Difference between revisions of "Part:BBa K2460007"
| Line 3: | Line 3: | ||
<partinfo>BBa_K2460007 short</partinfo> | <partinfo>BBa_K2460007 short</partinfo> | ||
| − | This ORF codes for the | + | This ORF codes for the steptomyces phage TG1 integrase, which catalyzes a site specific recombination between TG1 attB ([[Part:BBa_K2460008|BBa_K2460008]]) and attP ([[Part:BBa_K2460009|BBa_K2460009]]) sequences. This recombinase can be used to flip a DNA sequence flanked by attB and attP sites, as well as integrating a circular DNA with a attB/P site into a linear/circular DNA with a attP/B site. |
=Biological Characterisation= | =Biological Characterisation= | ||
| Line 21: | Line 21: | ||
==Results== | ==Results== | ||
| − | Due to the leakage before inducing, the LR rate changing is not obvious, which means the TG1 integrase is | + | Due to the leakage before inducing, the LR rate changing is not obvious, which means the TG1 integrase is highly efficient, and the leakage may be reduced by improving the genetic design. |
[[File:Fudan_China_TG1.jpg]] | [[File:Fudan_China_TG1.jpg]] | ||
| Line 38: | Line 38: | ||
==Results== | ==Results== | ||
| − | Expect for the TG1 attB/P site, the | + | Expect for the TG1 attB/P site,none of the attB/P sites flipped, indicating that TG1 integrase has a good orthogonality with other attB/P sites and compatible with other integrases. |
[[File:Fudan_China_Orthogonality.jpg]] | [[File:Fudan_China_Orthogonality.jpg]] | ||
Revision as of 18:40, 1 November 2017
TG1 integrase
This ORF codes for the steptomyces phage TG1 integrase, which catalyzes a site specific recombination between TG1 attB (BBa_K2460008) and attP (BBa_K2460009) sequences. This recombinase can be used to flip a DNA sequence flanked by attB and attP sites, as well as integrating a circular DNA with a attB/P site into a linear/circular DNA with a attP/B site.
Biological Characterisation
TG1 integrase is a member of the large serine recombinases(LSRs) family. LSRs are recombinases encoded by temperate phage genomes, which precisely cut and recombine DNA in a highly controllable and predictable way. The recombinases recognize and bind to attB/P sites as dimers, and cut and recombinate on the attB/P sites, with two dimers forming a tetramer. The recombinated attB/P sites form attL and attR sites.
Kinetic Characterization
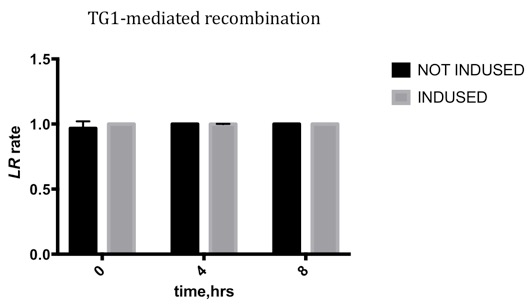
We conducted qPCR to record the ratio of attL/R to the sum of attL/R and attB/P, to characterize the efficiency varying with time.
Genetic design
- The integrase is expressed under a PBad promoter (BBa_I13453). Considering the leakage of PBad promoter would lead to flipping,the promoter is followed by a weaker RBS (BBa_B0033). Ahead of the promoter,an attB and attP are opposite to each other,flanking a 50 bp random sequence. This construct was cloned on a medium copy plasmid with p15A replication of origin and kanamycin resistance.
- To ensure a low leakage when not induced, AraC is expressed under a constitutive promoter, cloned on a high copy plasmid with pUC19 replication of origin and Ampicillin resistance.
Experimental Setup
- Since transforming the two plasmid above at a time was unable to prevent the flipping, the plasmid with AraC was first transformed into E.coli DH10b, then the plasmid with the integrase was transformed into the competent cell with the AraC plasmid.
- After culturing for over 2 hours in 2×YT borth, the cells were induced in 1% Arabinose for 8 hours. We sampled the culture at 0 hour, the 4th hour and the 8th hour.
- We calculated flipped and unflipped random sequence by qPCR.
Results
Due to the leakage before inducing, the LR rate changing is not obvious, which means the TG1 integrase is highly efficient, and the leakage may be reduced by improving the genetic design.
Orthogonality Characterization
Since our circuit are using multiple integrases, it's important that the attB/P site of one integrase won't be flipped by one another integrase. We conducted orthogonality tests to see the compatibility between our integrases, including phiBT1 (BBa_K2460001), Bxb1 (BBa_K907000), phiC31 (BBa_K1039012), phiRv1 (BBa_K2460004) and TG1 (BBa_K2460007) integrase.
Genetic design
- 5 pairs of attB/P sites are each flanking a 50 bp random sequence, lining up ahead of the PBad promoter (BBa_I13453), an RBS (BBa_B0033) and an integrase. This construct was cloned on a medium copy plasmid with p15A replication of origin and kanamycin resistance.
- The same AraC plasmid in Kinetic Characterization was used in orthogonality Characterization.
Experimental Setup
- The same method of transformation in Kinetic Characterization was used in Kinetic Characterization.
- After culturing for over 2 hours in 2×YT borth, the cells were induced in 1% Arabinose 16 hours.
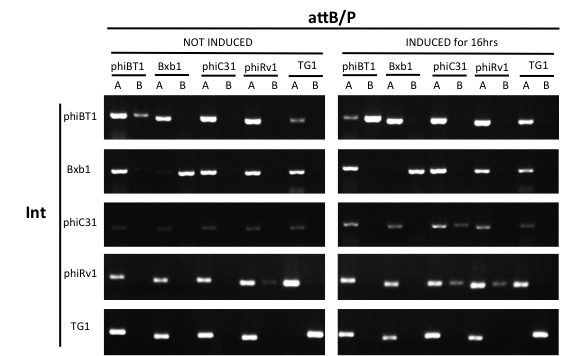
- Conduct PCR for each integrase's test and control group with primers from 5 pairs of attB/P sites, 2 pairs of primers for each pair of attB/P site.
Results
Expect for the TG1 attB/P site,none of the attB/P sites flipped, indicating that TG1 integrase has a good orthogonality with other attB/P sites and compatible with other integrases.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 1725
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 1547
- 1000INCOMPATIBLE WITH RFC[1000]Illegal SapI site found at 1761
Illegal SapI.rc site found at 352
Illegal SapI.rc site found at 1513