Difference between revisions of "Part:BBa K3823008"
Jiayi Chen (Talk | contribs) |
Jiayi Chen (Talk | contribs) |
||
| Line 18: | Line 18: | ||
[[Image:BBa K3823008-2.jpeg|400px|center|thumb|Figure 2. Reduced CstR binds to cst operator (CstO) sites with high affinity in a manner dependent of the central run of four GC base pairs. A, schematic of the cstR-cstA intergenic region highlighting the positions of the two tandem candidate CstR operator sites. B, nucleotide sequence of the cstR-cstA intergenic region, highlighting the OP1 and OP2 operator sequences (green). Underlined bases correspond to the 5′-fluorescein-labeled duplex oligonucleotides used for DNA binding experiments. C, CstR binding isotherms for OP1 (solid squares, ■), OP2 (open circles, ○), and OP1_5GC (filled circles, ●) in which an additional GC base pair was inserted into the run of four GC base pairs. Inset, competition dissociation experiments with fluorescein-labeled apo-CstR-OP1 complexes with unlabeled wild-type OP1 (solid squares, ■), OP1-GC3 (open triangles, △), and OP1_1GC (open squares, □) duplexes. Conditions used were pH 7.0, 0.2 m NaCl, at 25.0 °C.<sup>[4]</sup>]] | [[Image:BBa K3823008-2.jpeg|400px|center|thumb|Figure 2. Reduced CstR binds to cst operator (CstO) sites with high affinity in a manner dependent of the central run of four GC base pairs. A, schematic of the cstR-cstA intergenic region highlighting the positions of the two tandem candidate CstR operator sites. B, nucleotide sequence of the cstR-cstA intergenic region, highlighting the OP1 and OP2 operator sequences (green). Underlined bases correspond to the 5′-fluorescein-labeled duplex oligonucleotides used for DNA binding experiments. C, CstR binding isotherms for OP1 (solid squares, ■), OP2 (open circles, ○), and OP1_5GC (filled circles, ●) in which an additional GC base pair was inserted into the run of four GC base pairs. Inset, competition dissociation experiments with fluorescein-labeled apo-CstR-OP1 complexes with unlabeled wild-type OP1 (solid squares, ■), OP1-GC3 (open triangles, △), and OP1_1GC (open squares, □) duplexes. Conditions used were pH 7.0, 0.2 m NaCl, at 25.0 °C.<sup>[4]</sup>]] | ||
| − | To improve its performance in sensitivity, leaky strength, maximal strength, and strength amplitude, Liu, H., et al <sup>[ | + | To improve its performance in sensitivity, leaky strength, maximal strength, and strength amplitude, Liu, H., et al <sup>[1]</sup>constructed a series of combinations of PL and PR(Pcstr), and found the expression from elements 1, 4, and 5 was low and relatively stable (<b>Figure 3</b>), indicating that they are stringent enough for reducing leaky expression caused by background HS<sub>n</sub>H. |
Here is the sequence of the promoter <b>P<sub>R3</sub></b> in Figure 3. | Here is the sequence of the promoter <b>P<sub>R3</sub></b> in Figure 3. | ||
| − | [[Image:BBa K3823008-3.jpeg|thumb|center|400px|Figure 3. Construction and test of HS<sub>n</sub>H sensor-actuator elements. (A) Seven elements were constructed by changing CstR and mKate expression promoters (PL and PR). These elements were transformed into <i>E. coli</i> BL21. (B) Response of these elements to HS<sub>n</sub>H induction. The data were fitted with the Hill equation. The Hill coefficient (n) and the dose inducing half turn-on (Km) were listed. (C) Leaky expression strength (without HS<sub>n</sub>H induction) and maximal induced strength by 300 μM HS<sub>n</sub>H of the elements. The folds of changes were given in numbers. (D) Performance of the elements in a whole growth circle. <i>E. coli</i> BL21 harboring these elements were cultivated in 200 μL of LB medium in a 96-well plate (37 °C with vibration). mKate fluorescence and cell density were automatically recorded at 30 min intervals using the Synergy H1Microplate reader. Data shown are normalized fluorescence (fluorescence intensity/OD<sub>600 nm</sub>).<sup>[ | + | [[Image:BBa K3823008-3.jpeg|thumb|center|400px|Figure 3. Construction and test of HS<sub>n</sub>H sensor-actuator elements. (A) Seven elements were constructed by changing CstR and mKate expression promoters (PL and PR). These elements were transformed into <i>E. coli</i> BL21. (B) Response of these elements to HS<sub>n</sub>H induction. The data were fitted with the Hill equation. The Hill coefficient (n) and the dose inducing half turn-on (Km) were listed. (C) Leaky expression strength (without HS<sub>n</sub>H induction) and maximal induced strength by 300 μM HS<sub>n</sub>H of the elements. The folds of changes were given in numbers. (D) Performance of the elements in a whole growth circle. <i>E. coli</i> BL21 harboring these elements were cultivated in 200 μL of LB medium in a 96-well plate (37 °C with vibration). mKate fluorescence and cell density were automatically recorded at 30 min intervals using the Synergy H1Microplate reader. Data shown are normalized fluorescence (fluorescence intensity/OD<sub>600 nm</sub>).<sup>[1]</sup>]] |
| Line 39: | Line 39: | ||
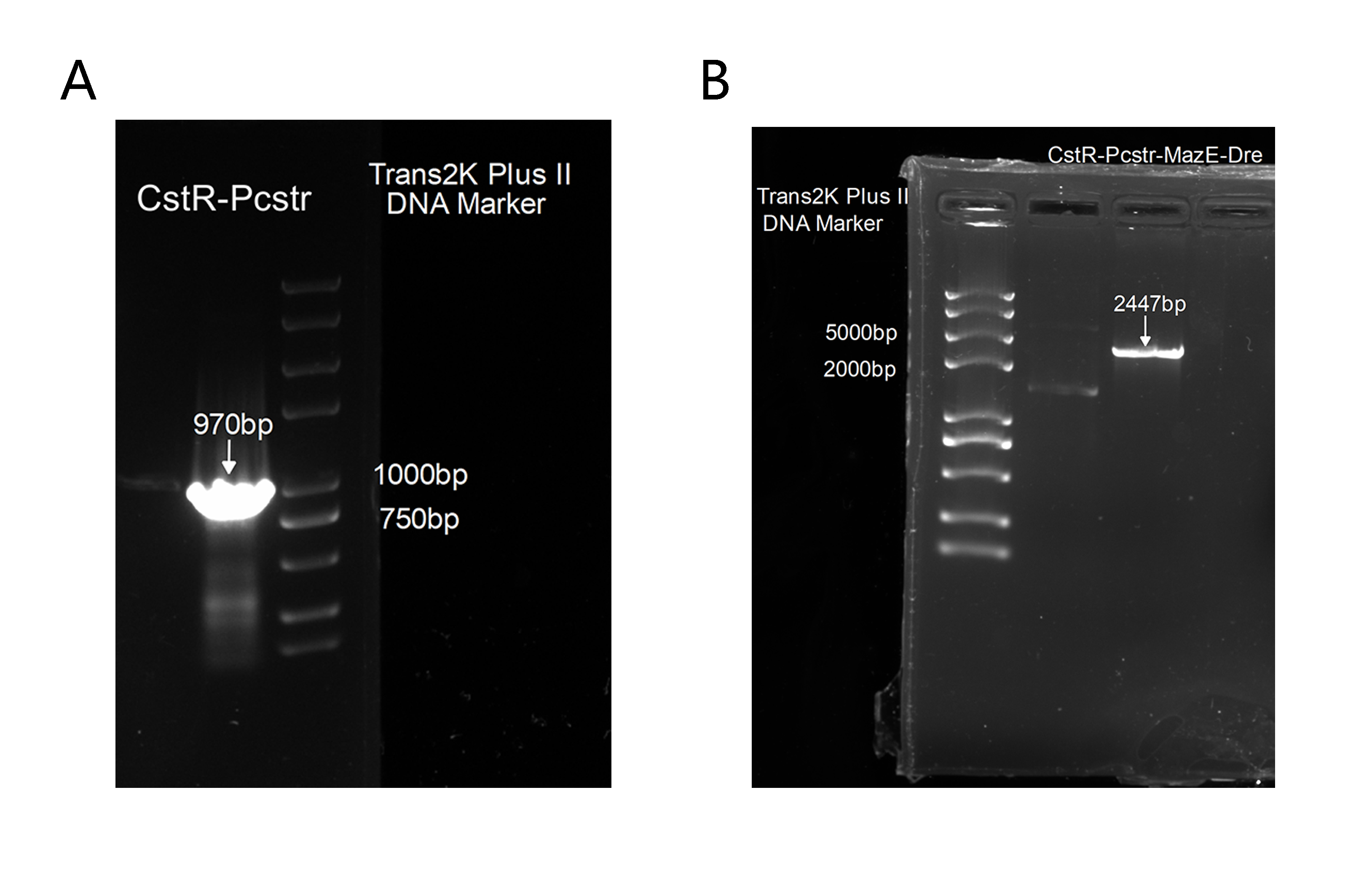
2. Also, to get this part for our plasmid construction, we successfully amplified CstR-Pcstr(<b>Figure 6A</b>), which was proved correct by sequencing and we has cloned it into our backbone to construct CstR-Pcstr-MazE-Dre(<b>Figure 6B</b>),which was also proved correct by sequencing. | 2. Also, to get this part for our plasmid construction, we successfully amplified CstR-Pcstr(<b>Figure 6A</b>), which was proved correct by sequencing and we has cloned it into our backbone to construct CstR-Pcstr-MazE-Dre(<b>Figure 6B</b>),which was also proved correct by sequencing. | ||
[[Image:BBa K3823006-4.png|600px|center|thumb|Figure 6. Agarose electrophoresis]] | [[Image:BBa K3823006-4.png|600px|center|thumb|Figure 6. Agarose electrophoresis]] | ||
| + | |||
| + | *[1]Grossoehme, N., et al., Control of Copper Resistance and Inorganic Sulfur Metabolism by Paralogous Regulators in Staphylococcus aureus. Journal of Biological Chemistry, 2011. 286(15): p. 13522-13531. | ||
| + | *[2]Giedroc, D.P., A new player in bacterial sulfide-inducible transcriptional regulation. Mol Microbiol, 2017. 105(3): p. 347-352. | ||
| + | *[3]Liu, H., et al., Synthetic Gene Circuits Enable Escherichia coli To Use Endogenous H(2)S as a Signaling Molecule for Quorum Sensing. ACS Synth Biol, 2019. 8(9): p. 2113-2120. | ||
| + | *[4]Grossoehme, N., et al., Control of Copper Resistance and Inorganic Sulfur Metabolism by Paralogous Regulators in Staphylococcus aureus. Journal of Biological Chemistry, 2011. 286(15): p. 13522-13531. | ||
<!-- Add more about the biology of this part here | <!-- Add more about the biology of this part here | ||
Revision as of 14:37, 20 October 2021
Pcstr: an artificial hydrogen sulfide sensitive promoter with binding sequences of CstR
Pcstr: An artificial promoter with two kinds of binding sequences of CstR from the natural Pcstr.It is sensitive to hydrogen sulfide density.
Introduction
CstR
CstR(BBa_K3823006)is a CsoR-like sulfur transferase repressor found in Staphylococcus aureus[1], reported to react with HSnH and form sulfhydrated proteins, which can turn on their regulated genes[2] . Compared with other HSnH sensors, it shows significant response to HSnH induction.And it shows no response to H2O2, a structural analogue of HSSH and a common intracellular metabolite(Figure 1.).[3]

The structure of the Pcstr
Grossoehme, N., et al. found two strong candidate tandemly repeated CstR operator sites, CstO OP1 and OP2, between the cstR and cstA genes. These sites were characterized by a run of four consecutive GC-base pairs and flanked by AT-rich regions(Figure 2.) [4].

To improve its performance in sensitivity, leaky strength, maximal strength, and strength amplitude, Liu, H., et al [1]constructed a series of combinations of PL and PR(Pcstr), and found the expression from elements 1, 4, and 5 was low and relatively stable (Figure 3), indicating that they are stringent enough for reducing leaky expression caused by background HSnH. Here is the sequence of the promoter PR3 in Figure 3.

Results
1. To figure out how CstR works by sensing H2S, we characterized the function of CstR with Pcstr using pTrchis2A-CstR-Pcstr-mKate-CpSQR(donated by Professor Liu from Shandong University). The expression level regulated by Pcstr at different concentrations of S2- was shown by the fluorescence intensity of mKate(/OD600 nm).
At the beginning, bacteria cultured in 5 mL were treated with 0, 6.5, 13.0, 26.0 and 39.0 mg/L Na2S (0, 20, 40, 80 and 120 mg/L Na2S·9H2O) respectively, and then the fluorescence intensity of mKate was measured with a microplate reader(Figure 4.).
It can be seen that the presence of S2- at different concentrations has no significant effect on the growth of bacteria. However, with the increase of S2-, the fluorescence intensity increases at first and then decreases, indicating that when S2- is too high, the lifting effect on CstR inhibition is weakened. We hope to find out an appropriate concentration range in which the strength of Pcstr is positively correlated with the concentration of S2-, which tells us that we need to reduce the concentration gradient and concentration range for further characterization.
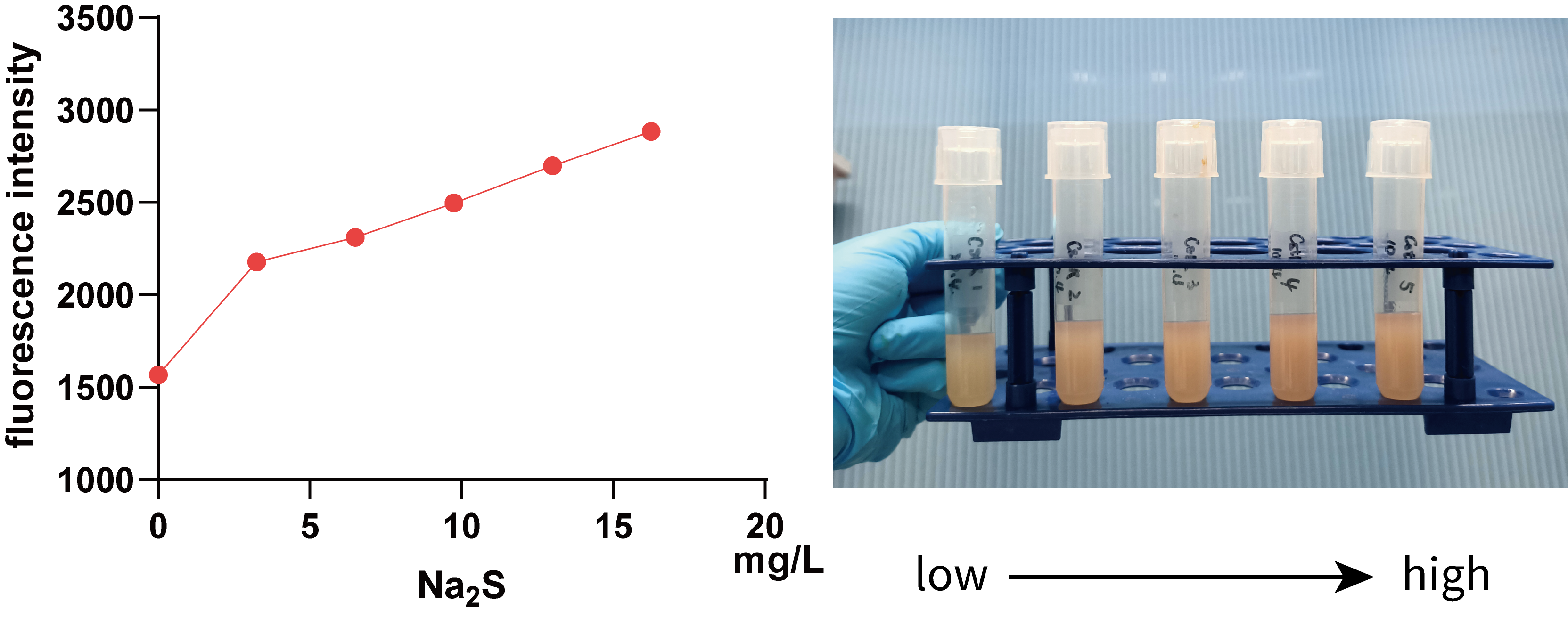
We further used 0, 3.25, 6.50, 9.75, 13.00, 16.25 mg/L Na2S (0, 10, 20, 30, 40 and 50 mg/L Na2S·9H2O) to treat 5 mL bacterial solution and then measured the fluorescence intensity of mKate using a microplate reader(Figure 5.).
It can be seen from the figure that in a smaller concentration range, there is a positive correlation between fluorescence intensity and Na2S concentration, visible to naked eyes, which implies that we can use data within this range to regulate Pcstr by giving different concentration H2S.
2. Also, to get this part for our plasmid construction, we successfully amplified CstR-Pcstr(Figure 6A), which was proved correct by sequencing and we has cloned it into our backbone to construct CstR-Pcstr-MazE-Dre(Figure 6B),which was also proved correct by sequencing.
- [1]Grossoehme, N., et al., Control of Copper Resistance and Inorganic Sulfur Metabolism by Paralogous Regulators in Staphylococcus aureus. Journal of Biological Chemistry, 2011. 286(15): p. 13522-13531.
- [2]Giedroc, D.P., A new player in bacterial sulfide-inducible transcriptional regulation. Mol Microbiol, 2017. 105(3): p. 347-352.
- [3]Liu, H., et al., Synthetic Gene Circuits Enable Escherichia coli To Use Endogenous H(2)S as a Signaling Molecule for Quorum Sensing. ACS Synth Biol, 2019. 8(9): p. 2113-2120.
- [4]Grossoehme, N., et al., Control of Copper Resistance and Inorganic Sulfur Metabolism by Paralogous Regulators in Staphylococcus aureus. Journal of Biological Chemistry, 2011. 286(15): p. 13522-13531.
Sequence and Features
- 10INCOMPATIBLE WITH RFC[10]Illegal EcoRI site found at 31
- 12INCOMPATIBLE WITH RFC[12]Illegal EcoRI site found at 31
- 21INCOMPATIBLE WITH RFC[21]Illegal EcoRI site found at 31
- 23INCOMPATIBLE WITH RFC[23]Illegal EcoRI site found at 31
- 25INCOMPATIBLE WITH RFC[25]Illegal EcoRI site found at 31
- 1000COMPATIBLE WITH RFC[1000]