Difference between revisions of "Part:BBa K2656023"
| Line 1: | Line 1: | ||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K2656023 short</partinfo> | <partinfo>BBa_K2656023 short</partinfo> | ||
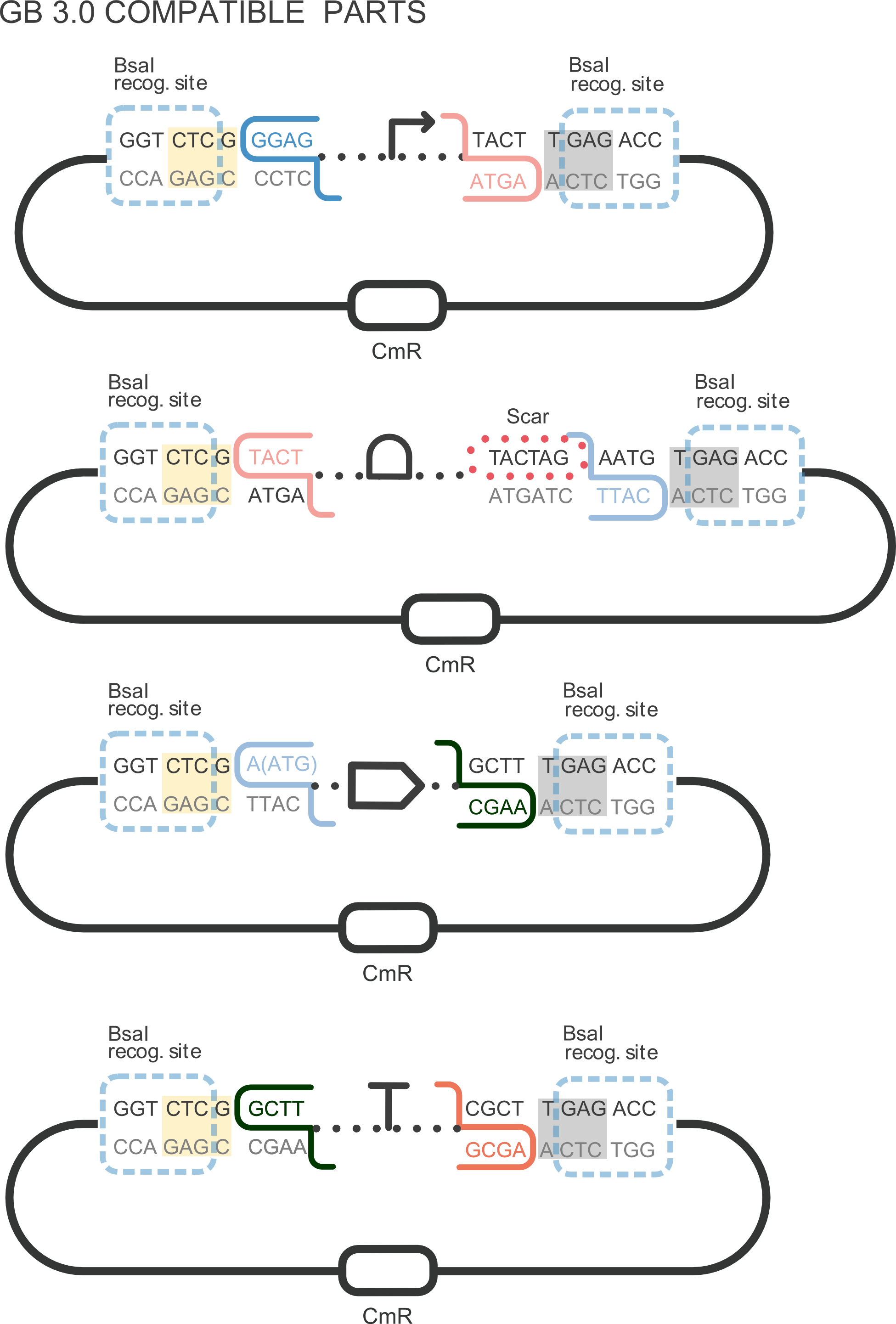
| − | + | [[File:T--Valencia_UPV--im11UPV2018.png|250px|thumb|right|alt=domestication.|Figure 1. DNA basic parts domestication. Third construction refers to CDS GB domestication. ]] | |
Part BBa_K2656013 is the GFPmut3b coding sequence with the LVA degradation tag. It is the [https://parts.igem.org/Part:BBa_K1399004 BBa_K1399004] original sequence adapted to be Golden Gate compatible. Thus, it can be used with Biobrick and [http://2018.igem.org/Team:Valencia_UPV/Design Golden Braid] assembly methods. | Part BBa_K2656013 is the GFPmut3b coding sequence with the LVA degradation tag. It is the [https://parts.igem.org/Part:BBa_K1399004 BBa_K1399004] original sequence adapted to be Golden Gate compatible. Thus, it can be used with Biobrick and [http://2018.igem.org/Team:Valencia_UPV/Design Golden Braid] assembly methods. | ||
The LVA tag causes an increase in protease activity and, therefore, a faster degradation of the reporter protein in the cell. | The LVA tag causes an increase in protease activity and, therefore, a faster degradation of the reporter protein in the cell. | ||
Latest revision as of 03:19, 18 October 2018
GFPmut3b_LVA Coding Sequence
Part BBa_K2656013 is the GFPmut3b coding sequence with the LVA degradation tag. It is the BBa_K1399004 original sequence adapted to be Golden Gate compatible. Thus, it can be used with Biobrick and [http://2018.igem.org/Team:Valencia_UPV/Design Golden Braid] assembly methods. The LVA tag causes an increase in protease activity and, therefore, a faster degradation of the reporter protein in the cell.
It can be combined with other compatible parts from our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection Valencia UPV IGEM 2018 Printeria Collection] to assemble transcriptional units with a BsaI one-step reaction with the Golden Gate assembly protocol .
Sequence and Features
Assembly Compatibility:
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]