Difference between revisions of "Part:BBa K3457042"
| Line 11: | Line 11: | ||
[[File:T--QHFZ-China--LEA-11.png|400px|thumb|left|Figure 1. The Schematic cartoon of the DNA construct of BBa_K3457042.]] | [[File:T--QHFZ-China--LEA-11.png|400px|thumb|left|Figure 1. The Schematic cartoon of the DNA construct of BBa_K3457042.]] | ||
| − | <p style="clear:left;"> This part should be composed of a T7 promoter, RBS B0034, CDS of LEA [https://parts.igem.org/Part:BBa_K3457040 BBa_K3457040] | + | <p style="clear:left;"> This part should be composed of a T7 promoter, RBS B0034, CDS of LEA [https://parts.igem.org/Part:BBa_K3457040 BBa_K3457040], and a T7 terminator. However, we added another terminator B0015 before the T7 terminator. The reason is that if the T7 promoter is changed into other promoters, such as J23100, the corresponding RNA polymerase changed from T7 RNAP to endogenous RNAP of <i>E. coli</i>, which can be stopped by terminator B0015. In other words, the added terminator made the part easier to be modified. In the cartoon (Fig. 1), there are two points around the T7 promoter. They are two <i>Bsa</i>I cutting site, which can be used to change the T7 promoter by Golden Gate technique.</p> |
<h3><b>Documentation:</b></h3> | <h3><b>Documentation:</b></h3> | ||
Latest revision as of 00:48, 27 October 2020
T7-Olac-RBS-LEA
This biological part is the expression sequence of a Late Embryogenesis Abundant (LEA) protein, LEA3 protein, from soybean (Glycine max). It is a protein which help seeds resist dessication.
Contribution
Group: QHFZ-China iGEM 2020
Author: Yixian Yang
Design
This part should be composed of a T7 promoter, RBS B0034, CDS of LEA BBa_K3457040, and a T7 terminator. However, we added another terminator B0015 before the T7 terminator. The reason is that if the T7 promoter is changed into other promoters, such as J23100, the corresponding RNA polymerase changed from T7 RNAP to endogenous RNAP of E. coli, which can be stopped by terminator B0015. In other words, the added terminator made the part easier to be modified. In the cartoon (Fig. 1), there are two points around the T7 promoter. They are two BsaI cutting site, which can be used to change the T7 promoter by Golden Gate technique.
Documentation:
Introduction:
This year, we tried to introduce a new biopreservation method. We used freeze-drying to make the engineered into dry powder. Then the powder can be stored at room temperature for a long time. This method can make the storage of bacteria get rid of ultra-low temperature freezer, so that it will promote the practical application of engineered bacteria out of laboratory. However, the stresses during freeze-drying and subsequent dry storage, including freeze, dry and vacuum, are lethal to bacteria. We use TDPs to help bacteria survive the situation. We also tested the effect of LEA3 for comparison.
Protocol:
To test the effect of LEA3, we modified a frequently and widely used vector, pet28a+ and put this part or TDP genes into it (Fig. 2). Then we transformed the plasmid into E. coli BL21 strain.
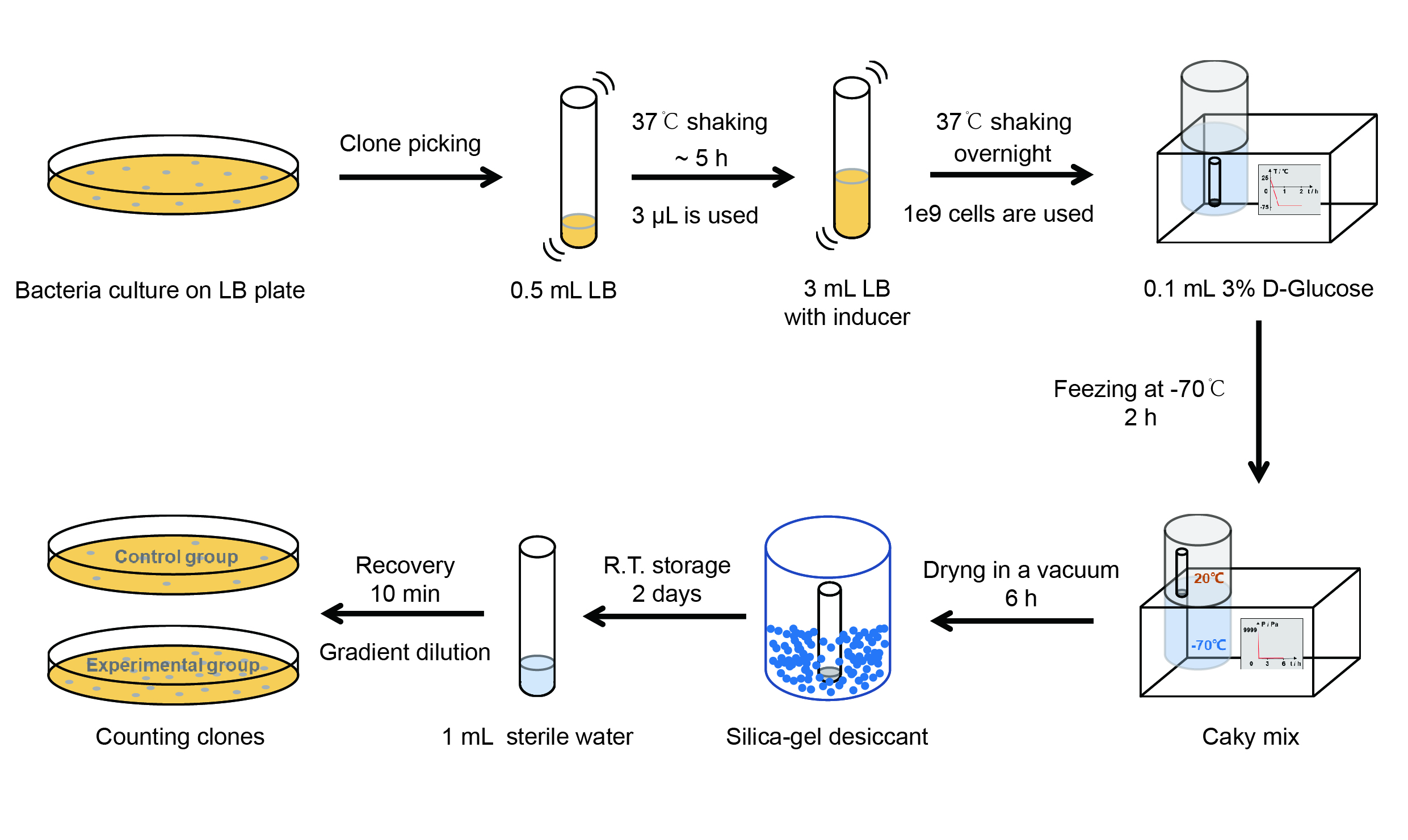
Then we used the following protocols to verify its function (Fig. 3):
【Day 1】Induction culture
(1) Pick clones which are in good condition and put them into 500 μL LB medium containing antibiotics. Shake them to grow at 37℃ for 5~7 hours until the bacteria solution becomes turbid.
(2) Add 2 mM iPTG into 3 mL LB medium containing antibiotics. Add 3 μL of the bacteria solution mentioned in step 1 to dilute the bacteria by the ratio of 1:1000. Shake the solution to grow the bacteria at 37℃ overnight.
【Day 2】Freeze-dried
(1) If fluorescence induced by the iPTG is detectable in the control group (GFP), continue conducting the experiment.
(2) Use spectrophotometer to measure the OD600 of the bacteria solution, OD600 = 1 equals to 109 cells. If the OD600 value is between 0.1 and 1, There is a linear relationship between OD600 and bacterial density. Calculate the volume of bacterial solution for 109 cells by using the formula V = 100 / (OD600 × Dilution ratio).
(3) Take out a measured amount of 109 cells and centrifuge it at 8000 rpm for 3 min. Then pour out the supernatant.
(4) Resuspend the bacteria in a 15 mL tube with pre-refrigerated 100 μL 3% glucose solution.
(5) Take off the cover of the tube and put the bacteria into the cold trap. Open the compressor of the lyophilization machine and freeze the shake tube for 2 h at -70℃.
(6) Put the caky bacteria solution into the drying chamber of the lyophilization machine. Open the vacuum pump to dry it in vacuum for 6h at 1 Pa vacuum degree.
(7) Turn off the vacuum pump, place it at seal box filled with silica-gel desiccant a for 2 days at room temperature.
【Day 3】Room temperature storage
【Day 4】Detect the survival rate
(1) Add 1 mL of sterile water to the tube, vortex for 15 s, placed it at room temperature for 10 min.
(2) Adjust the density of the bacteria solution by gradient dilution, then spread 100 μL of the bacteria solution on the LB plate.
(3) If the density above is not suitable, take 100μL of the solution and spread it on the LB plate after several gradient dilutions.
(4) Culture the bacteria overnight at 37℃.
【Day 5】Cell Count
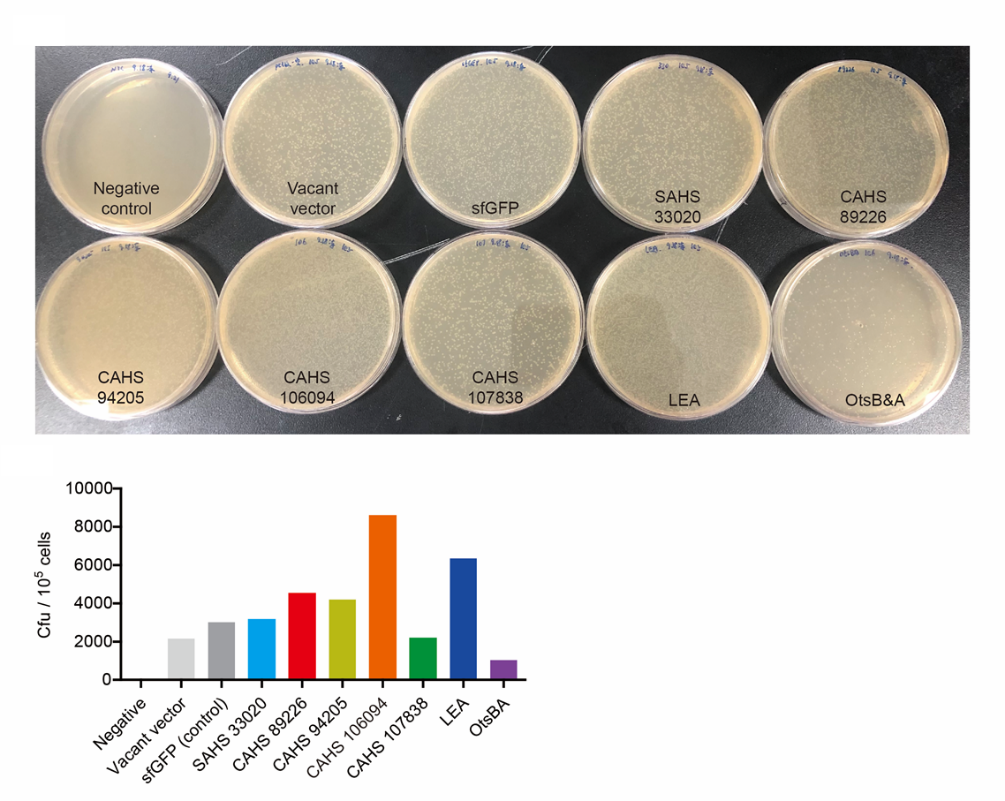
(1) Take out the LB plate and take photos to record experimental results.
(2) Use the automatic cell counting function of Image J to count the colone number on the LB plate, then compare the results between each group.
Results:
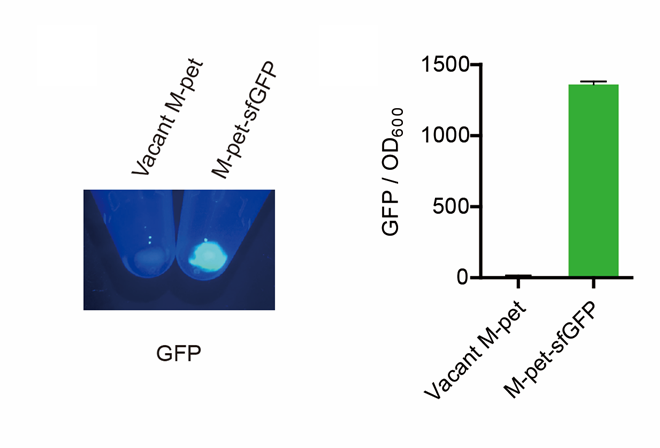
First, by a reporter, sfGFP, we confirmed that the plasmid can normally expressed exogenous proteins in E. coli BL21 strain (Fig. 4).
Summary:
LEA3 helps the bacteria survive freeze-drying and subsequent dry storage. To achieve that, you need only to transform the sequence into your bacteria. You can also use the part to express and purify LEA3 through Ni-chelating affinity chromatography.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 1647
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 27
Illegal BsaI.rc site found at 2
Illegal SapI.rc site found at 129