yaraksa, N., Daduang, S., Temsiripong, Y., Svasti, J., Araki, T. and Thammasirirak, S. (2011). Characterization of the novel antibacterial peptide Leucrocin from crocodile (Crocodylus siamensis) white blood cell extracts. Developmental & Comparative Immunology, 35(5), pp.545-553.
| None |